Nassir Navab
Computer Aided Medical Procedures, Technische Universit Munchen, Germany, Johns Hopkins University, Baltimore MD, USA
From EHRs to Patient Pathways: Scalable Modeling of Longitudinal Health Trajectories with LLMs
Jun 05, 2025



Abstract:Healthcare systems face significant challenges in managing and interpreting vast, heterogeneous patient data for personalized care. Existing approaches often focus on narrow use cases with a limited feature space, overlooking the complex, longitudinal interactions needed for a holistic understanding of patient health. In this work, we propose a novel approach to patient pathway modeling by transforming diverse electronic health record (EHR) data into a structured representation and designing a holistic pathway prediction model, EHR2Path, optimized to predict future health trajectories. Further, we introduce a novel summary mechanism that embeds long-term temporal context into topic-specific summary tokens, improving performance over text-only models, while being much more token-efficient. EHR2Path demonstrates strong performance in both next time-step prediction and longitudinal simulation, outperforming competitive baselines. It enables detailed simulations of patient trajectories, inherently targeting diverse evaluation tasks, such as forecasting vital signs, lab test results, or length-of-stay, opening a path towards predictive and personalized healthcare.
EgoExOR: An Ego-Exo-Centric Operating Room Dataset for Surgical Activity Understanding
May 30, 2025Abstract:Operating rooms (ORs) demand precise coordination among surgeons, nurses, and equipment in a fast-paced, occlusion-heavy environment, necessitating advanced perception models to enhance safety and efficiency. Existing datasets either provide partial egocentric views or sparse exocentric multi-view context, but do not explore the comprehensive combination of both. We introduce EgoExOR, the first OR dataset and accompanying benchmark to fuse first-person and third-person perspectives. Spanning 94 minutes (84,553 frames at 15 FPS) of two emulated spine procedures, Ultrasound-Guided Needle Insertion and Minimally Invasive Spine Surgery, EgoExOR integrates egocentric data (RGB, gaze, hand tracking, audio) from wearable glasses, exocentric RGB and depth from RGB-D cameras, and ultrasound imagery. Its detailed scene graph annotations, covering 36 entities and 22 relations (568,235 triplets), enable robust modeling of clinical interactions, supporting tasks like action recognition and human-centric perception. We evaluate the surgical scene graph generation performance of two adapted state-of-the-art models and offer a new baseline that explicitly leverages EgoExOR's multimodal and multi-perspective signals. This new dataset and benchmark set a new foundation for OR perception, offering a rich, multimodal resource for next-generation clinical perception.
Does Machine Unlearning Truly Remove Model Knowledge? A Framework for Auditing Unlearning in LLMs
May 29, 2025



Abstract:In recent years, Large Language Models (LLMs) have achieved remarkable advancements, drawing significant attention from the research community. Their capabilities are largely attributed to large-scale architectures, which require extensive training on massive datasets. However, such datasets often contain sensitive or copyrighted content sourced from the public internet, raising concerns about data privacy and ownership. Regulatory frameworks, such as the General Data Protection Regulation (GDPR), grant individuals the right to request the removal of such sensitive information. This has motivated the development of machine unlearning algorithms that aim to remove specific knowledge from models without the need for costly retraining. Despite these advancements, evaluating the efficacy of unlearning algorithms remains a challenge due to the inherent complexity and generative nature of LLMs. In this work, we introduce a comprehensive auditing framework for unlearning evaluation, comprising three benchmark datasets, six unlearning algorithms, and five prompt-based auditing methods. By using various auditing algorithms, we evaluate the effectiveness and robustness of different unlearning strategies. To explore alternatives beyond prompt-based auditing, we propose a novel technique that leverages intermediate activation perturbations, addressing the limitations of auditing methods that rely solely on model inputs and outputs.
Temporal Differential Fields for 4D Motion Modeling via Image-to-Video Synthesis
May 22, 2025Abstract:Temporal modeling on regular respiration-induced motions is crucial to image-guided clinical applications. Existing methods cannot simulate temporal motions unless high-dose imaging scans including starting and ending frames exist simultaneously. However, in the preoperative data acquisition stage, the slight movement of patients may result in dynamic backgrounds between the first and last frames in a respiratory period. This additional deviation can hardly be removed by image registration, thus affecting the temporal modeling. To address that limitation, we pioneeringly simulate the regular motion process via the image-to-video (I2V) synthesis framework, which animates with the first frame to forecast future frames of a given length. Besides, to promote the temporal consistency of animated videos, we devise the Temporal Differential Diffusion Model to generate temporal differential fields, which measure the relative differential representations between adjacent frames. The prompt attention layer is devised for fine-grained differential fields, and the field augmented layer is adopted to better interact these fields with the I2V framework, promoting more accurate temporal variation of synthesized videos. Extensive results on ACDC cardiac and 4D Lung datasets reveal that our approach simulates 4D videos along the intrinsic motion trajectory, rivaling other competitive methods on perceptual similarity and temporal consistency. Codes will be available soon.
ORQA: A Benchmark and Foundation Model for Holistic Operating Room Modeling
May 19, 2025Abstract:The real-world complexity of surgeries necessitates surgeons to have deep and holistic comprehension to ensure precision, safety, and effective interventions. Computational systems are required to have a similar level of comprehension within the operating room. Prior works, limited to single-task efforts like phase recognition or scene graph generation, lack scope and generalizability. In this work, we introduce ORQA, a novel OR question answering benchmark and foundational multimodal model to advance OR intelligence. By unifying all four public OR datasets into a comprehensive benchmark, we enable our approach to concurrently address a diverse range of OR challenges. The proposed multimodal large language model fuses diverse OR signals such as visual, auditory, and structured data, for a holistic modeling of the OR. Finally, we propose a novel, progressive knowledge distillation paradigm, to generate a family of models optimized for different speed and memory requirements. We show the strong performance of ORQA on our proposed benchmark, and its zero-shot generalization, paving the way for scalable, unified OR modeling and significantly advancing multimodal surgical intelligence. We will release our code and data upon acceptance.
VeLU: Variance-enhanced Learning Unit for Deep Neural Networks
Apr 21, 2025



Abstract:Activation functions are fundamental in deep neural networks and directly impact gradient flow, optimization stability, and generalization. Although ReLU remains standard because of its simplicity, it suffers from vanishing gradients and lacks adaptability. Alternatives like Swish and GELU introduce smooth transitions, but fail to dynamically adjust to input statistics. We propose VeLU, a Variance-enhanced Learning Unit as an activation function that dynamically scales based on input variance by integrating ArcTan-Sin transformations and Wasserstein-2 regularization, effectively mitigating covariate shifts and stabilizing optimization. Extensive experiments on ViT_B16, VGG19, ResNet50, DenseNet121, MobileNetV2, and EfficientNetB3 confirm VeLU's superiority over ReLU, ReLU6, Swish, and GELU on six vision benchmarks. The codes of VeLU are publicly available on GitHub.
LiteTracker: Leveraging Temporal Causality for Accurate Low-latency Tissue Tracking
Apr 14, 2025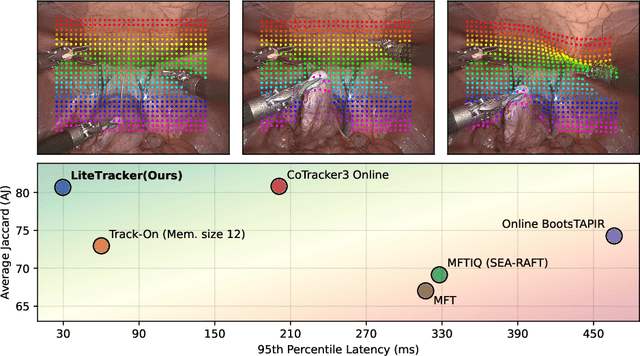
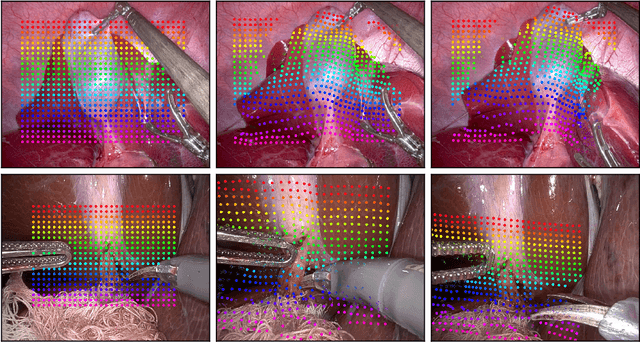
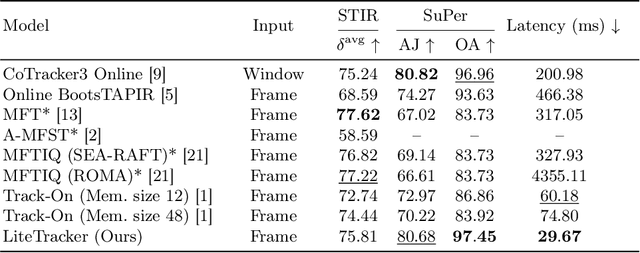
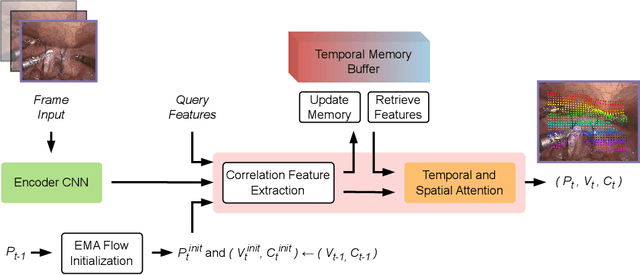

Abstract:Tissue tracking plays a critical role in various surgical navigation and extended reality (XR) applications. While current methods trained on large synthetic datasets achieve high tracking accuracy and generalize well to endoscopic scenes, their runtime performances fail to meet the low-latency requirements necessary for real-time surgical applications. To address this limitation, we propose LiteTracker, a low-latency method for tissue tracking in endoscopic video streams. LiteTracker builds on a state-of-the-art long-term point tracking method, and introduces a set of training-free runtime optimizations. These optimizations enable online, frame-by-frame tracking by leveraging a temporal memory buffer for efficient feature reuse and utilizing prior motion for accurate track initialization. LiteTracker demonstrates significant runtime improvements being around 7x faster than its predecessor and 2x than the state-of-the-art. Beyond its primary focus on efficiency, LiteTracker delivers high-accuracy tracking and occlusion prediction, performing competitively on both the STIR and SuPer datasets. We believe LiteTracker is an important step toward low-latency tissue tracking for real-time surgical applications in the operating room.
PRISM-0: A Predicate-Rich Scene Graph Generation Framework for Zero-Shot Open-Vocabulary Tasks
Apr 01, 2025Abstract:In Scene Graphs Generation (SGG) one extracts structured representation from visual inputs in the form of objects nodes and predicates connecting them. This facilitates image-based understanding and reasoning for various downstream tasks. Although fully supervised SGG approaches showed steady performance improvements, they suffer from a severe training bias. This is caused by the availability of only small subsets of curated data and exhibits long-tail predicate distribution issues with a lack of predicate diversity adversely affecting downstream tasks. To overcome this, we introduce PRISM-0, a framework for zero-shot open-vocabulary SGG that bootstraps foundation models in a bottom-up approach to capture the whole spectrum of diverse, open-vocabulary predicate prediction. Detected object pairs are filtered and passed to a Vision Language Model (VLM) that generates descriptive captions. These are used to prompt an LLM to generate fine-andcoarse-grained predicates for the pair. The predicates are then validated using a VQA model to provide a final SGG. With the modular and dataset-independent PRISM-0, we can enrich existing SG datasets such as Visual Genome (VG). Experiments illustrate that PRIMS-0 generates semantically meaningful graphs that improve downstream tasks such as Image Captioning and Sentence-to-Graph Retrieval with a performance on par to the best fully supervised methods.
Adversarial Wear and Tear: Exploiting Natural Damage for Generating Physical-World Adversarial Examples
Mar 27, 2025



Abstract:The presence of adversarial examples in the physical world poses significant challenges to the deployment of Deep Neural Networks in safety-critical applications such as autonomous driving. Most existing methods for crafting physical-world adversarial examples are ad-hoc, relying on temporary modifications like shadows, laser beams, or stickers that are tailored to specific scenarios. In this paper, we introduce a new class of physical-world adversarial examples, AdvWT, which draws inspiration from the naturally occurring phenomenon of `wear and tear', an inherent property of physical objects. Unlike manually crafted perturbations, `wear and tear' emerges organically over time due to environmental degradation, as seen in the gradual deterioration of outdoor signboards. To achieve this, AdvWT follows a two-step approach. First, a GAN-based, unsupervised image-to-image translation network is employed to model these naturally occurring damages, particularly in the context of outdoor signboards. The translation network encodes the characteristics of damaged signs into a latent `damage style code'. In the second step, we introduce adversarial perturbations into the style code, strategically optimizing its transformation process. This manipulation subtly alters the damage style representation, guiding the network to generate adversarial images where the appearance of damages remains perceptually realistic, while simultaneously ensuring their effectiveness in misleading neural networks. Through comprehensive experiments on two traffic sign datasets, we show that AdvWT effectively misleads DNNs in both digital and physical domains. AdvWT achieves an effective attack success rate, greater robustness, and a more natural appearance compared to existing physical-world adversarial examples. Additionally, integrating AdvWT into training enhances a model's generalizability to real-world damaged signs.
Learning to Efficiently Adapt Foundation Models for Self-Supervised Endoscopic 3D Scene Reconstruction from Any Cameras
Mar 20, 2025



Abstract:Accurate 3D scene reconstruction is essential for numerous medical tasks. Given the challenges in obtaining ground truth data, there has been an increasing focus on self-supervised learning (SSL) for endoscopic depth estimation as a basis for scene reconstruction. While foundation models have shown remarkable progress in visual tasks, their direct application to the medical domain often leads to suboptimal results. However, the visual features from these models can still enhance endoscopic tasks, emphasizing the need for efficient adaptation strategies, which still lack exploration currently. In this paper, we introduce Endo3DAC, a unified framework for endoscopic scene reconstruction that efficiently adapts foundation models. We design an integrated network capable of simultaneously estimating depth maps, relative poses, and camera intrinsic parameters. By freezing the backbone foundation model and training only the specially designed Gated Dynamic Vector-Based Low-Rank Adaptation (GDV-LoRA) with separate decoder heads, Endo3DAC achieves superior depth and pose estimation while maintaining training efficiency. Additionally, we propose a 3D scene reconstruction pipeline that optimizes depth maps' scales, shifts, and a few parameters based on our integrated network. Extensive experiments across four endoscopic datasets demonstrate that Endo3DAC significantly outperforms other state-of-the-art methods while requiring fewer trainable parameters. To our knowledge, we are the first to utilize a single network that only requires surgical videos to perform both SSL depth estimation and scene reconstruction tasks. The code will be released upon acceptance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge