Reflective-AR Display: An Interaction Methodology for Virtual-Real Alignment in Medical Robotics
Jul 23, 2019Javad Fotouhi, Tianyu Song, Arian Mehrfard, Giacomo Taylor, Alejandro Martin-Gomez, Bernhard Fuerst, Mehran Armand, Mathias Unberath, Nassir Navab
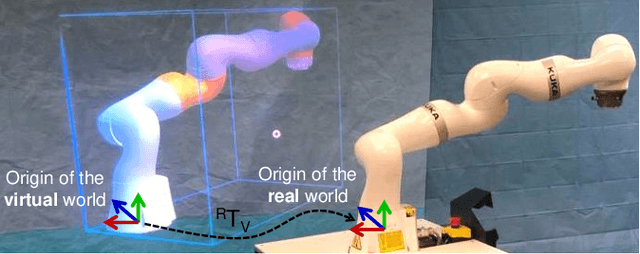
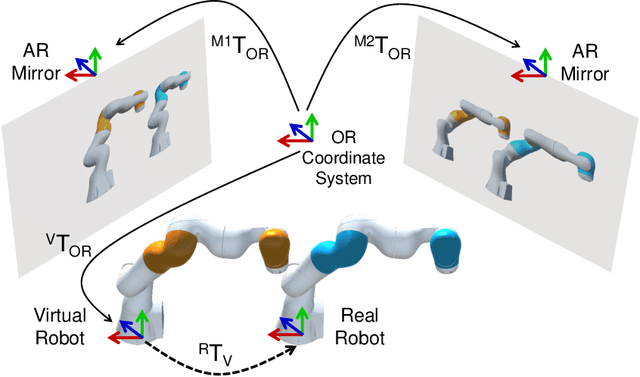
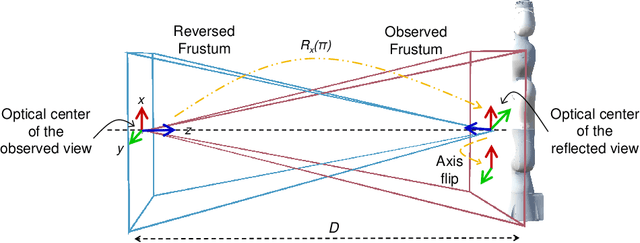
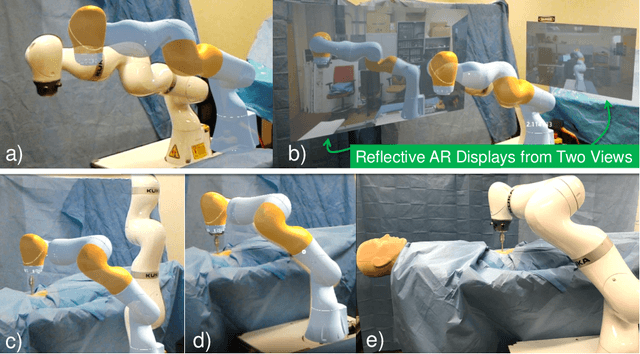
Robot-assisted minimally invasive surgery has shown to improve patient outcomes, as well as reduce complications and recovery time for several clinical applications. However, increasingly configurable robotic arms require careful setup by surgical staff to maximize anatomical reach and avoid collisions. Furthermore, safety regulations prevent automatically driving robotic arms to this optimal positioning. We propose a Head-Mounted Display (HMD) based augmented reality (AR) guidance system for optimal surgical arm setup. In this case, the staff equipped with HMD aligns the robot with its planned virtual counterpart. The main challenge, however, is the perspective ambiguities hindering such collaborative robotic solution. To overcome this challenge, we introduce a novel registration concept for intuitive alignment of such AR content by providing a multi-view AR experience via reflective-AR displays that show the augmentations from multiple viewpoints. Using this system, operators can visualize different perspectives simultaneously while actively adjusting the pose to determine the registration transformation that most closely superimposes the virtual onto real. The experimental results demonstrate improvement in the interactive alignment of a virtual and real robot when using a reflective-AR display. We also present measurements from configuring a robotic manipulator in a simulated trocar placement surgery using the AR guidance methodology.
Domain-Specific Priors and Meta Learning for Low-shot First-Person Action Recognition
Jul 22, 2019Huseyin Coskun, Zeeshan Zia, Bugra Tekin, Federica Bogo, Nassir Navab, Federico Tombari, Harpreet Sawhney
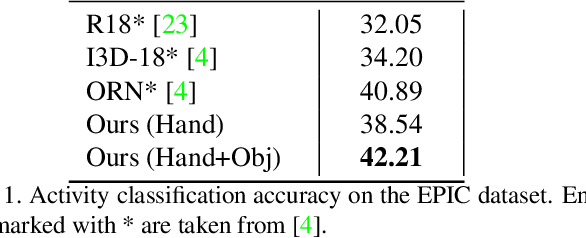
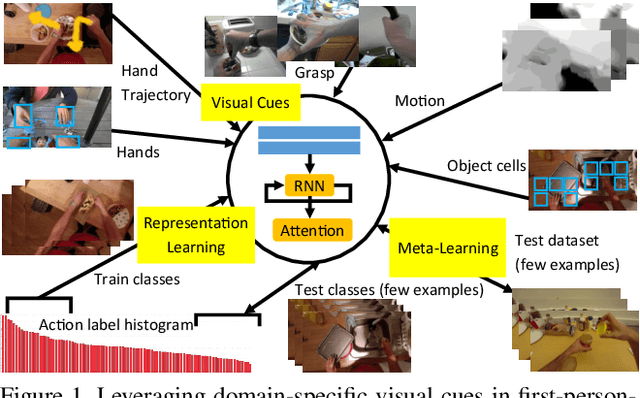

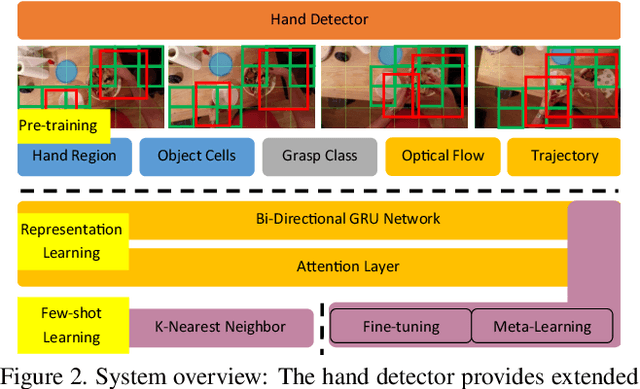
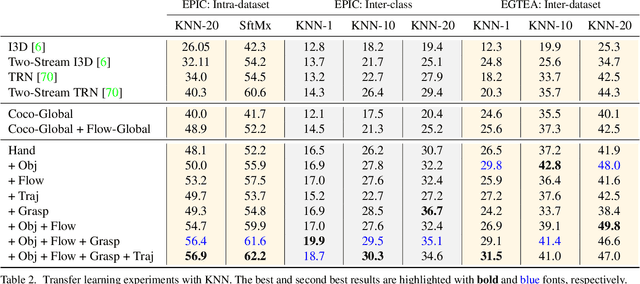
The lack of large-scale real datasets with annotationsmakes transfer learning a necessity for video activity under-standing. Within this scope, we aim at developing an effec-tive method for low-shot transfer learning for first-personaction classification. We leverage independently trained lo-cal visual cues to learn representations that can be trans-ferred from a source domain providing primitive action la-bels to a target domain with only a handful of examples.Such visual cues include object-object interactions, handgrasps and motion within regions that are a function of handlocations. We suggest a framework based on meta-learningto appropriately extract the distinctive and domain invari-ant components of the deployed visual cues, so to be able totransfer action classification models across public datasetscaptured with different scene configurations. We thoroughlyevaluate our methodology and report promising results overstate-of-the-art action classification approaches for bothinter-class and inter-dataset transfer.
Image to Images Translation for Multi-Task Organ Segmentation and Bone Suppression in Chest X-Ray Radiography
Jun 24, 2019Mohammad Eslami, Solale Tabarestani, Shadi Albarqouni, Ehsan Adeli, Nassir Navab, Malek Adjouadi
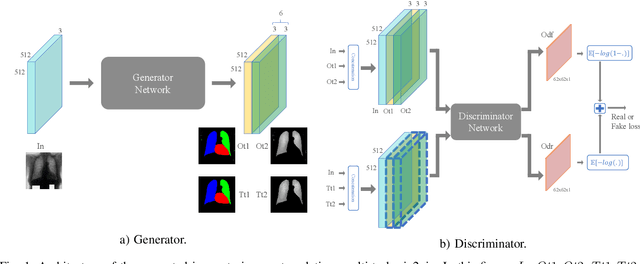
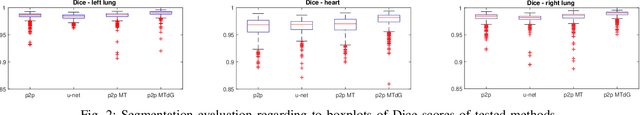
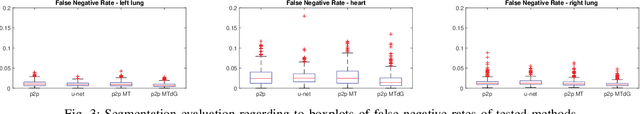
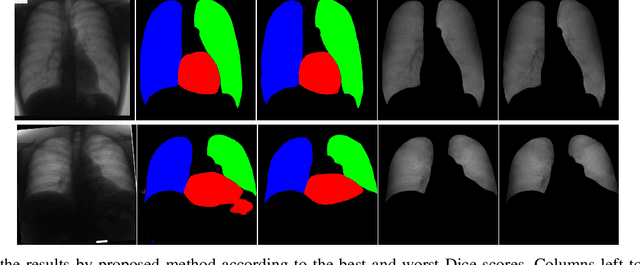
Chest X-ray radiography is one of the earliest medical imaging technologies and remains one of the most widely-used for the diagnosis, screening and treatment follow up of diseases related to lungs and heart. The literature in this field of research reports many interesting studies dealing with the challenging tasks of bone suppression and organ segmentation but performed separately, limiting any learning that comes with the consolidation of parameters that could optimize both processes. Although image processing could facilitate computer aided diagnosis, machine learning seems more amenable in dealing with the many parameters one would have to contend with to yield an near optimal classification or decision-making process. This study, and for the first time, introduces a multitask deep learning model that generates simultaneously the bone-suppressed image and the organ segmented image, minimizing as a consequence the number of parameters the model has to deal with and optimizing the processing time as well; while at the same time exploiting the interplay in these parameters so as to benefit the performance of both tasks. The design architecture of this model, which relies on a conditional generative adversarial network, reveals the process on how we managed to modify the well-established pix2pix network to fit the need for multitasking and hence extending the standard image-to-image network to the new image-to-images architecture. Dilated convolutions are also used to improve the results through a more effective receptive field assessment. A comparison of the proposed approach to state-of-the-art algorithms is provided to gauge the merits of the proposed approach.
`Project & Excite' Modules for Segmentation of Volumetric Medical Scans
Jun 12, 2019Anne-Marie Rickmann, Abhijit Guha Roy, Ignacio Sarasua, Nassir Navab, Christian Wachinger
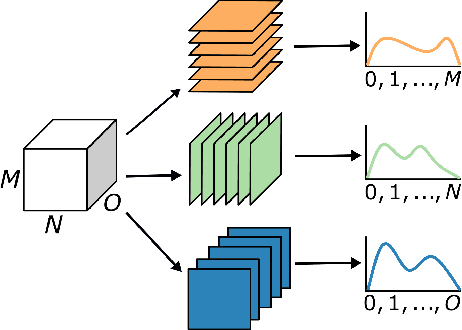
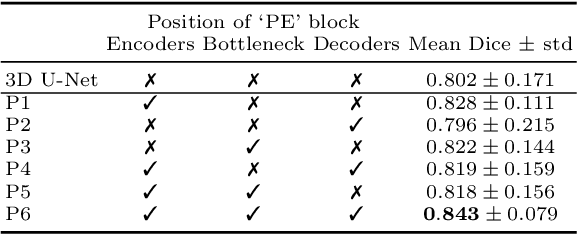
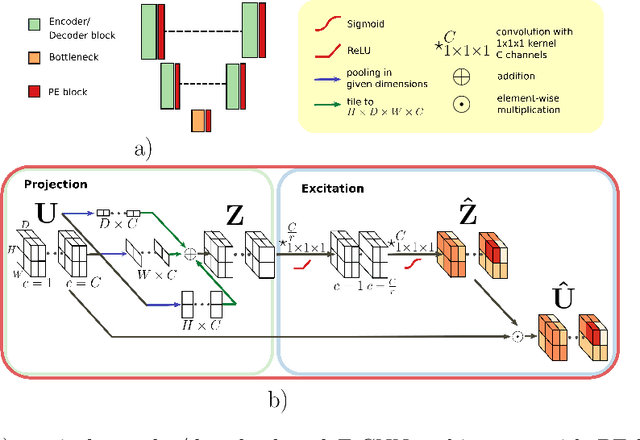
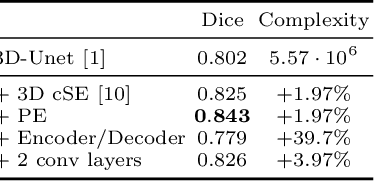
Fully Convolutional Neural Networks (F-CNNs) achieve state-of-the-art performance for image segmentation in medical imaging. Recently, squeeze and excitation (SE) modules and variations thereof have been introduced to recalibrate feature maps channel- and spatial-wise, which can boost performance while only minimally increasing model complexity. So far, the development of SE has focused on 2D images. In this paper, we propose `Project & Excite' (PE) modules that base upon the ideas of SE and extend them to operating on 3D volumetric images. `Project & Excite' does not perform global average pooling, but squeezes feature maps along different slices of a tensor separately to retain more spatial information that is subsequently used in the excitation step. We demonstrate that PE modules can be easily integrated in 3D U-Net, boosting performance by 5% Dice points, while only increasing the model complexity by 2%. We evaluate the PE module on two challenging tasks, whole-brain segmentation of MRI scans and whole-body segmentation of CT scans. Code: https://github.com/ai-med/squeeze_and_excitation
An Uncertainty-Driven GCN Refinement Strategy for Organ Segmentation
Jun 05, 2019Roger D. Soberanis-Mukul, Shadi Albarqouni, Nassir Navab
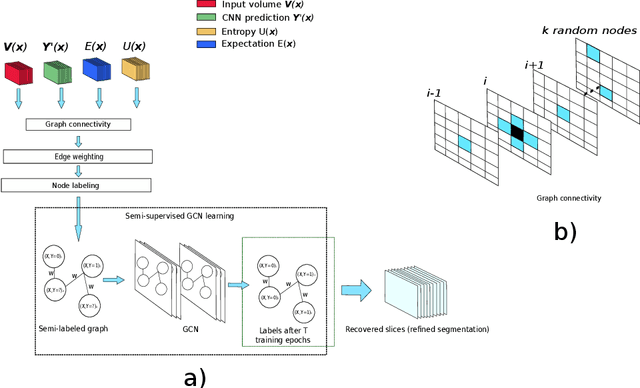
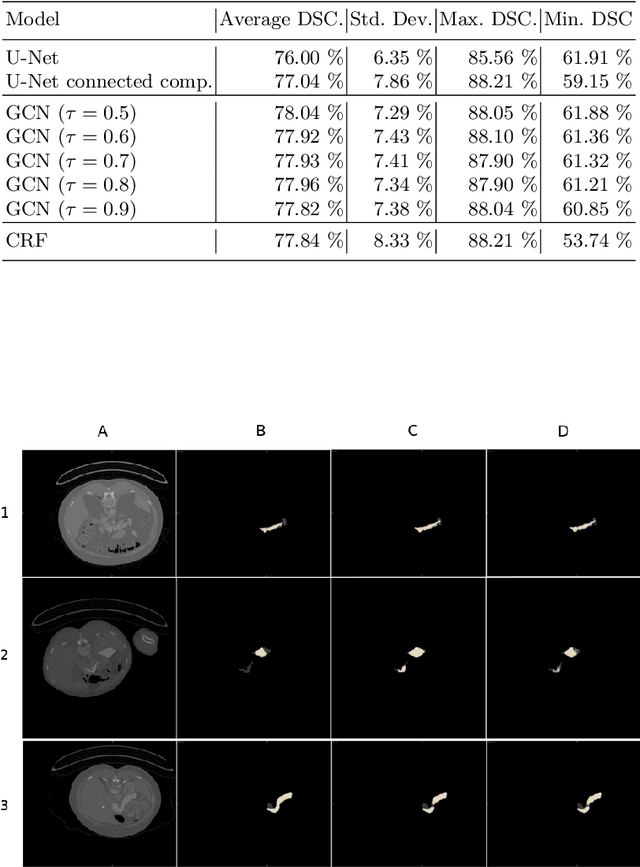
Organ segmentation is an important pre-processing step in many computer assisted intervention and computer assisted diagnosis methods. In recent years, CNNs have dominated the state of the art in this task. Organ segmentation scenarios present a challenging environment for these methods due to high variability in shape, similarity with background, etc. This leads to the generation of false negative and false positive regions in the output segmentation. In this context, the uncertainty analysis of the model can provide us with useful information about potentially misclassified elements. In this work we propose a method based on uncertainty analysis and graph convolutional networks as a post-processing step for segmentation. For this, we employ the uncertainty levels of the CNN to formulate a semi-supervised graph learning problem that is solved by training a GCN on the low uncertainty elements. Finally, we evaluate the full graph on the trained GCN to get the refined segmentation. We compare our framework with CRF on a graph-like data representation as refinement strategy.
Perceptual Embedding Consistency for Seamless Reconstruction of Tilewise Style Transfer
Jun 03, 2019Amal Lahiani, Nassir Navab, Shadi Albarqouni, Eldad Klaiman
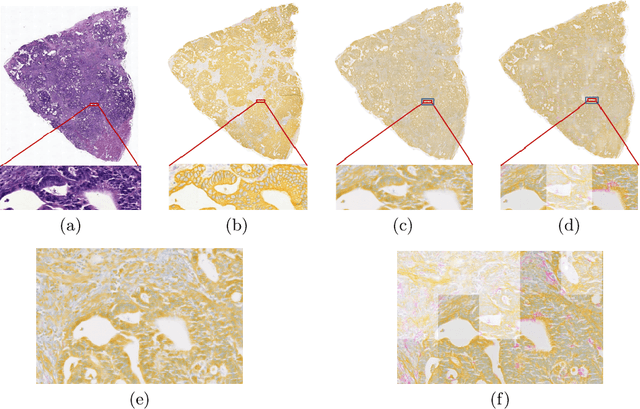
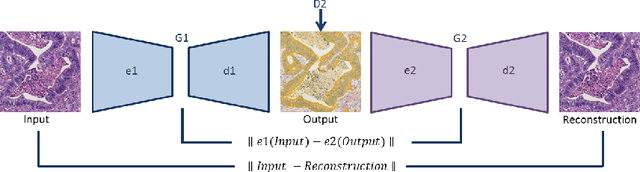


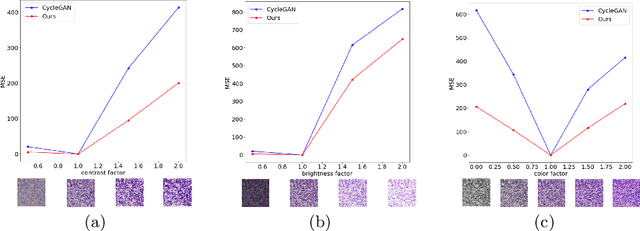
Style transfer is a field with growing interest and use cases in deep learning. Recent work has shown Generative Adversarial Networks(GANs) can be used to create realistic images of virtually stained slide images in digital pathology with clinically validated interpretability. Digital pathology images are typically of extremely high resolution, making tilewise analysis necessary for deep learning applications. It has been shown that image generators with instance normalization can cause a tiling artifact when a large image is reconstructed from the tilewise analysis. We introduce a novel perceptual embedding consistency loss significantly reducing the tiling artifact created in the reconstructed whole slide image (WSI). We validate our results by comparing virtually stained slide images with consecutive real stained tissue slide images. We also demonstrate that our model is more robust to contrast, color and brightness perturbations by running comparative sensitivity analysis tests.
BrainTorrent: A Peer-to-Peer Environment for Decentralized Federated Learning
May 16, 2019Abhijit Guha Roy, Shayan Siddiqui, Sebastian Pölsterl, Nassir Navab, Christian Wachinger
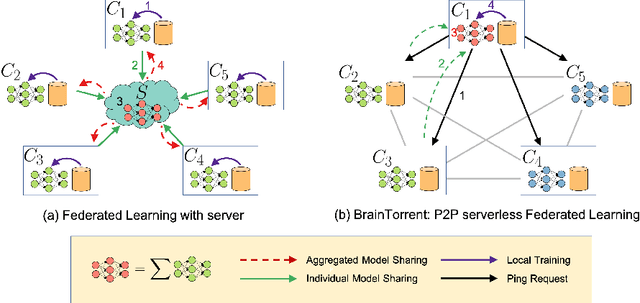
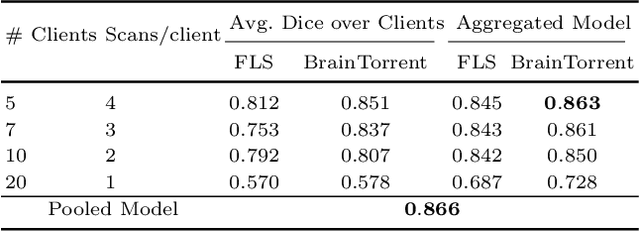
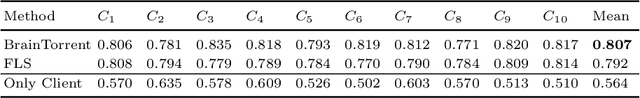
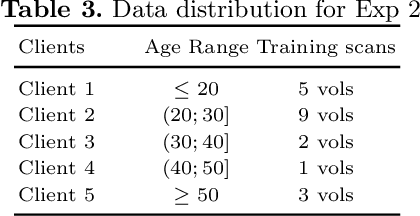
Access to sufficient annotated data is a common challenge in training deep neural networks on medical images. As annotating data is expensive and time-consuming, it is difficult for an individual medical center to reach large enough sample sizes to build their own, personalized models. As an alternative, data from all centers could be pooled to train a centralized model that everyone can use. However, such a strategy is often infeasible due to the privacy-sensitive nature of medical data. Recently, federated learning (FL) has been introduced to collaboratively learn a shared prediction model across centers without the need for sharing data. In FL, clients are locally training models on site-specific datasets for a few epochs and then sharing their model weights with a central server, which orchestrates the overall training process. Importantly, the sharing of models does not compromise patient privacy. A disadvantage of FL is the dependence on a central server, which requires all clients to agree on one trusted central body, and whose failure would disrupt the training process of all clients. In this paper, we introduce BrainTorrent, a new FL framework without a central server, particularly targeted towards medical applications. BrainTorrent presents a highly dynamic peer-to-peer environment, where all centers directly interact with each other without depending on a central body. We demonstrate the overall effectiveness of FL for the challenging task of whole brain segmentation and observe that the proposed server-less BrainTorrent approach does not only outperform the traditional server-based one but reaches a similar performance to a model trained on pooled data.
Learning Interpretable Features via Adversarially Robust Optimization
May 09, 2019Ashkan Khakzar, Shadi Albarqouni, Nassir Navab
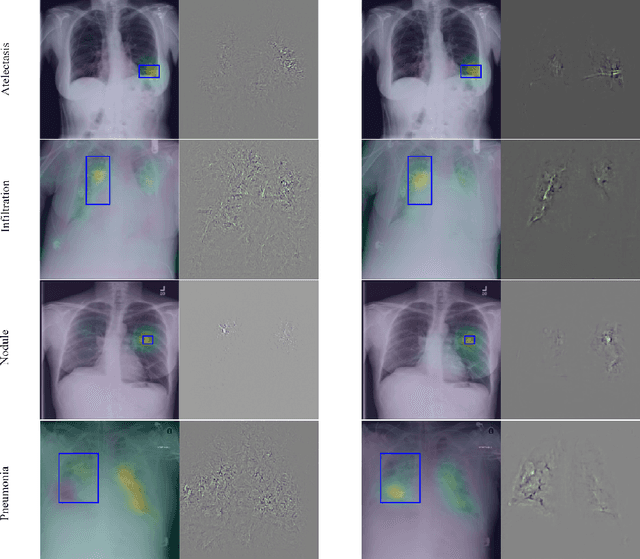
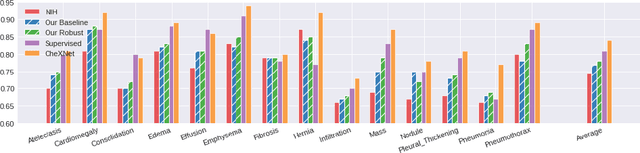
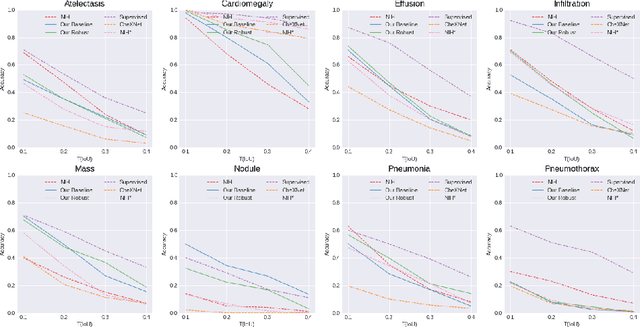

Neural networks are proven to be remarkably successful for classification and diagnosis in medical applications. However, the ambiguity in the decision-making process and the interpretability of the learned features is a matter of concern. In this work, we propose a method for improving the feature interpretability of neural network classifiers. Initially, we propose a baseline convolutional neural network with state of the art performance in terms of accuracy and weakly supervised localization. Subsequently, the loss is modified to integrate robustness to adversarial examples into the training process. In this work, feature interpretability is quantified via evaluating the weakly supervised localization using the ground truth bounding boxes. Interpretability is also visually assessed using class activation maps and saliency maps. The method is applied to NIH ChestX-ray14, the largest publicly available chest x-rays dataset. We demonstrate that the adversarially robust optimization paradigm improves feature interpretability both quantitatively and visually.
Multi-modal Graph Fusion for Inductive Disease Classification in Incomplete Datasets
May 08, 2019Gerome Vivar, Hendrik Burwinkel, Anees Kazi, Andreas Zwergal, Nassir Navab, Seyed-Ahmad Ahmadi
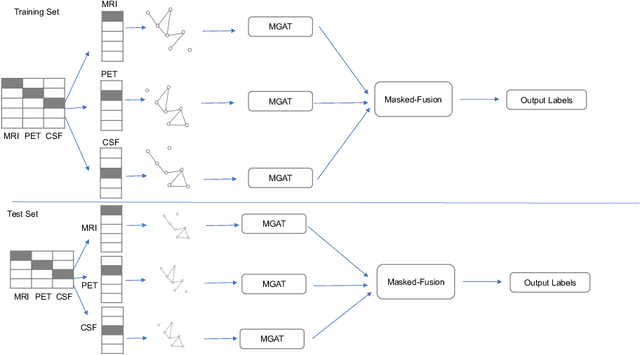
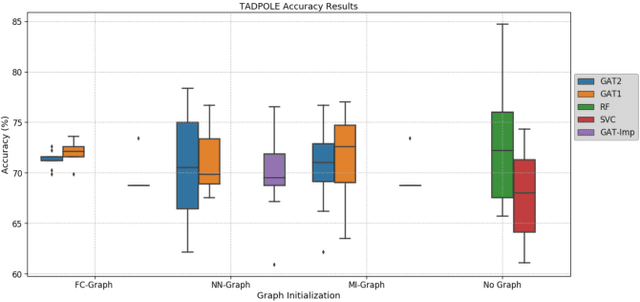
Clinical diagnostic decision making and population-based studies often rely on multi-modal data which is noisy and incomplete. Recently, several works proposed geometric deep learning approaches to solve disease classification, by modeling patients as nodes in a graph, along with graph signal processing of multi-modal features. Many of these approaches are limited by assuming modality- and feature-completeness, and by transductive inference, which requires re-training of the entire model for each new test sample. In this work, we propose a novel inductive graph-based approach that can generalize to out-of-sample patients, despite missing features from entire modalities per patient. We propose multi-modal graph fusion which is trained end-to-end towards node-level classification. We demonstrate the fundamental working principle of this method on a simplified MNIST toy dataset. In experiments on medical data, our method outperforms single static graph approach in multi-modal disease classification.
Adaptive image-feature learning for disease classification using inductive graph networks
May 08, 2019Hendrik Burwinkel, Anees Kazi, Gerome Vivar, Shadi Albarqouni, Guillaume Zahnd, Nassir Navab, Seyed-Ahmad Ahmadi
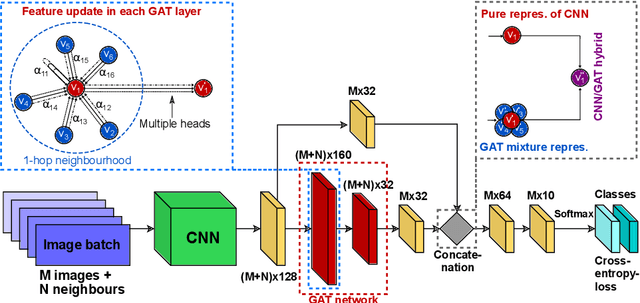
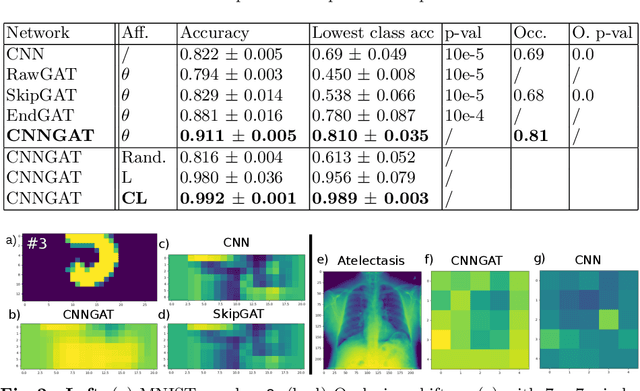
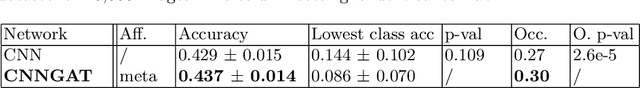
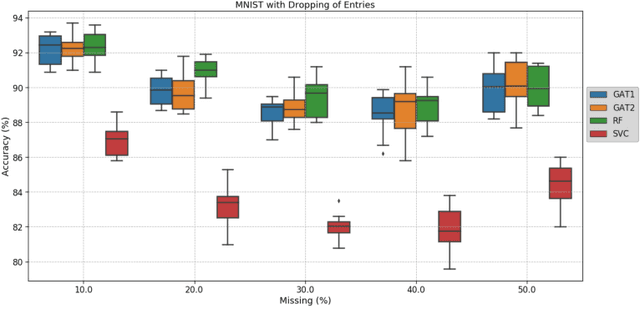
Recently, Geometric Deep Learning (GDL) has been introduced as a novel and versatile framework for computer-aided disease classification. GDL uses patient meta-information such as age and gender to model patient cohort relations in a graph structure. Concepts from graph signal processing are leveraged to learn the optimal mapping of multi-modal features, e.g. from images to disease classes. Related studies so far have considered image features that are extracted in a pre-processing step. We hypothesize that such an approach prevents the network from optimizing feature representations towards achieving the best performance in the graph network. We propose a new network architecture that exploits an inductive end-to-end learning approach for disease classification, where filters from both the CNN and the graph are trained jointly. We validate this architecture against state-of-the-art inductive graph networks and demonstrate significantly improved classification scores on a modified MNIST toy dataset, as well as comparable classification results with higher stability on a chest X-ray image dataset. Additionally, we explain how the structural information of the graph affects both the image filters and the feature learning.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge