Shuiwang Ji
FlowX: Towards Explainable Graph Neural Networks via Message Flows
Jun 26, 2022
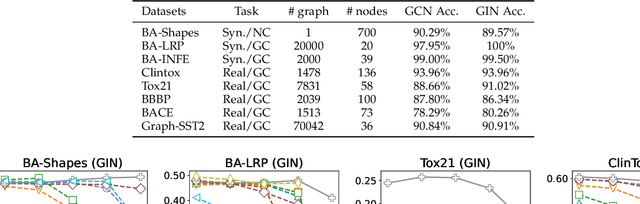
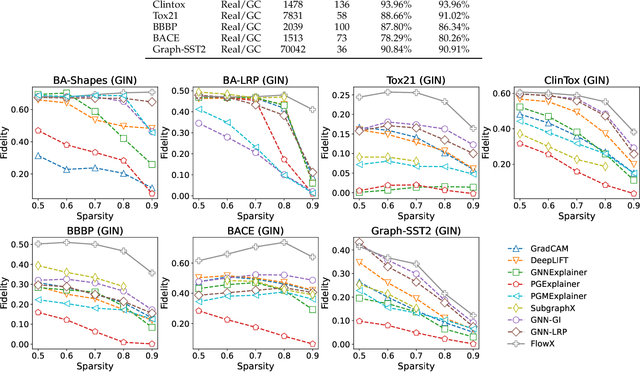
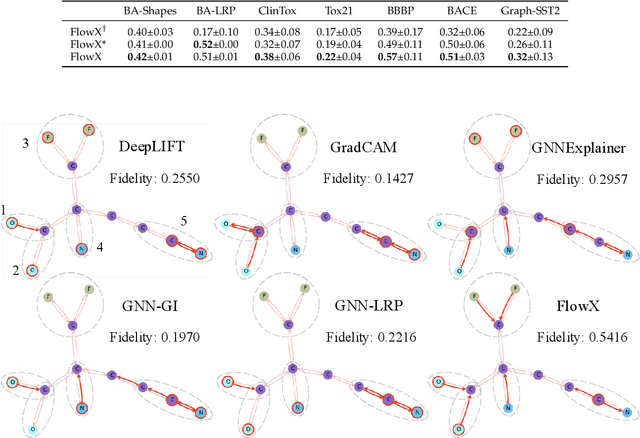
Abstract:We investigate the explainability of graph neural networks (GNNs) as a step towards elucidating their working mechanisms. While most current methods focus on explaining graph nodes, edges, or features, we argue that, as the inherent functional mechanism of GNNs, message flows are more natural for performing explainability. To this end, we propose a novel method here, known as FlowX, to explain GNNs by identifying important message flows. To quantify the importance of flows, we propose to follow the philosophy of Shapley values from cooperative game theory. To tackle the complexity of computing all coalitions' marginal contributions, we propose an approximation scheme to compute Shapley-like values as initial assessments of further redistribution training. We then propose a learning algorithm to train flow scores and improve explainability. Experimental studies on both synthetic and real-world datasets demonstrate that our proposed FlowX leads to improved explainability of GNNs.
GraphFM: Improving Large-Scale GNN Training via Feature Momentum
Jun 18, 2022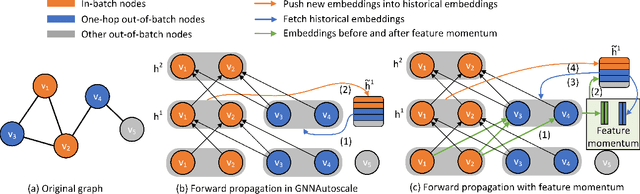
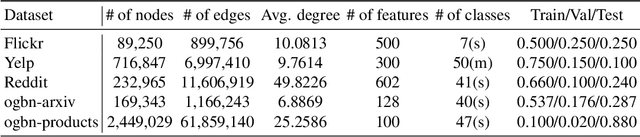
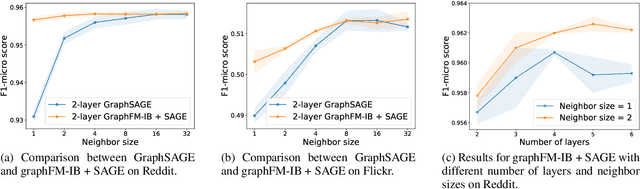
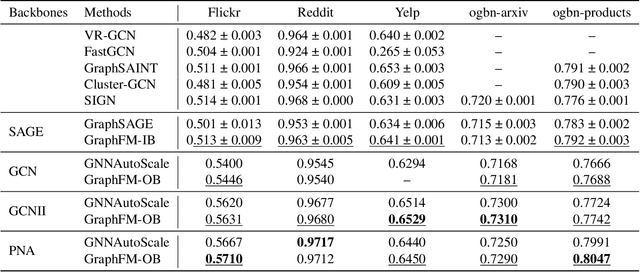
Abstract:Training of graph neural networks (GNNs) for large-scale node classification is challenging. A key difficulty lies in obtaining accurate hidden node representations while avoiding the neighborhood explosion problem. Here, we propose a new technique, named feature momentum (FM), that uses a momentum step to incorporate historical embeddings when updating feature representations. We develop two specific algorithms, known as GraphFM-IB and GraphFM-OB, that consider in-batch and out-of-batch data, respectively. GraphFM-IB applies FM to in-batch sampled data, while GraphFM-OB applies FM to out-of-batch data that are 1-hop neighborhood of in-batch data. We provide a convergence analysis for GraphFM-IB and some theoretical insight for GraphFM-OB. Empirically, we observe that GraphFM-IB can effectively alleviate the neighborhood explosion problem of existing methods. In addition, GraphFM-OB achieves promising performance on multiple large-scale graph datasets.
ComENet: Towards Complete and Efficient Message Passing for 3D Molecular Graphs
Jun 17, 2022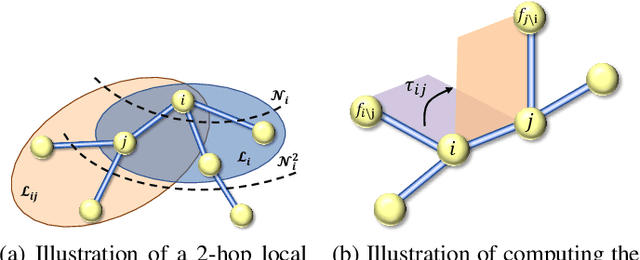

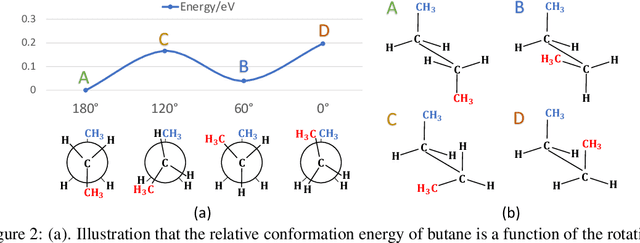
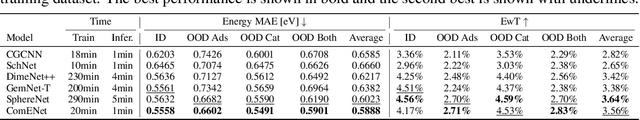
Abstract:Many real-world data can be modeled as 3D graphs, but learning representations that incorporates 3D information completely and efficiently is challenging. Existing methods either use partial 3D information, or suffer from excessive computational cost. To incorporate 3D information completely and efficiently, we propose a novel message passing scheme that operates within 1-hop neighborhood. Our method guarantees full completeness of 3D information on 3D graphs by achieving global and local completeness. Notably, we propose the important rotation angles to fulfill global completeness. Additionally, we show that our method is orders of magnitude faster than prior methods. We provide rigorous proof of completeness and analysis of time complexity for our methods. As molecules are in essence quantum systems, we build the \underline{com}plete and \underline{e}fficient graph neural network (ComENet) by combing quantum inspired basis functions and the proposed message passing scheme. Experimental results demonstrate the capability and efficiency of ComENet, especially on real-world datasets that are large in both numbers and sizes of graphs. Our code is publicly available as part of the DIG library (\url{https://github.com/divelab/DIG}).
GOOD: A Graph Out-of-Distribution Benchmark
Jun 16, 2022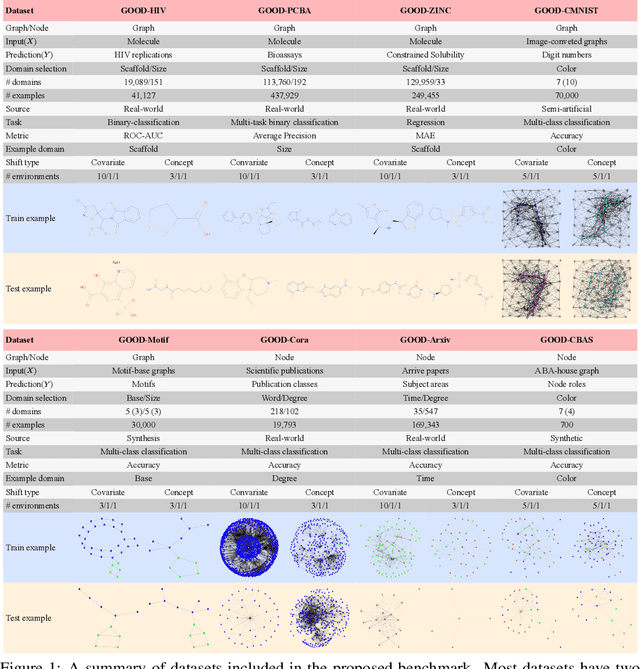
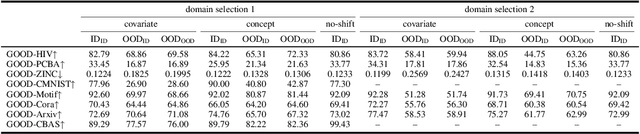
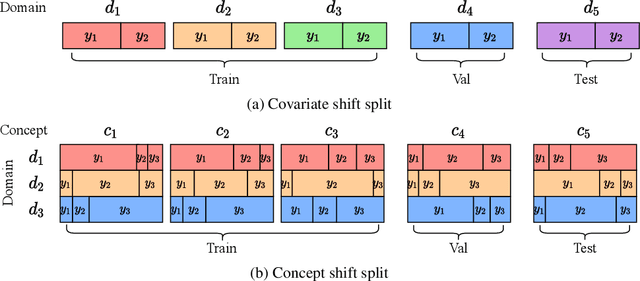
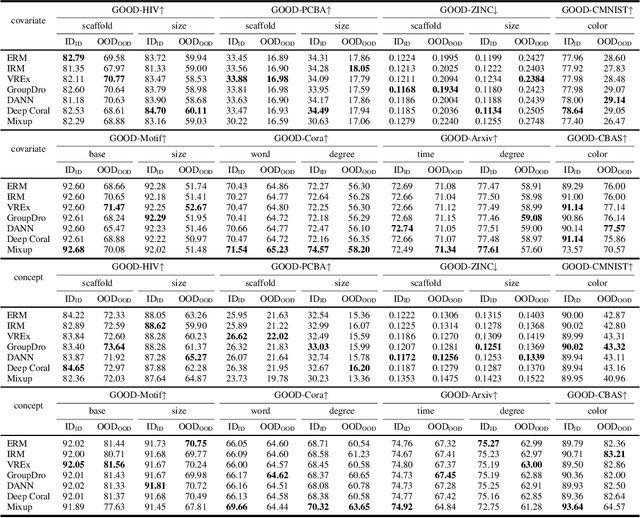
Abstract:Out-of-distribution (OOD) learning deals with scenarios in which training and test data follow different distributions. Although general OOD problems have been intensively studied in machine learning, graph OOD is only an emerging area of research. Currently, there lacks a systematic benchmark tailored to graph OOD method evaluation. In this work, we aim at developing an OOD benchmark, known as GOOD, for graphs specifically. We explicitly make distinctions between covariate and concept shifts and design data splits that accurately reflect different shifts. We consider both graph and node prediction tasks as there are key differences when designing shifts. Overall, GOOD contains 8 datasets with 14 domain selections. When combined with covariate, concept, and no shifts, we obtain 42 different splits. We provide performance results on 7 commonly used baseline methods with 10 random runs. This results in 294 dataset-model combinations in total. Our results show significant performance gaps between in-distribution and OOD settings. Our results also shed light on different performance trends between covariate and concept shifts by different methods. Our GOOD benchmark is a growing project and expects to expand in both quantity and variety of resources as the area develops. The GOOD benchmark can be accessed via $\href{https://github.com/divelab/GOOD/}{\text{https://github.com/divelab/GOOD/}}$.
Self-Adaptive Label Augmentation for Semi-supervised Few-shot Classification
Jun 16, 2022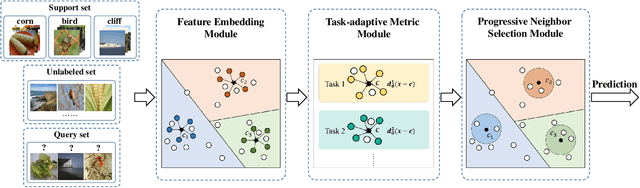
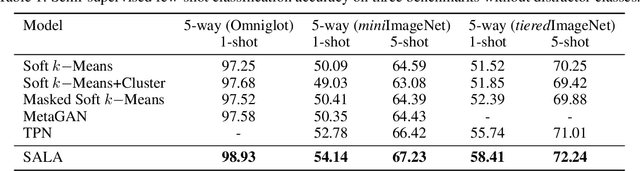
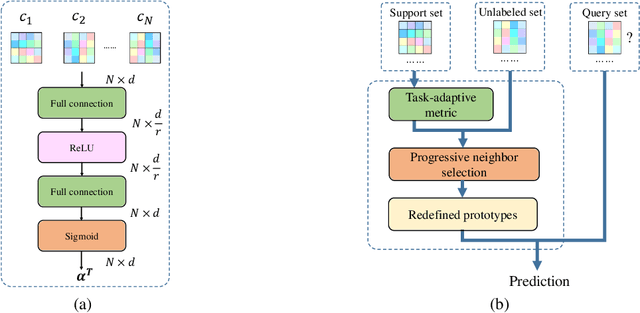
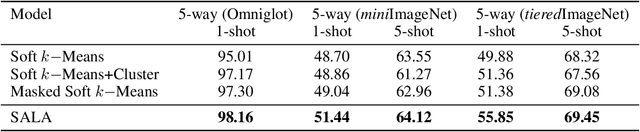
Abstract:Few-shot classification aims to learn a model that can generalize well to new tasks when only a few labeled samples are available. To make use of unlabeled data that are more abundantly available in real applications, Ren et al. \shortcite{ren2018meta} propose a semi-supervised few-shot classification method that assigns an appropriate label to each unlabeled sample by a manually defined metric. However, the manually defined metric fails to capture the intrinsic property in data. In this paper, we propose a \textbf{S}elf-\textbf{A}daptive \textbf{L}abel \textbf{A}ugmentation approach, called \textbf{SALA}, for semi-supervised few-shot classification. A major novelty of SALA is the task-adaptive metric, which can learn the metric adaptively for different tasks in an end-to-end fashion. Another appealing feature of SALA is a progressive neighbor selection strategy, which selects unlabeled data with high confidence progressively through the training phase. Experiments demonstrate that SALA outperforms several state-of-the-art methods for semi-supervised few-shot classification on benchmark datasets.
Lattice Convolutional Networks for Learning Ground States of Quantum Many-Body Systems
Jun 15, 2022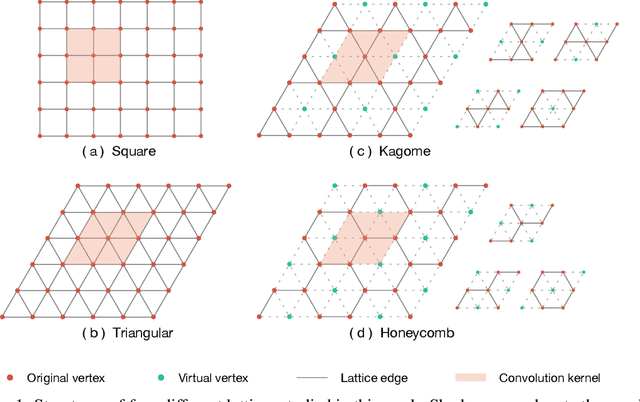
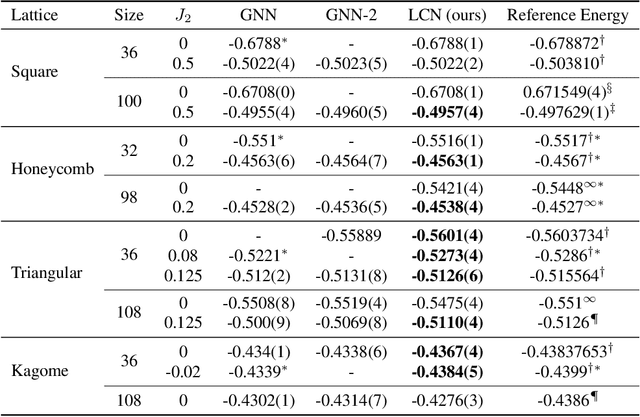
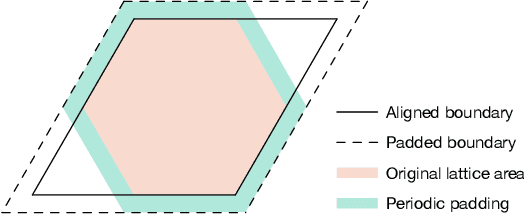
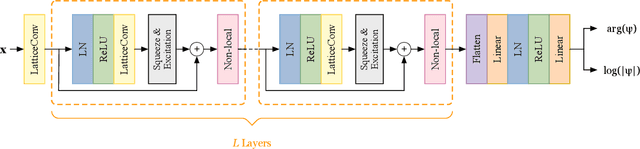
Abstract:Deep learning methods have been shown to be effective in representing ground-state wave functions of quantum many-body systems. Existing methods use convolutional neural networks (CNNs) for square lattices due to their image-like structures. For non-square lattices, existing method uses graph neural network (GNN) in which structure information is not precisely captured, thereby requiring additional hand-crafted sublattice encoding. In this work, we propose lattice convolutions in which a set of proposed operations are used to convert non-square lattices into grid-like augmented lattices on which regular convolution can be applied. Based on the proposed lattice convolutions, we design lattice convolutional networks (LCN) that use self-gating and attention mechanisms. Experimental results show that our method achieves performance on par or better than existing methods on spin 1/2 $J_1$-$J_2$ Heisenberg model over the square, honeycomb, triangular, and kagome lattices while without using hand-crafted encoding.
Your Neighbors Are Communicating: Towards Powerful and Scalable Graph Neural Networks
Jun 04, 2022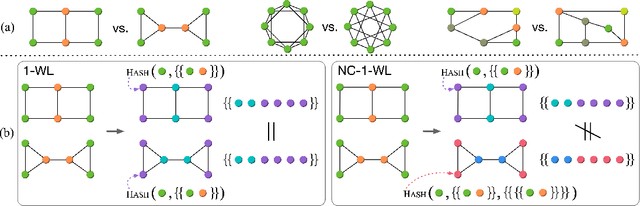
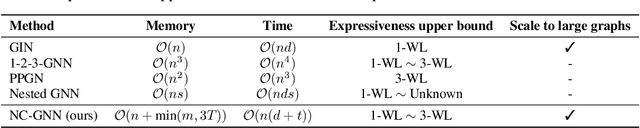

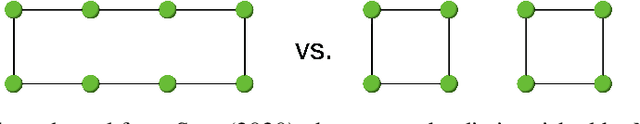
Abstract:Message passing graph neural networks (GNNs) are known to have their expressiveness upper-bounded by 1-dimensional Weisfeiler-Lehman (1-WL) algorithm. To achieve more powerful GNNs, existing attempts either require ad hoc features, or involve operations that incur high time and space complexities. In this work, we propose a general and provably powerful GNN framework that preserves the scalability of message passing scheme. In particular, we first propose to empower 1-WL for graph isomorphism test by considering edges among neighbors, giving rise to NC-1-WL. The expressiveness of NC-1-WL is shown to be strictly above 1-WL but below 3-WL theoretically. Further, we propose the NC-GNN framework as a differentiable neural version of NC-1-WL. Our simple implementation of NC-GNN is provably as powerful as NC-1-WL. Experiments demonstrate that our NC-GNN achieves remarkable performance on various benchmarks.
Learning Task-relevant Representations for Generalization via Characteristic Functions of Reward Sequence Distributions
May 20, 2022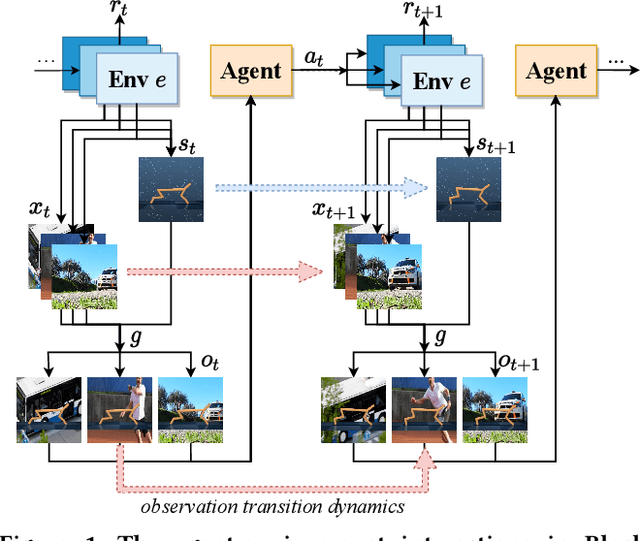
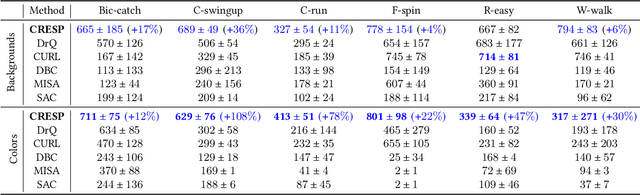
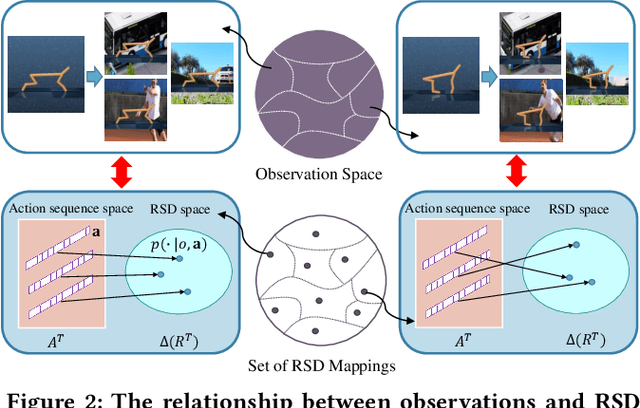
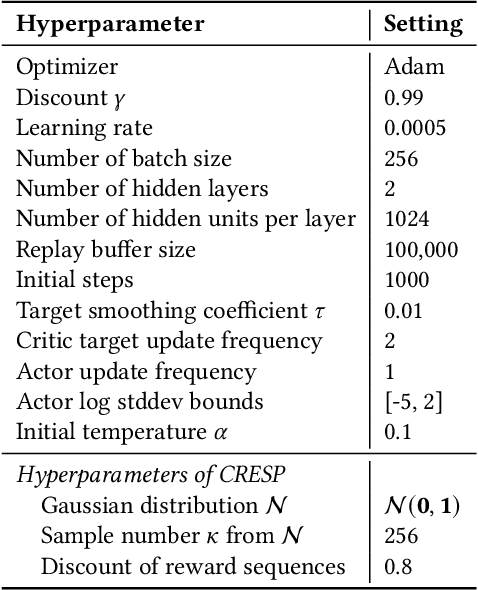
Abstract:Generalization across different environments with the same tasks is critical for successful applications of visual reinforcement learning (RL) in real scenarios. However, visual distractions -- which are common in real scenes -- from high-dimensional observations can be hurtful to the learned representations in visual RL, thus degrading the performance of generalization. To tackle this problem, we propose a novel approach, namely Characteristic Reward Sequence Prediction (CRESP), to extract the task-relevant information by learning reward sequence distributions (RSDs), as the reward signals are task-relevant in RL and invariant to visual distractions. Specifically, to effectively capture the task-relevant information via RSDs, CRESP introduces an auxiliary task -- that is, predicting the characteristic functions of RSDs -- to learn task-relevant representations, because we can well approximate the high-dimensional distributions by leveraging the corresponding characteristic functions. Experiments demonstrate that CRESP significantly improves the performance of generalization on unseen environments, outperforming several state-of-the-arts on DeepMind Control tasks with different visual distractions.
Generating 3D Molecules for Target Protein Binding
Apr 19, 2022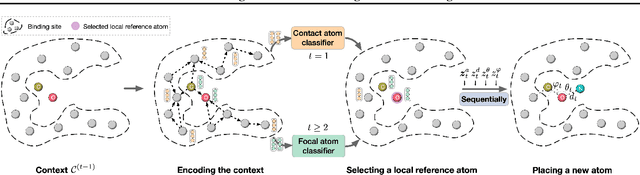
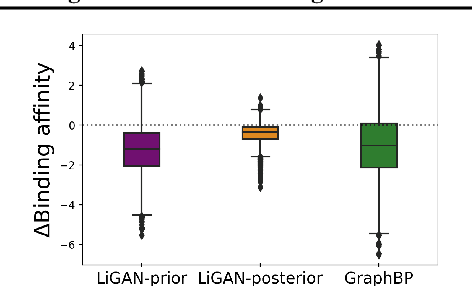
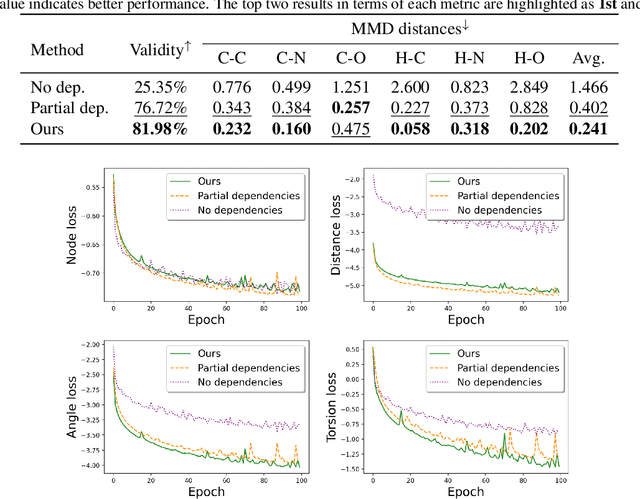
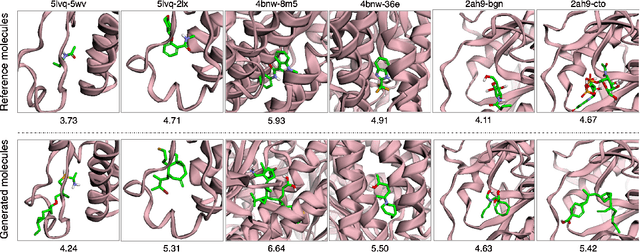
Abstract:A fundamental problem in drug discovery is to design molecules that bind to specific proteins. To tackle this problem using machine learning methods, here we propose a novel and effective framework, known as GraphBP, to generate 3D molecules that bind to given proteins by placing atoms of specific types and locations to the given binding site one by one. In particular, at each step, we first employ a 3D graph neural network to obtain geometry-aware and chemically informative representations from the intermediate contextual information. Such context includes the given binding site and atoms placed in the previous steps. Second, to preserve the desirable equivariance property, we select a local reference atom according to the designed auxiliary classifiers and then construct a local spherical coordinate system. Finally, to place a new atom, we generate its atom type and relative location w.r.t. the constructed local coordinate system via a flow model. We also consider generating the variables of interest sequentially to capture the underlying dependencies among them. Experiments demonstrate that our GraphBP is effective to generate 3D molecules with binding ability to target protein binding sites. Our implementation is available at https://github.com/divelab/GraphBP.
Duality-Induced Regularizer for Semantic Matching Knowledge Graph Embeddings
Apr 06, 2022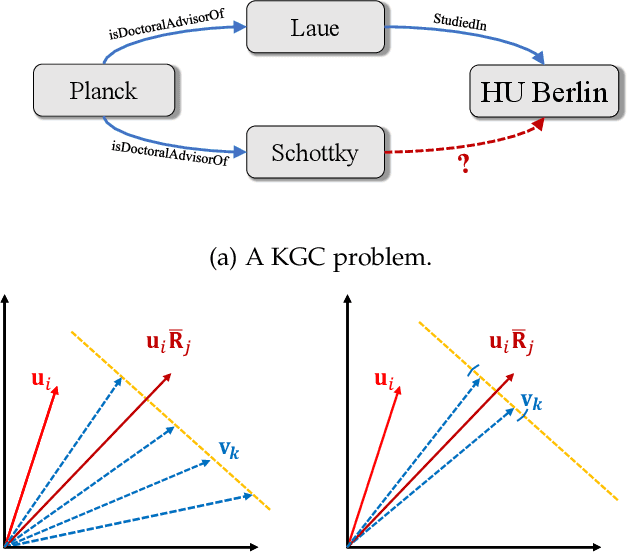
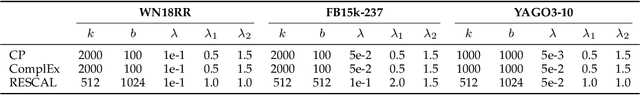
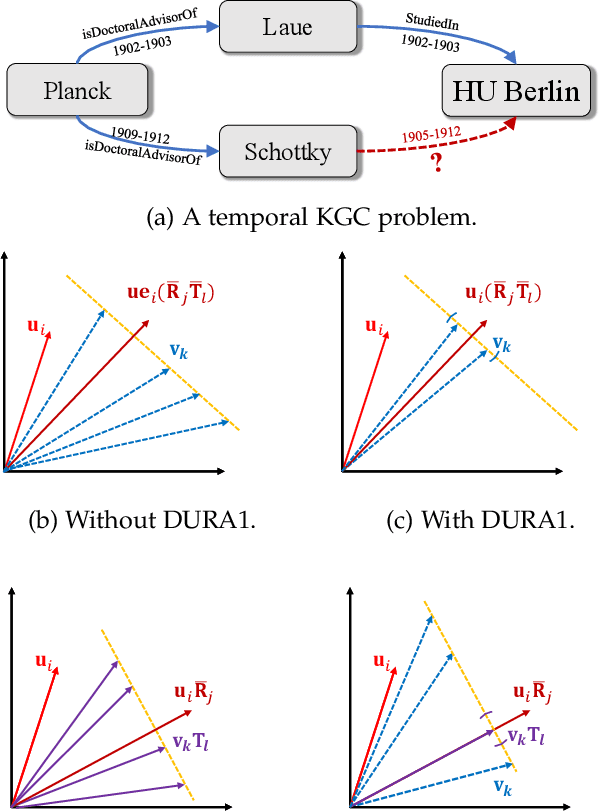
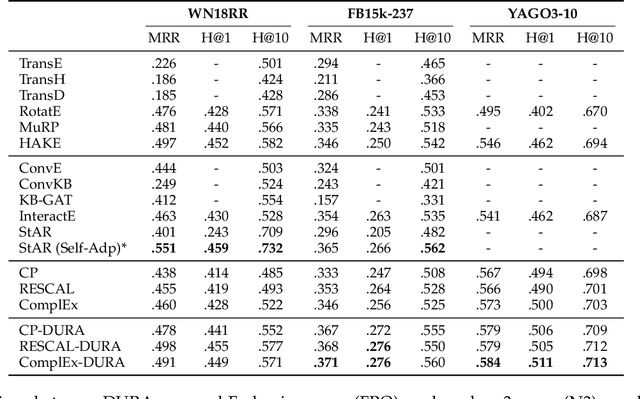
Abstract:Semantic matching models -- which assume that entities with similar semantics have similar embeddings -- have shown great power in knowledge graph embeddings (KGE). Many existing semantic matching models use inner products in embedding spaces to measure the plausibility of triples and quadruples in static and temporal knowledge graphs. However, vectors that have the same inner products with another vector can still be orthogonal to each other, which implies that entities with similar semantics may have dissimilar embeddings. This property of inner products significantly limits the performance of semantic matching models. To address this challenge, we propose a novel regularizer -- namely, DUality-induced RegulArizer (DURA) -- which effectively encourages the entities with similar semantics to have similar embeddings. The major novelty of DURA is based on the observation that, for an existing semantic matching KGE model (primal), there is often another distance based KGE model (dual) closely associated with it, which can be used as effective constraints for entity embeddings. Experiments demonstrate that DURA consistently and significantly improves the performance of state-of-the-art semantic matching models on both static and temporal knowledge graph benchmarks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge