Xiaosong Wang
Cost-effective Instruction Learning for Pathology Vision and Language Analysis
Jul 25, 2024



Abstract:The advent of vision-language models fosters the interactive conversations between AI-enabled models and humans. Yet applying these models into clinics must deal with daunting challenges around large-scale training data, financial, and computational resources. Here we propose a cost-effective instruction learning framework for conversational pathology named as CLOVER. CLOVER only trains a lightweight module and uses instruction tuning while freezing the parameters of the large language model. Instead of using costly GPT-4, we propose well-designed prompts on GPT-3.5 for building generation-based instructions, emphasizing the utility of pathological knowledge derived from the Internet source. To augment the use of instructions, we construct a high-quality set of template-based instructions in the context of digital pathology. From two benchmark datasets, our findings reveal the strength of hybrid-form instructions in the visual question-answer in pathology. Extensive results show the cost-effectiveness of CLOVER in answering both open-ended and closed-ended questions, where CLOVER outperforms strong baselines that possess 37 times more training parameters and use instruction data generated from GPT-4. Through the instruction tuning, CLOVER exhibits robustness of few-shot learning in the external clinical dataset. These findings demonstrate that cost-effective modeling of CLOVER could accelerate the adoption of rapid conversational applications in the landscape of digital pathology.
Pathology-knowledge Enhanced Multi-instance Prompt Learning for Few-shot Whole Slide Image Classification
Jul 15, 2024



Abstract:Current multi-instance learning algorithms for pathology image analysis often require a substantial number of Whole Slide Images for effective training but exhibit suboptimal performance in scenarios with limited learning data. In clinical settings, restricted access to pathology slides is inevitable due to patient privacy concerns and the prevalence of rare or emerging diseases. The emergence of the Few-shot Weakly Supervised WSI Classification accommodates the significant challenge of the limited slide data and sparse slide-level labels for diagnosis. Prompt learning based on the pre-trained models (\eg, CLIP) appears to be a promising scheme for this setting; however, current research in this area is limited, and existing algorithms often focus solely on patch-level prompts or confine themselves to language prompts. This paper proposes a multi-instance prompt learning framework enhanced with pathology knowledge, \ie, integrating visual and textual prior knowledge into prompts at both patch and slide levels. The training process employs a combination of static and learnable prompts, effectively guiding the activation of pre-trained models and further facilitating the diagnosis of key pathology patterns. Lightweight Messenger (self-attention) and Summary (attention-pooling) layers are introduced to model relationships between patches and slides within the same patient data. Additionally, alignment-wise contrastive losses ensure the feature-level alignment between visual and textual learnable prompts for both patches and slides. Our method demonstrates superior performance in three challenging clinical tasks, significantly outperforming comparative few-shot methods.
OpenMEDLab: An Open-source Platform for Multi-modality Foundation Models in Medicine
Mar 04, 2024



Abstract:The emerging trend of advancing generalist artificial intelligence, such as GPTv4 and Gemini, has reshaped the landscape of research (academia and industry) in machine learning and many other research areas. However, domain-specific applications of such foundation models (e.g., in medicine) remain untouched or often at their very early stages. It will require an individual set of transfer learning and model adaptation techniques by further expanding and injecting these models with domain knowledge and data. The development of such technologies could be largely accelerated if the bundle of data, algorithms, and pre-trained foundation models were gathered together and open-sourced in an organized manner. In this work, we present OpenMEDLab, an open-source platform for multi-modality foundation models. It encapsulates not only solutions of pioneering attempts in prompting and fine-tuning large language and vision models for frontline clinical and bioinformatic applications but also building domain-specific foundation models with large-scale multi-modal medical data. Importantly, it opens access to a group of pre-trained foundation models for various medical image modalities, clinical text, protein engineering, etc. Inspiring and competitive results are also demonstrated for each collected approach and model in a variety of benchmarks for downstream tasks. We welcome researchers in the field of medical artificial intelligence to continuously contribute cutting-edge methods and models to OpenMEDLab, which can be accessed via https://github.com/openmedlab.
Learning A Multi-Task Transformer Via Unified And Customized Instruction Tuning For Chest Radiograph Interpretation
Nov 02, 2023Abstract:The emergence of multi-modal deep learning models has made significant impacts on clinical applications in the last decade. However, the majority of models are limited to single-tasking, without considering disease diagnosis is indeed a multi-task procedure. Here, we demonstrate a unified transformer model specifically designed for multi-modal clinical tasks by incorporating customized instruction tuning. We first compose a multi-task training dataset comprising 13.4 million instruction and ground-truth pairs (with approximately one million radiographs) for the customized tuning, involving both image- and pixel-level tasks. Thus, we can unify the various vision-intensive tasks in a single training framework with homogeneous model inputs and outputs to increase clinical interpretability in one reading. Finally, we demonstrate the overall superior performance of our model compared to prior arts on various chest X-ray benchmarks across multi-tasks in both direct inference and finetuning settings. Three radiologists further evaluate the generated reports against the recorded ones, which also exhibit the enhanced explainability of our multi-task model.
Boosting Dermatoscopic Lesion Segmentation via Diffusion Models with Visual and Textual Prompts
Oct 04, 2023



Abstract:Image synthesis approaches, e.g., generative adversarial networks, have been popular as a form of data augmentation in medical image analysis tasks. It is primarily beneficial to overcome the shortage of publicly accessible data and associated quality annotations. However, the current techniques often lack control over the detailed contents in generated images, e.g., the type of disease patterns, the location of lesions, and attributes of the diagnosis. In this work, we adapt the latest advance in the generative model, i.e., the diffusion model, with the added control flow using lesion-specific visual and textual prompts for generating dermatoscopic images. We further demonstrate the advantage of our diffusion model-based framework over the classical generation models in both the image quality and boosting the segmentation performance on skin lesions. It can achieve a 9% increase in the SSIM image quality measure and an over 5% increase in Dice coefficients over the prior arts.
Text-guided Foundation Model Adaptation for Pathological Image Classification
Jul 27, 2023



Abstract:The recent surge of foundation models in computer vision and natural language processing opens up perspectives in utilizing multi-modal clinical data to train large models with strong generalizability. Yet pathological image datasets often lack biomedical text annotation and enrichment. Guiding data-efficient image diagnosis from the use of biomedical text knowledge becomes a substantial interest. In this paper, we propose to Connect Image and Text Embeddings (CITE) to enhance pathological image classification. CITE injects text insights gained from language models pre-trained with a broad range of biomedical texts, leading to adapt foundation models towards pathological image understanding. Through extensive experiments on the PatchGastric stomach tumor pathological image dataset, we demonstrate that CITE achieves leading performance compared with various baselines especially when training data is scarce. CITE offers insights into leveraging in-domain text knowledge to reinforce data-efficient pathological image classification. Code is available at https://github.com/Yunkun-Zhang/CITE.
Exploring Data Redundancy in Real-world Image Classification through Data Selection
Jun 25, 2023



Abstract:Deep learning models often require large amounts of data for training, leading to increased costs. It is particularly challenging in medical imaging, i.e., gathering distributed data for centralized training, and meanwhile, obtaining quality labels remains a tedious job. Many methods have been proposed to address this issue in various training paradigms, e.g., continual learning, active learning, and federated learning, which indeed demonstrate certain forms of the data valuation process. However, existing methods are either overly intuitive or limited to common clean/toy datasets in the experiments. In this work, we present two data valuation metrics based on Synaptic Intelligence and gradient norms, respectively, to study the redundancy in real-world image data. Novel online and offline data selection algorithms are then proposed via clustering and grouping based on the examined data values. Our online approach effectively evaluates data utilizing layerwise model parameter updates and gradients in each epoch and can accelerate model training with fewer epochs and a subset (e.g., 19%-59%) of data while maintaining equivalent levels of accuracy in a variety of datasets. It also extends to the offline coreset construction, producing subsets of only 18%-30% of the original. The codes for the proposed adaptive data selection and coreset computation are available (https://github.com/ZhenyuTANG2023/data_selection).
MedFMC: A Real-world Dataset and Benchmark For Foundation Model Adaptation in Medical Image Classification
Jun 16, 2023Abstract:Foundation models, often pre-trained with large-scale data, have achieved paramount success in jump-starting various vision and language applications. Recent advances further enable adapting foundation models in downstream tasks efficiently using only a few training samples, e.g., in-context learning. Yet, the application of such learning paradigms in medical image analysis remains scarce due to the shortage of publicly accessible data and benchmarks. In this paper, we aim at approaches adapting the foundation models for medical image classification and present a novel dataset and benchmark for the evaluation, i.e., examining the overall performance of accommodating the large-scale foundation models downstream on a set of diverse real-world clinical tasks. We collect five sets of medical imaging data from multiple institutes targeting a variety of real-world clinical tasks (22,349 images in total), i.e., thoracic diseases screening in X-rays, pathological lesion tissue screening, lesion detection in endoscopy images, neonatal jaundice evaluation, and diabetic retinopathy grading. Results of multiple baseline methods are demonstrated using the proposed dataset from both accuracy and cost-effective perspectives.
T-AutoML: Automated Machine Learning for Lesion Segmentation using Transformers in 3D Medical Imaging
Nov 15, 2021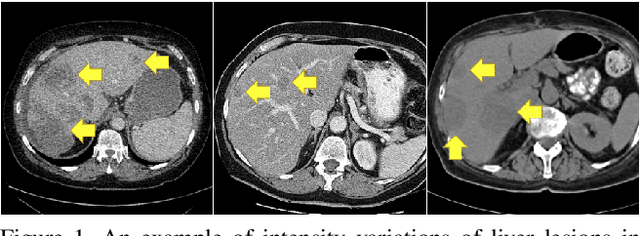
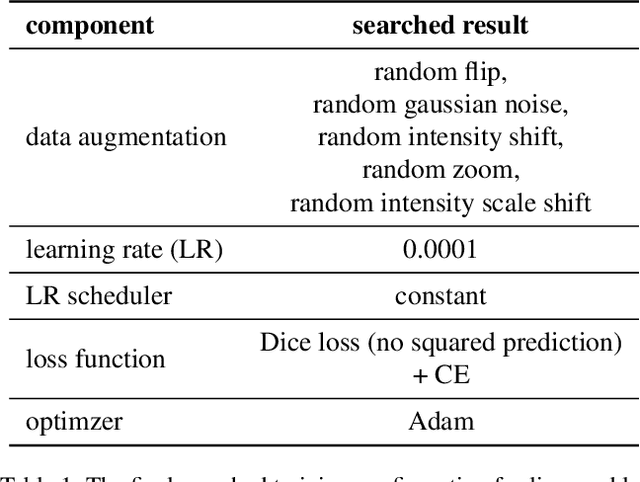
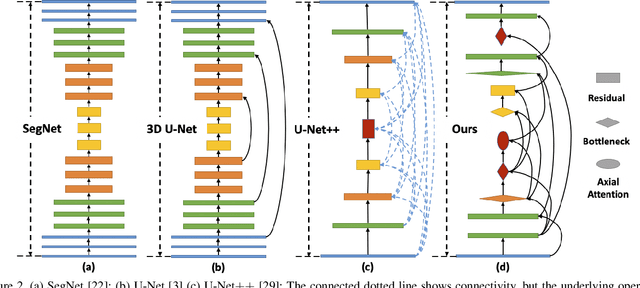
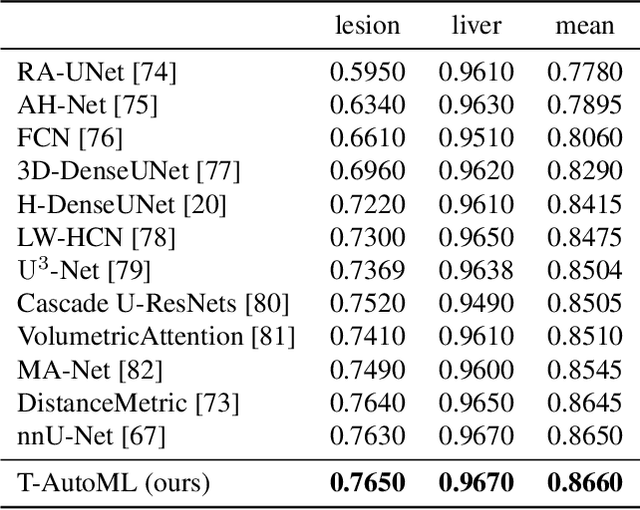
Abstract:Lesion segmentation in medical imaging has been an important topic in clinical research. Researchers have proposed various detection and segmentation algorithms to address this task. Recently, deep learning-based approaches have significantly improved the performance over conventional methods. However, most state-of-the-art deep learning methods require the manual design of multiple network components and training strategies. In this paper, we propose a new automated machine learning algorithm, T-AutoML, which not only searches for the best neural architecture, but also finds the best combination of hyper-parameters and data augmentation strategies simultaneously. The proposed method utilizes the modern transformer model, which is introduced to adapt to the dynamic length of the search space embedding and can significantly improve the ability of the search. We validate T-AutoML on several large-scale public lesion segmentation data-sets and achieve state-of-the-art performance.
Improving Pneumonia Localization via Cross-Attention on Medical Images and Reports
Oct 06, 2021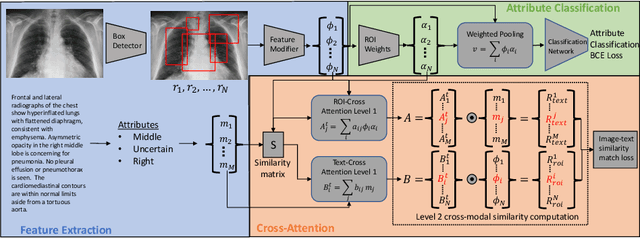
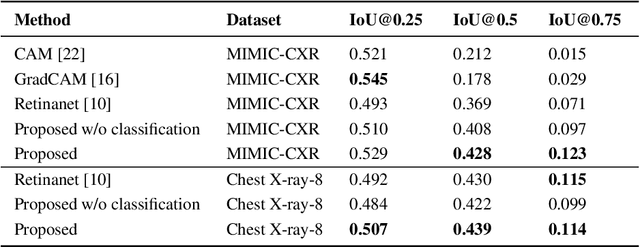
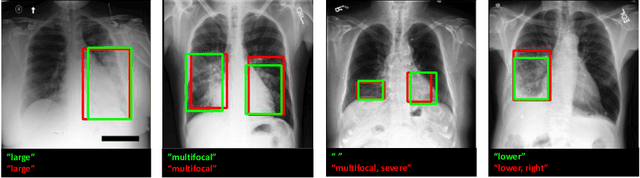
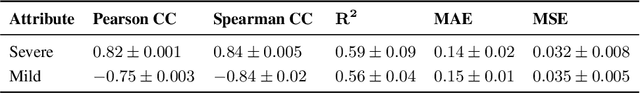
Abstract:Localization and characterization of diseases like pneumonia are primary steps in a clinical pipeline, facilitating detailed clinical diagnosis and subsequent treatment planning. Additionally, such location annotated datasets can provide a pathway for deep learning models to be used for downstream tasks. However, acquiring quality annotations is expensive on human resources and usually requires domain expertise. On the other hand, medical reports contain a plethora of information both about pneumonia characteristics and its location. In this paper, we propose a novel weakly-supervised attention-driven deep learning model that leverages encoded information in medical reports during training to facilitate better localization. Our model also performs classification of attributes that are associated to pneumonia and extracted from medical reports for supervision. Both the classification and localization are trained in conjunction and once trained, the model can be utilized for both the localization and characterization of pneumonia using only the input image. In this paper, we explore and analyze the model using chest X-ray datasets and demonstrate qualitatively and quantitatively that the introduction of textual information improves pneumonia localization. We showcase quantitative results on two datasets, MIMIC-CXR and Chest X-ray-8, and we also showcase severity characterization on the COVID-19 dataset.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge