U-GAT: Multimodal Graph Attention Network for COVID-19 Outcome Prediction
Jul 29, 2021Matthias Keicher, Hendrik Burwinkel, David Bani-Harouni, Magdalini Paschali, Tobias Czempiel, Egon Burian, Marcus R. Makowski, Rickmer Braren, Nassir Navab, Thomas Wendler
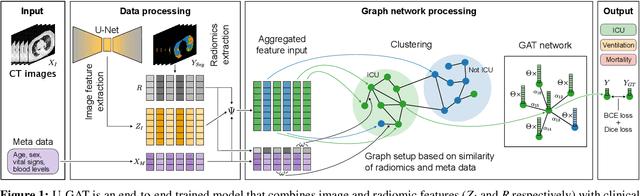
During the first wave of COVID-19, hospitals were overwhelmed with the high number of admitted patients. An accurate prediction of the most likely individual disease progression can improve the planning of limited resources and finding the optimal treatment for patients. However, when dealing with a newly emerging disease such as COVID-19, the impact of patient- and disease-specific factors (e.g. body weight or known co-morbidities) on the immediate course of disease is by and large unknown. In the case of COVID-19, the need for intensive care unit (ICU) admission of pneumonia patients is often determined only by acute indicators such as vital signs (e.g. breathing rate, blood oxygen levels), whereas statistical analysis and decision support systems that integrate all of the available data could enable an earlier prognosis. To this end, we propose a holistic graph-based approach combining both imaging and non-imaging information. Specifically, we introduce a multimodal similarity metric to build a population graph for clustering patients and an image-based end-to-end Graph Attention Network to process this graph and predict the COVID-19 patient outcomes: admission to ICU, need for ventilation and mortality. Additionally, the network segments chest CT images as an auxiliary task and extracts image features and radiomics for feature fusion with the available metadata. Results on a dataset collected in Klinikum rechts der Isar in Munich, Germany show that our approach outperforms single modality and non-graph baselines. Moreover, our clustering and graph attention allow for increased understanding of the patient relationships within the population graph and provide insight into the network's decision-making process.
Segmentation in Style: Unsupervised Semantic Image Segmentation with Stylegan and CLIP
Jul 26, 2021Daniil Pakhomov, Sanchit Hira, Narayani Wagle, Kemar E. Green, Nassir Navab
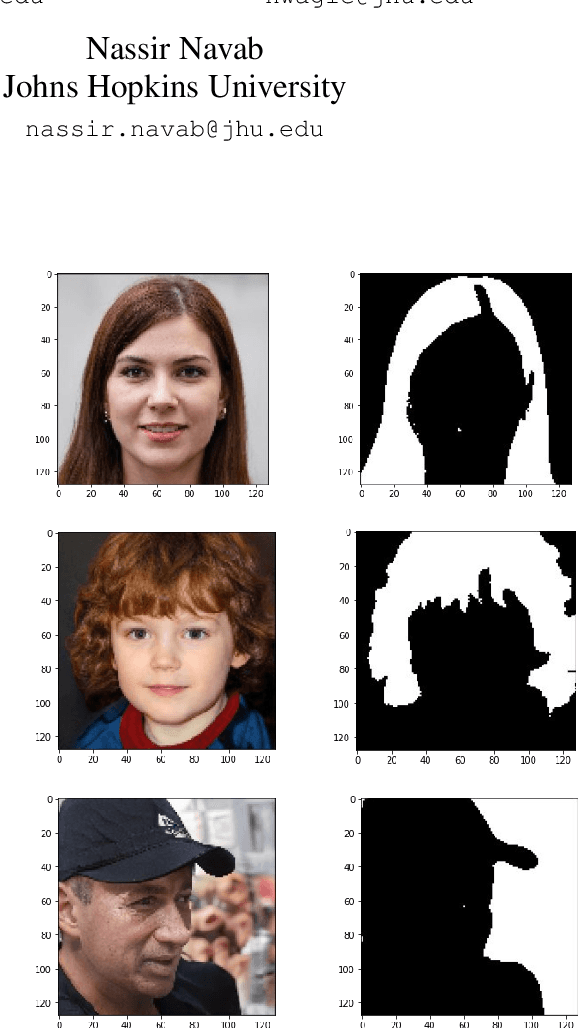
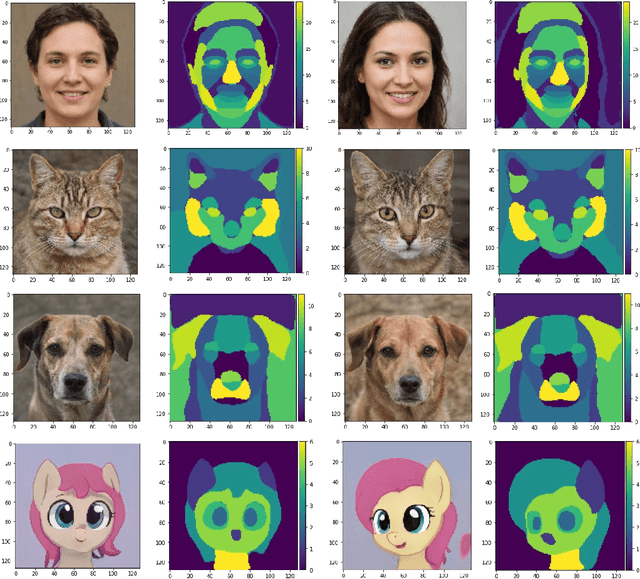


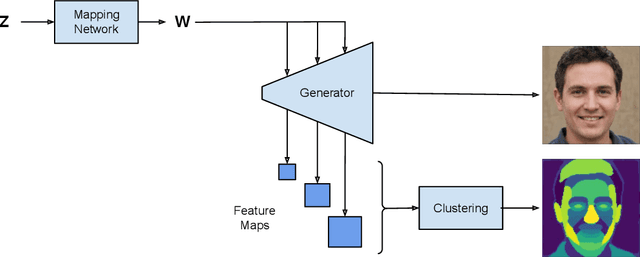
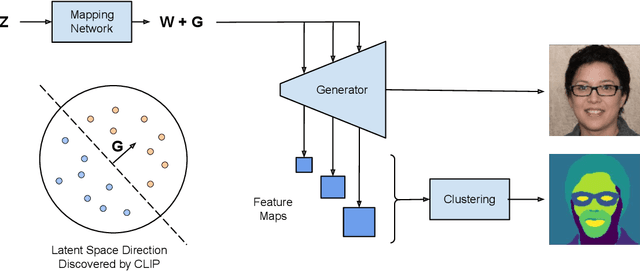
We introduce a method that allows to automatically segment images into semantically meaningful regions without human supervision. Derived regions are consistent across different images and coincide with human-defined semantic classes on some datasets. In cases where semantic regions might be hard for human to define and consistently label, our method is still able to find meaningful and consistent semantic classes. In our work, we use pretrained StyleGAN2~\cite{karras2020analyzing} generative model: clustering in the feature space of the generative model allows to discover semantic classes. Once classes are discovered, a synthetic dataset with generated images and corresponding segmentation masks can be created. After that a segmentation model is trained on the synthetic dataset and is able to generalize to real images. Additionally, by using CLIP~\cite{radford2021learning} we are able to use prompts defined in a natural language to discover some desired semantic classes. We test our method on publicly available datasets and show state-of-the-art results.
Deformation-Aware Robotic 3D Ultrasound
Jul 18, 2021Zhongliang Jiang, Yue Zhou, Yuan Bi, Mingchuan Zhou, Thomas Wendler, Nassir Navab
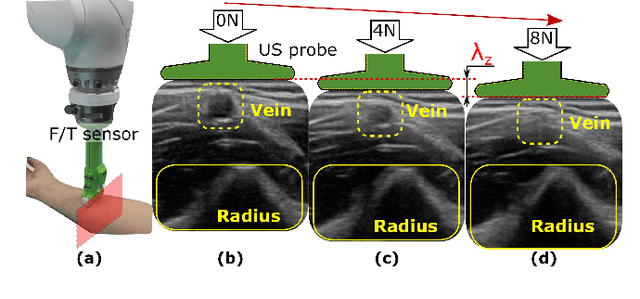
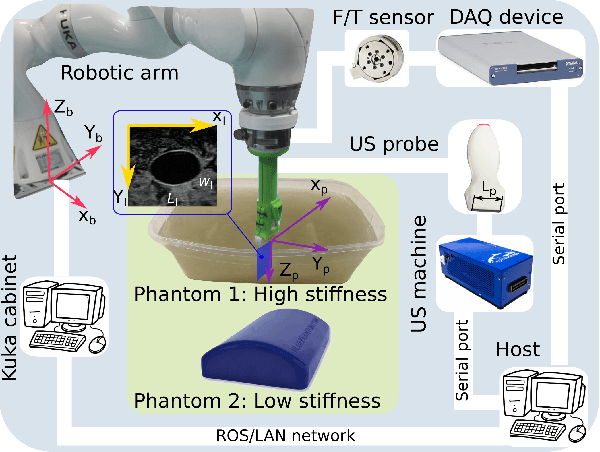
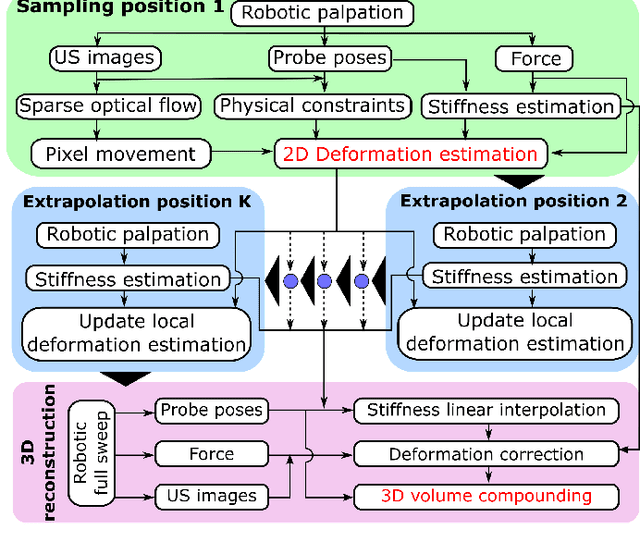
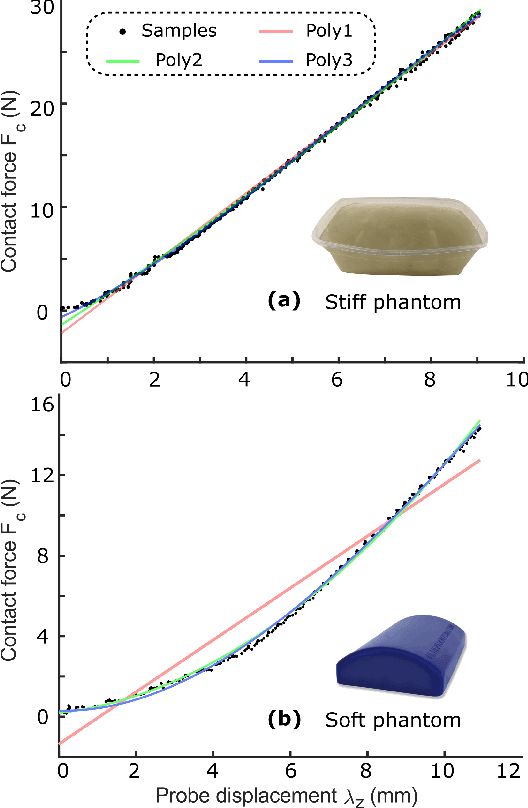
Tissue deformation in ultrasound (US) imaging leads to geometrical errors when measuring tissues due to the pressure exerted by probes. Such deformation has an even larger effect on 3D US volumes as the correct compounding is limited by the inconsistent location and geometry. This work proposes a patient-specified stiffness-based method to correct the tissue deformations in robotic 3D US acquisitions. To obtain the patient-specified model, robotic palpation is performed at sampling positions on the tissue. The contact force, US images and the probe poses of the palpation procedure are recorded. The contact force and the probe poses are used to estimate the nonlinear tissue stiffness. The images are fed to an optical flow algorithm to compute the pixel displacement. Then the pixel-wise tissue deformation under different forces is characterized by a coupled quadratic regression. To correct the deformation at unseen positions on the trajectory for building 3D volumes, an interpolation is performed based on the stiffness values computed at the sampling positions. With the stiffness and recorded force, the tissue displacement could be corrected. The method was validated on two blood vessel phantoms with different stiffness. The results demonstrate that the method can effectively correct the force-induced deformation and finally generate 3D tissue geometries
Motion-Aware Robotic 3D Ultrasound
Jul 13, 2021Zhongliang Jiang, Hanyu Wang, Zhenyu Li, Matthias Grimm, Mingchuan Zhou, Ulrich Eck, Sandra V. Brecht, Tim C. Lueth, Thomas Wendler, Nassir Navab
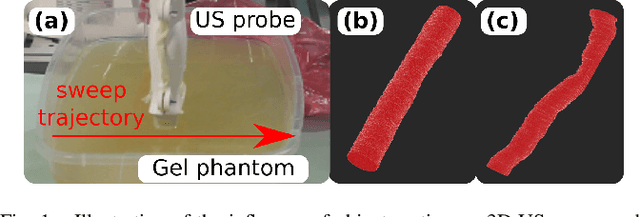

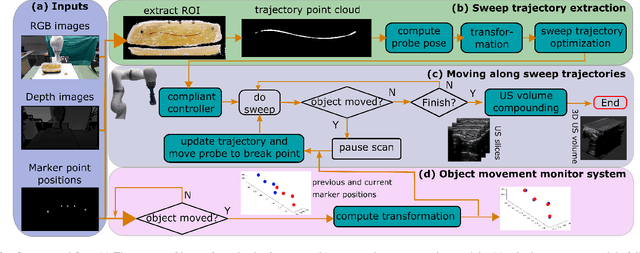
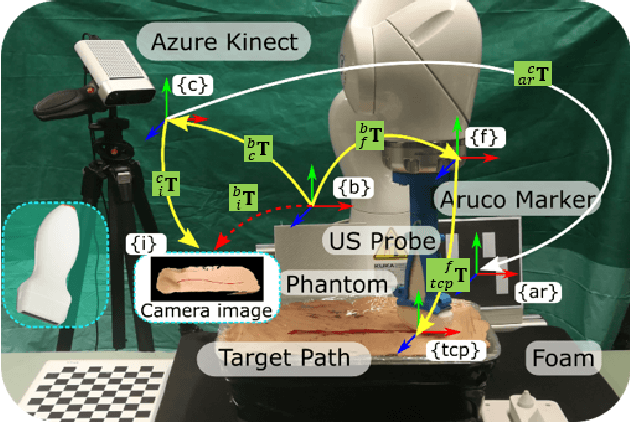
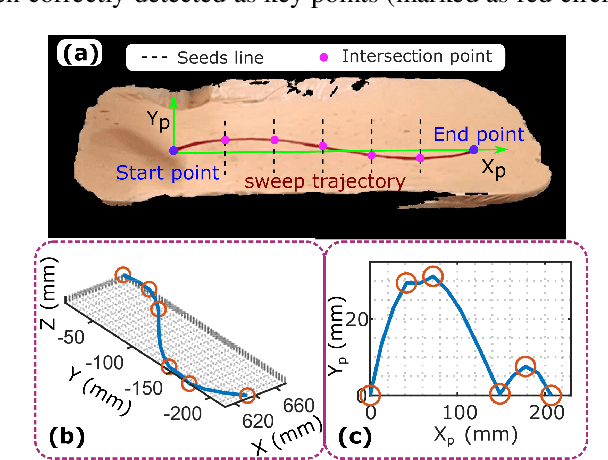
Robotic three-dimensional (3D) ultrasound (US) imaging has been employed to overcome the drawbacks of traditional US examinations, such as high inter-operator variability and lack of repeatability. However, object movement remains a challenge as unexpected motion decreases the quality of the 3D compounding. Furthermore, attempted adjustment of objects, e.g., adjusting limbs to display the entire limb artery tree, is not allowed for conventional robotic US systems. To address this challenge, we propose a vision-based robotic US system that can monitor the object's motion and automatically update the sweep trajectory to provide 3D compounded images of the target anatomy seamlessly. To achieve these functions, a depth camera is employed to extract the manually planned sweep trajectory after which the normal direction of the object is estimated using the extracted 3D trajectory. Subsequently, to monitor the movement and further compensate for this motion to accurately follow the trajectory, the position of firmly attached passive markers is tracked in real-time. Finally, a step-wise compounding was performed. The experiments on a gel phantom demonstrate that the system can resume a sweep when the object is not stationary during scanning.
Deep Direct Volume Rendering: Learning Visual Feature Mappings From Exemplary Images
Jun 09, 2021Jakob Weiss, Nassir Navab
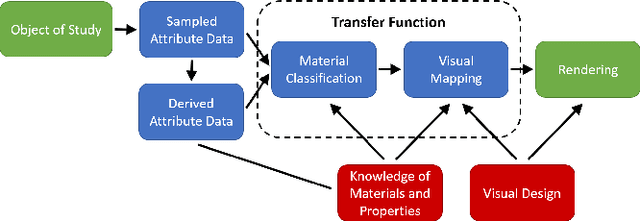
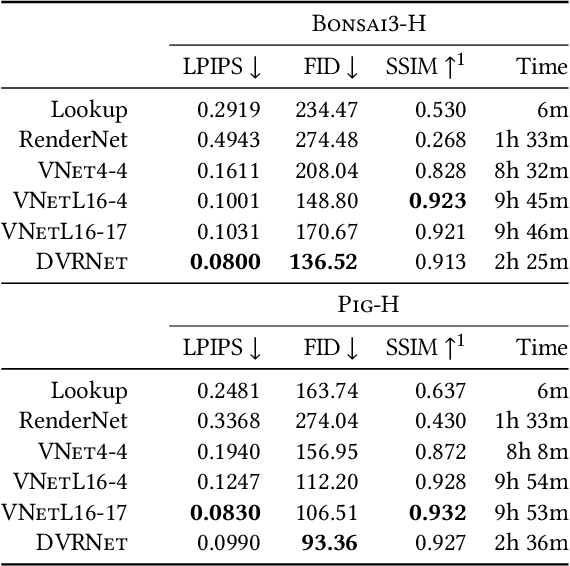
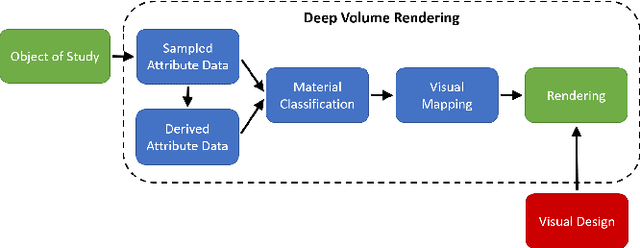
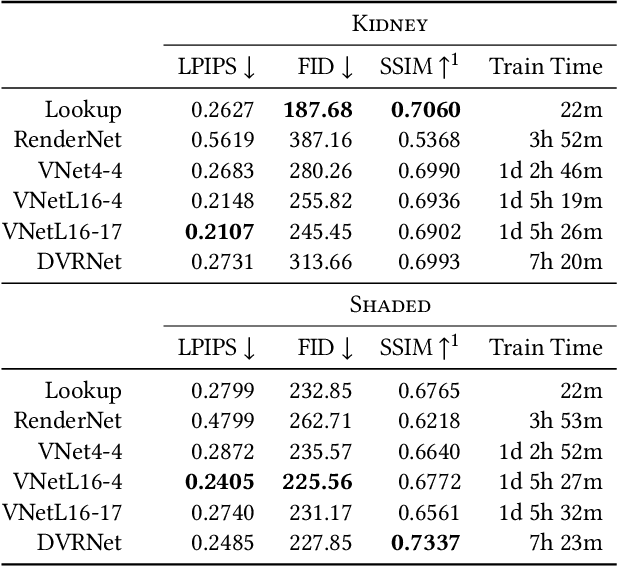
Volume Rendering is an important technique for visualizing three-dimensional scalar data grids and is commonly employed for scientific and medical image data. Direct Volume Rendering (DVR) is a well established and efficient rendering algorithm for volumetric data. Neural rendering uses deep neural networks to solve inverse rendering tasks and applies techniques similar to DVR. However, it has not been demonstrated successfully for the rendering of scientific volume data. In this work, we introduce Deep Direct Volume Rendering (DeepDVR), a generalization of DVR that allows for the integration of deep neural networks into the DVR algorithm. We conceptualize the rendering in a latent color space, thus enabling the use of deep architectures to learn implicit mappings for feature extraction and classification, replacing explicit feature design and hand-crafted transfer functions. Our generalization serves to derive novel volume rendering architectures that can be trained end-to-end directly from examples in image space, obviating the need to manually define and fine-tune multidimensional transfer functions while providing superior classification strength. We further introduce a novel stepsize annealing scheme to accelerate the training of DeepDVR models and validate its effectiveness in a set of experiments. We validate our architectures on two example use cases: (1) learning an optimized rendering from manually adjusted reference images for a single volume and (2) learning advanced visualization concepts like shading and semantic colorization that generalize to unseen volume data. We find that deep volume rendering architectures with explicit modeling of the DVR pipeline effectively enable end-to-end learning of scientific volume rendering tasks from target images.
Multimodal Semantic Scene Graphs for Holistic Modeling of Surgical Procedures
Jun 09, 2021Ege Özsoy, Evin Pınar Örnek, Ulrich Eck, Federico Tombari, Nassir Navab
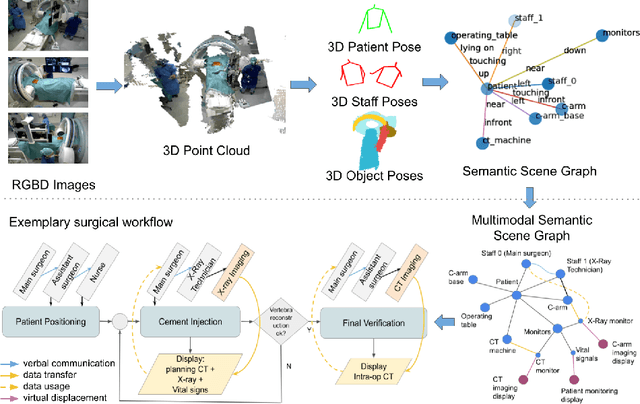
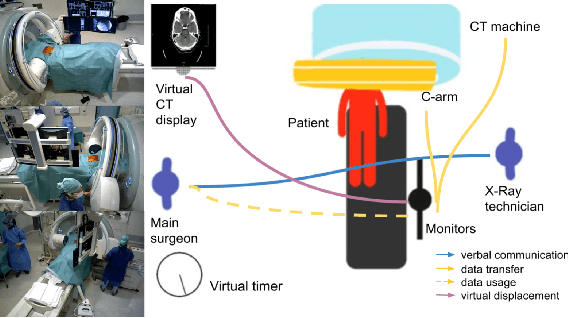
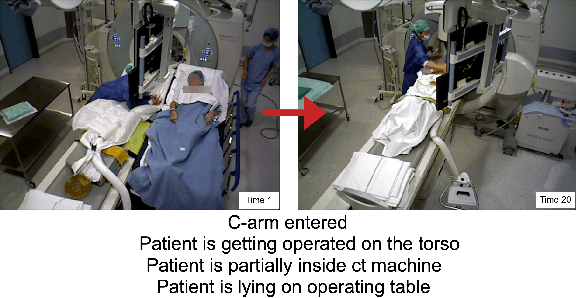
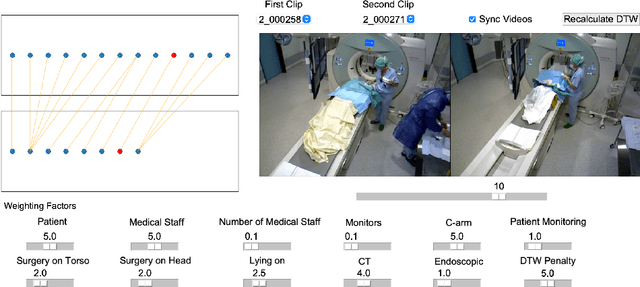
From a computer science viewpoint, a surgical domain model needs to be a conceptual one incorporating both behavior and data. It should therefore model actors, devices, tools, their complex interactions and data flow. To capture and model these, we take advantage of the latest computer vision methodologies for generating 3D scene graphs from camera views. We then introduce the Multimodal Semantic Scene Graph (MSSG) which aims at providing a unified symbolic, spatiotemporal and semantic representation of surgical procedures. This methodology aims at modeling the relationship between different components in surgical domain including medical staff, imaging systems, and surgical devices, opening the path towards holistic understanding and modeling of surgical procedures. We then use MSSG to introduce a dynamically generated graphical user interface tool for surgical procedure analysis which could be used for many applications including process optimization, OR design and automatic report generation. We finally demonstrate that the proposed MSSGs could also be used for synchronizing different complex surgical procedures. While the system still needs to be integrated into real operating rooms before getting validated, this conference paper aims mainly at providing the community with the basic principles of this novel concept through a first prototypal partial realization based on MVOR dataset.
Rethinking Ultrasound Augmentation: A Physics-Inspired Approach
May 05, 2021Maria Tirindelli, Christine Eilers, Walter Simson, Magdalini Paschali, Mohammad Farid Azampour, Nassir Navab
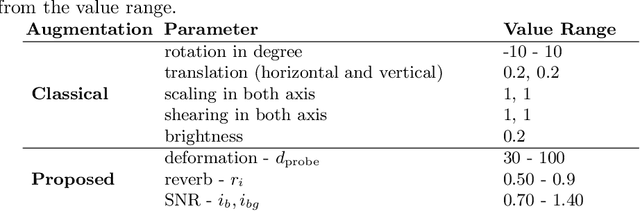
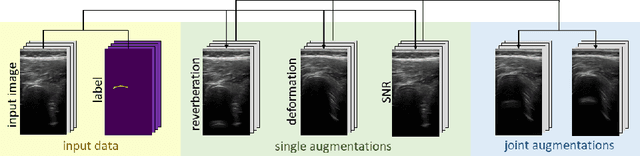

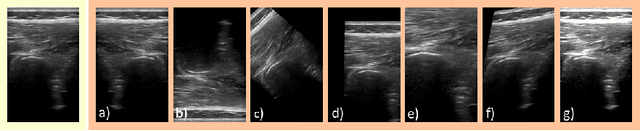
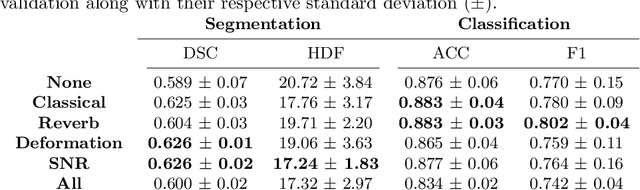
Medical Ultrasound (US), despite its wide use, is characterized by artifacts and operator dependency. Those attributes hinder the gathering and utilization of US datasets for the training of Deep Neural Networks used for Computer-Assisted Intervention Systems. Data augmentation is commonly used to enhance model generalization and performance. However, common data augmentation techniques, such as affine transformations do not align with the physics of US and, when used carelessly can lead to unrealistic US images. To this end, we propose a set of physics-inspired transformations, including deformation, reverb and Signal-to-Noise Ratio, that we apply on US B-mode images for data augmentation. We evaluate our method on a new spine US dataset for the tasks of bone segmentation and classification.
GKD: Semi-supervised Graph Knowledge Distillation for Graph-Independent Inference
Apr 08, 2021Mahsa Ghorbani, Mojtaba Bahrami, Anees Kazi, Mahdieh SoleymaniBaghshah, Hamid R. Rabiee, Nassir Navab
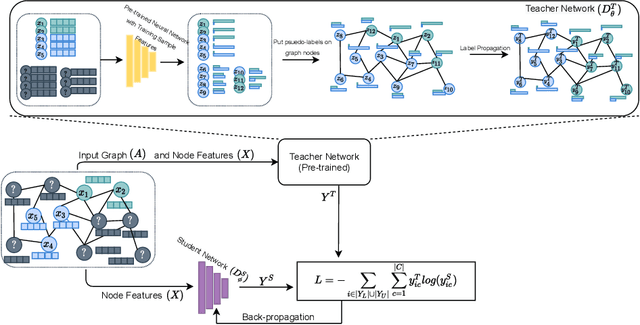
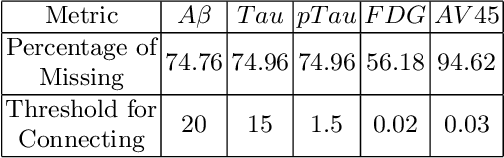
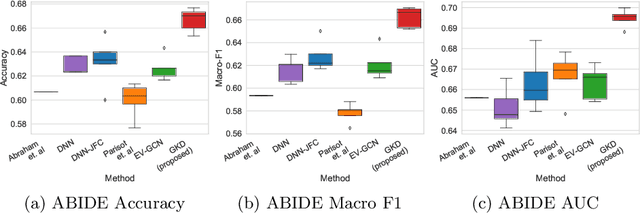
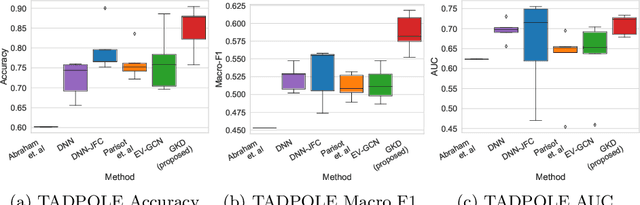
The increased amount of multi-modal medical data has opened the opportunities to simultaneously process various modalities such as imaging and non-imaging data to gain a comprehensive insight into the disease prediction domain. Recent studies using Graph Convolutional Networks (GCNs) provide novel semi-supervised approaches for integrating heterogeneous modalities while investigating the patients' associations for disease prediction. However, when the meta-data used for graph construction is not available at inference time (e.g., coming from a distinct population), the conventional methods exhibit poor performance. To address this issue, we propose a novel semi-supervised approach named GKD based on knowledge distillation. We train a teacher component that employs the label-propagation algorithm besides a deep neural network to benefit from the graph and non-graph modalities only in the training phase. The teacher component embeds all the available information into the soft pseudo-labels. The soft pseudo-labels are then used to train a deep student network for disease prediction of unseen test data for which the graph modality is unavailable. We perform our experiments on two public datasets for diagnosing Autism spectrum disorder, and Alzheimer's disease, along with a thorough analysis on synthetic multi-modal datasets. According to these experiments, GKD outperforms the previous graph-based deep learning methods in terms of accuracy, AUC, and Macro F1.
Towards Semantic Interpretation of Thoracic Disease and COVID-19 Diagnosis Models
Apr 04, 2021Ashkan Khakzar, Sabrina Musatian, Jonas Buchberger, Icxel Valeriano Quiroz, Nikolaus Pinger, Soroosh Baselizadeh, Seong Tae Kim, Nassir Navab
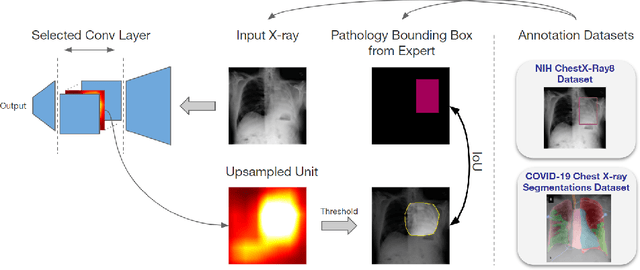
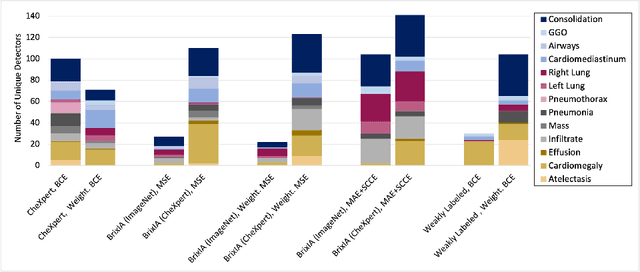
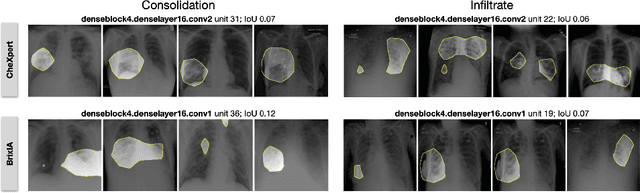
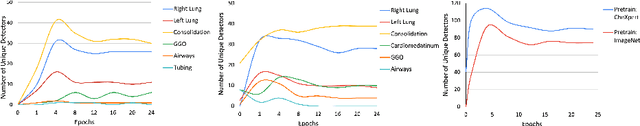
Convolutional neural networks are showing promise in the automatic diagnosis of thoracic pathologies on chest x-rays. Their black-box nature has sparked many recent works to explain the prediction via input feature attribution methods (aka saliency methods). However, input feature attribution methods merely identify the importance of input regions for the prediction and lack semantic interpretation of model behavior. In this work, we first identify the semantics associated with internal units (feature maps) of the network. We proceed to investigate the following questions; Does a regression model that is only trained with COVID-19 severity scores implicitly learn visual patterns associated with thoracic pathologies? Does a network that is trained on weakly labeled data (e.g. healthy, unhealthy) implicitly learn pathologies? Moreover, we investigate the effect of pretraining and data imbalance on the interpretability of learned features. In addition to the analysis, we propose semantic attribution to semantically explain each prediction. We present our findings using publicly available chest pathologies (CheXpert, NIH ChestX-ray8) and COVID-19 datasets (BrixIA, and COVID-19 chest X-ray segmentation dataset). The Code is publicly available.
Explaining COVID-19 and Thoracic Pathology Model Predictions by Identifying Informative Input Features
Apr 01, 2021Ashkan Khakzar, Yang Zhang, Wejdene Mansour, Yuezhi Cai, Yawei Li, Yucheng Zhang, Seong Tae Kim, Nassir Navab
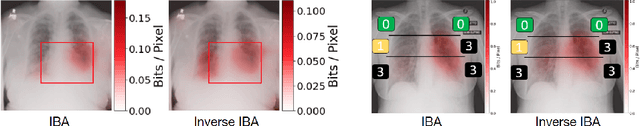
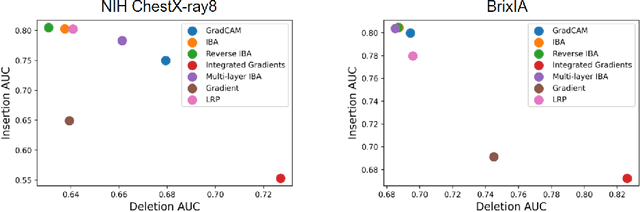
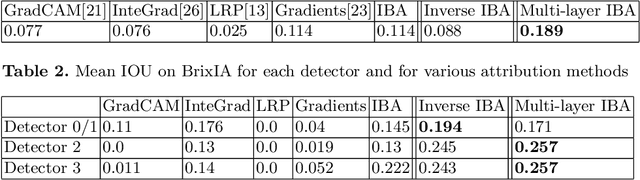
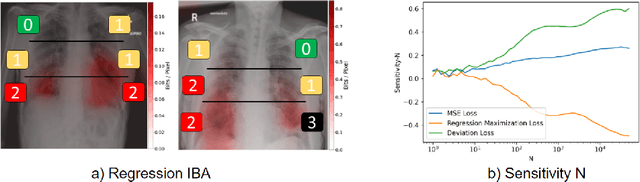
Neural networks have demonstrated remarkable performance in classification and regression tasks on chest X-rays. In order to establish trust in the clinical routine, the networks' prediction mechanism needs to be interpretable. One principal approach to interpretation is feature attribution. Feature attribution methods identify the importance of input features for the output prediction. Building on Information Bottleneck Attribution (IBA) method, for each prediction we identify the chest X-ray regions that have high mutual information with the network's output. Original IBA identifies input regions that have sufficient predictive information. We propose Inverse IBA to identify all informative regions. Thus all predictive cues for pathologies are highlighted on the X-rays, a desirable property for chest X-ray diagnosis. Moreover, we propose Regression IBA for explaining regression models. Using Regression IBA we observe that a model trained on cumulative severity score labels implicitly learns the severity of different X-ray regions. Finally, we propose Multi-layer IBA to generate higher resolution and more detailed attribution/saliency maps. We evaluate our methods using both human-centric (ground-truth-based) interpretability metrics, and human-independent feature importance metrics on NIH Chest X-ray8 and BrixIA datasets. The Code is publicly available.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge