Shuiwang Ji
Genetic InfoMax: Exploring Mutual Information Maximization in High-Dimensional Imaging Genetics Studies
Sep 26, 2023Abstract:Genome-wide association studies (GWAS) are used to identify relationships between genetic variations and specific traits. When applied to high-dimensional medical imaging data, a key step is to extract lower-dimensional, yet informative representations of the data as traits. Representation learning for imaging genetics is largely under-explored due to the unique challenges posed by GWAS in comparison to typical visual representation learning. In this study, we tackle this problem from the mutual information (MI) perspective by identifying key limitations of existing methods. We introduce a trans-modal learning framework Genetic InfoMax (GIM), including a regularized MI estimator and a novel genetics-informed transformer to address the specific challenges of GWAS. We evaluate GIM on human brain 3D MRI data and establish standardized evaluation protocols to compare it to existing approaches. Our results demonstrate the effectiveness of GIM and a significantly improved performance on GWAS.
Video Timeline Modeling For News Story Understanding
Sep 23, 2023



Abstract:In this paper, we present a novel problem, namely video timeline modeling. Our objective is to create a video-associated timeline from a set of videos related to a specific topic, thereby facilitating the content and structure understanding of the story being told. This problem has significant potential in various real-world applications, such as news story summarization. To bootstrap research in this area, we curate a realistic benchmark dataset, YouTube-News-Timeline, consisting of over $12$k timelines and $300$k YouTube news videos. Additionally, we propose a set of quantitative metrics as the protocol to comprehensively evaluate and compare methodologies. With such a testbed, we further develop and benchmark exploratory deep learning approaches to tackle this problem. We anticipate that this exploratory work will pave the way for further research in video timeline modeling. The assets are available via https://github.com/google-research/google-research/tree/master/video_timeline_modeling.
Artificial Intelligence for Science in Quantum, Atomistic, and Continuum Systems
Jul 17, 2023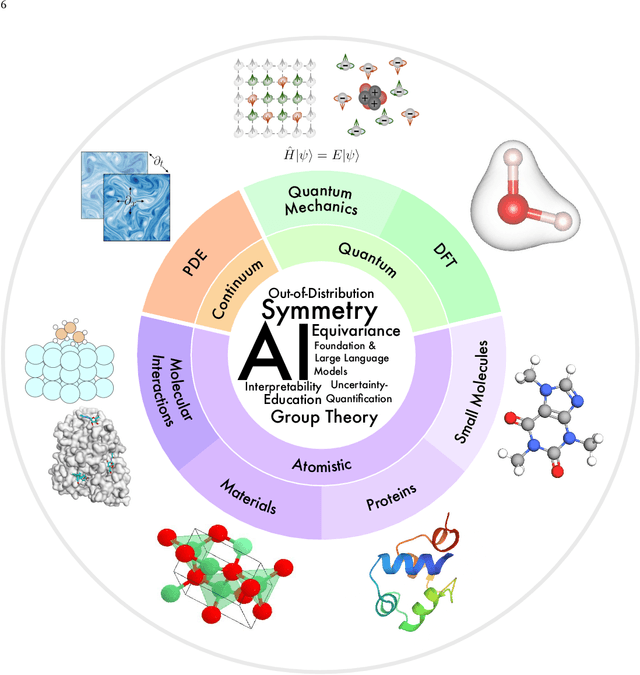



Abstract:Advances in artificial intelligence (AI) are fueling a new paradigm of discoveries in natural sciences. Today, AI has started to advance natural sciences by improving, accelerating, and enabling our understanding of natural phenomena at a wide range of spatial and temporal scales, giving rise to a new area of research known as AI for science (AI4Science). Being an emerging research paradigm, AI4Science is unique in that it is an enormous and highly interdisciplinary area. Thus, a unified and technical treatment of this field is needed yet challenging. This paper aims to provide a technically thorough account of a subarea of AI4Science; namely, AI for quantum, atomistic, and continuum systems. These areas aim at understanding the physical world from the subatomic (wavefunctions and electron density), atomic (molecules, proteins, materials, and interactions), to macro (fluids, climate, and subsurface) scales and form an important subarea of AI4Science. A unique advantage of focusing on these areas is that they largely share a common set of challenges, thereby allowing a unified and foundational treatment. A key common challenge is how to capture physics first principles, especially symmetries, in natural systems by deep learning methods. We provide an in-depth yet intuitive account of techniques to achieve equivariance to symmetry transformations. We also discuss other common technical challenges, including explainability, out-of-distribution generalization, knowledge transfer with foundation and large language models, and uncertainty quantification. To facilitate learning and education, we provide categorized lists of resources that we found to be useful. We strive to be thorough and unified and hope this initial effort may trigger more community interests and efforts to further advance AI4Science.
Towards Symmetry-Aware Generation of Periodic Materials
Jul 06, 2023



Abstract:We consider the problem of generating periodic materials with deep models. While symmetry-aware molecule generation has been studied extensively, periodic materials possess different symmetries, which have not been completely captured by existing methods. In this work, we propose SyMat, a novel material generation approach that can capture physical symmetries of periodic material structures. SyMat generates atom types and lattices of materials through generating atom type sets, lattice lengths and lattice angles with a variational auto-encoder model. In addition, SyMat employs a score-based diffusion model to generate atom coordinates of materials, in which a novel symmetry-aware probabilistic model is used in the coordinate diffusion process. We show that SyMat is theoretically invariant to all symmetry transformations on materials and demonstrate that SyMat achieves promising performance on random generation and property optimization tasks.
Efficient Approximations of Complete Interatomic Potentials for Crystal Property Prediction
Jun 26, 2023



Abstract:We study property prediction for crystal materials. A crystal structure consists of a minimal unit cell that is repeated infinitely in 3D space. How to accurately represent such repetitive structures in machine learning models remains unresolved. Current methods construct graphs by establishing edges only between nearby nodes, thereby failing to faithfully capture infinite repeating patterns and distant interatomic interactions. In this work, we propose several innovations to overcome these limitations. First, we propose to model physics-principled interatomic potentials directly instead of only using distances as in many existing methods. These potentials include the Coulomb potential, London dispersion potential, and Pauli repulsion potential. Second, we model the complete set of potentials among all atoms, instead of only between nearby atoms as in existing methods. This is enabled by our approximations of infinite potential summations with provable error bounds. We further develop efficient algorithms to compute the approximations. Finally, we propose to incorporate our computations of complete interatomic potentials into message passing neural networks for representation learning. We perform experiments on the JARVIS and Materials Project benchmarks for evaluation. Results show that the use of interatomic potentials and complete interatomic potentials leads to consistent performance improvements with reasonable computational costs. Our code is publicly available as part of the AIRS library (https://github.com/divelab/AIRS/tree/main/OpenMat/PotNet).
QH9: A Quantum Hamiltonian Prediction Benchmark for QM9 Molecules
Jun 15, 2023
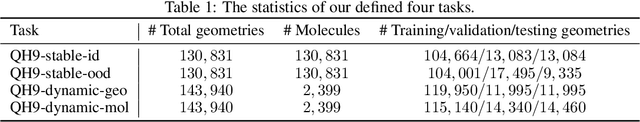


Abstract:Supervised machine learning approaches have been increasingly used in accelerating electronic structure prediction as surrogates of first-principle computational methods, such as density functional theory (DFT). While numerous quantum chemistry datasets focus on chemical properties and atomic forces, the ability to achieve accurate and efficient prediction of the Hamiltonian matrix is highly desired, as it is the most important and fundamental physical quantity that determines the quantum states of physical systems and chemical properties. In this work, we generate a new Quantum Hamiltonian dataset, named as QH9, to provide precise Hamiltonian matrices for 2,399 molecular dynamics trajectories and 130,831 stable molecular geometries, based on the QM9 dataset. By designing benchmark tasks with various molecules, we show that current machine learning models have the capacity to predict Hamiltonian matrices for arbitrary molecules. Both the QH9 dataset and the baseline models are provided to the community through an open-source benchmark, which can be highly valuable for developing machine learning methods and accelerating molecular and materials design for scientific and technological applications. Our benchmark is publicly available at https://github.com/divelab/AIRS/tree/main/OpenDFT/QHBench.
Graph Structure and Feature Extrapolation for Out-of-Distribution Generalization
Jun 13, 2023



Abstract:Out-of-distribution (OOD) generalization deals with the prevalent learning scenario where test distribution shifts from training distribution. With rising application demands and inherent complexity, graph OOD problems call for specialized solutions. While data-centric methods exhibit performance enhancements on many generic machine learning tasks, there is a notable absence of data augmentation methods tailored for graph OOD generalization. In this work, we propose to achieve graph OOD generalization with the novel design of non-Euclidean-space linear extrapolation. The proposed augmentation strategy extrapolates both structure and feature spaces to generate OOD graph data. Our design tailors OOD samples for specific shifts without corrupting underlying causal mechanisms. Theoretical analysis and empirical results evidence the effectiveness of our method in solving target shifts, showing substantial and constant improvements across various graph OOD tasks.
Graph Mixup with Soft Alignments
Jun 11, 2023



Abstract:We study graph data augmentation by mixup, which has been used successfully on images. A key operation of mixup is to compute a convex combination of a pair of inputs. This operation is straightforward for grid-like data, such as images, but challenging for graph data. The key difficulty lies in the fact that different graphs typically have different numbers of nodes, and thus there lacks a node-level correspondence between graphs. In this work, we propose S-Mixup, a simple yet effective mixup method for graph classification by soft alignments. Specifically, given a pair of graphs, we explicitly obtain node-level correspondence via computing a soft assignment matrix to match the nodes between two graphs. Based on the soft assignments, we transform the adjacency and node feature matrices of one graph, so that the transformed graph is aligned with the other graph. In this way, any pair of graphs can be mixed directly to generate an augmented graph. We conduct systematic experiments to show that S-Mixup can improve the performance and generalization of graph neural networks (GNNs) on various graph classification tasks. In addition, we show that S-Mixup can increase the robustness of GNNs against noisy labels.
Group Equivariant Fourier Neural Operators for Partial Differential Equations
Jun 09, 2023



Abstract:We consider solving partial differential equations (PDEs) with Fourier neural operators (FNOs), which operate in the frequency domain. Since the laws of physics do not depend on the coordinate system used to describe them, it is desirable to encode such symmetries in the neural operator architecture for better performance and easier learning. While encoding symmetries in the physical domain using group theory has been studied extensively, how to capture symmetries in the frequency domain is under-explored. In this work, we extend group convolutions to the frequency domain and design Fourier layers that are equivariant to rotations, translations, and reflections by leveraging the equivariance property of the Fourier transform. The resulting $G$-FNO architecture generalizes well across input resolutions and performs well in settings with varying levels of symmetry. Our code is publicly available as part of the AIRS library (https://github.com/divelab/AIRS).
Joint Learning of Label and Environment Causal Independence for Graph Out-of-Distribution Generalization
Jun 08, 2023Abstract:We tackle the problem of graph out-of-distribution (OOD) generalization. Existing graph OOD algorithms either rely on restricted assumptions or fail to exploit environment information in training data. In this work, we propose to simultaneously incorporate label and environment causal independence (LECI) to fully make use of label and environment information, thereby addressing the challenges faced by prior methods on identifying causal and invariant subgraphs. We further develop an adversarial training strategy to jointly optimize these two properties for causal subgraph discovery with theoretical guarantees. Extensive experiments and analysis show that LECI significantly outperforms prior methods on both synthetic and real-world datasets, establishing LECI as a practical and effective solution for graph OOD generalization.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge