Marco Lorenzi
EPIONE, UCA,3iA Côte d'Azur
Sequential Informed Federated Unlearning: Efficient and Provable Client Unlearning in Federated Optimization
Nov 21, 2022



Abstract:The aim of Machine Unlearning (MU) is to provide theoretical guarantees on the removal of the contribution of a given data point from a training procedure. Federated Unlearning (FU) consists in extending MU to unlearn a given client's contribution from a federated training routine. Current FU approaches are generally not scalable, and do not come with sound theoretical quantification of the effectiveness of unlearning. In this work we present Informed Federated Unlearning (IFU), a novel efficient and quantifiable FU approach. Upon unlearning request from a given client, IFU identifies the optimal FL iteration from which FL has to be reinitialized, with unlearning guarantees obtained through a randomized perturbation mechanism. The theory of IFU is also extended to account for sequential unlearning requests. Experimental results on different tasks and dataset show that IFU leads to more efficient unlearning procedures as compared to basic re-training and state-of-the-art FU approaches.
FLamby: Datasets and Benchmarks for Cross-Silo Federated Learning in Realistic Healthcare Settings
Oct 10, 2022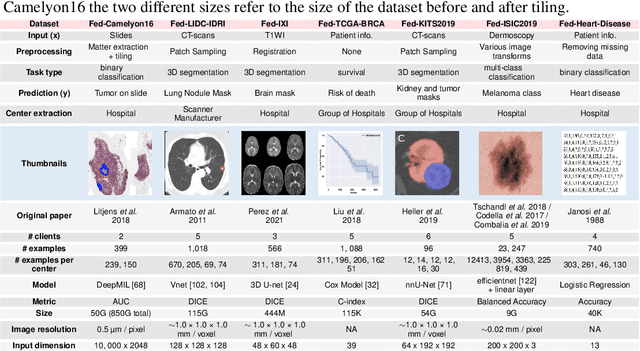
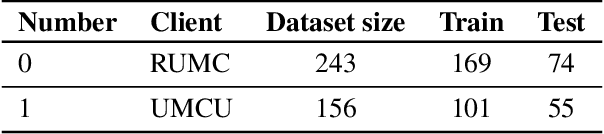
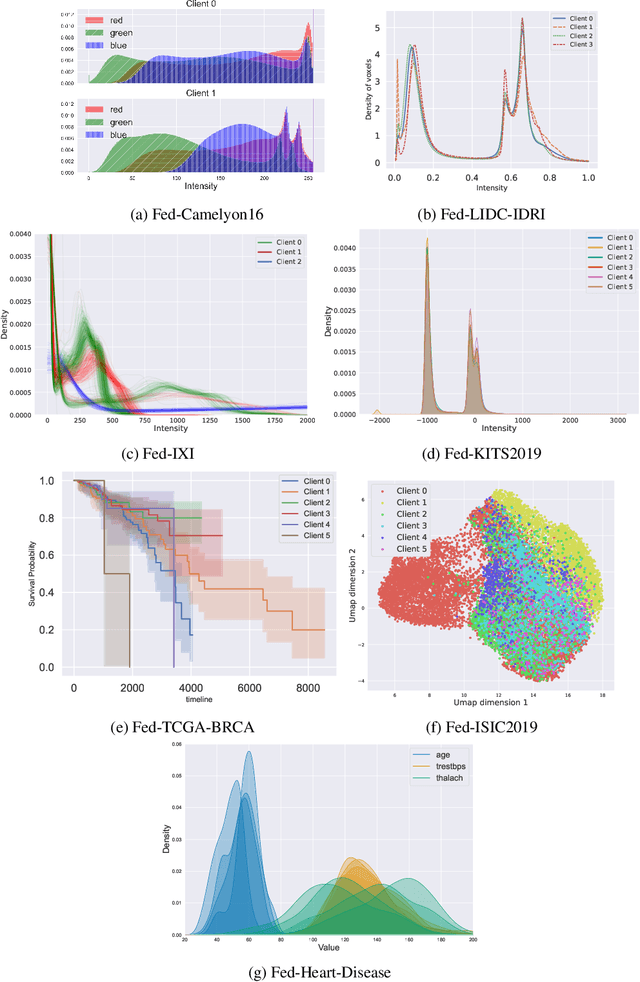
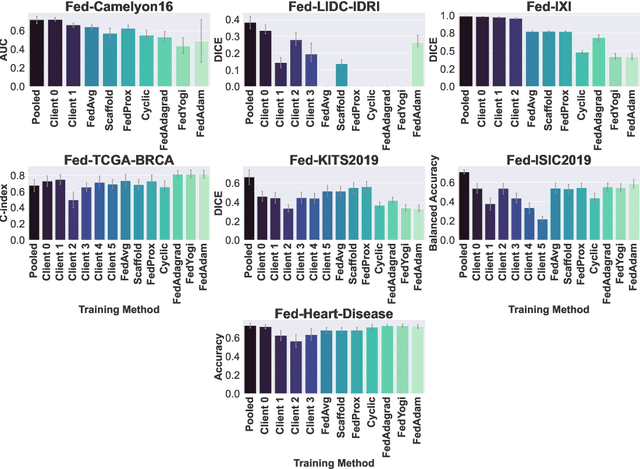

Abstract:Federated Learning (FL) is a novel approach enabling several clients holding sensitive data to collaboratively train machine learning models, without centralizing data. The cross-silo FL setting corresponds to the case of few ($2$--$50$) reliable clients, each holding medium to large datasets, and is typically found in applications such as healthcare, finance, or industry. While previous works have proposed representative datasets for cross-device FL, few realistic healthcare cross-silo FL datasets exist, thereby slowing algorithmic research in this critical application. In this work, we propose a novel cross-silo dataset suite focused on healthcare, FLamby (Federated Learning AMple Benchmark of Your cross-silo strategies), to bridge the gap between theory and practice of cross-silo FL. FLamby encompasses 7 healthcare datasets with natural splits, covering multiple tasks, modalities, and data volumes, each accompanied with baseline training code. As an illustration, we additionally benchmark standard FL algorithms on all datasets. Our flexible and modular suite allows researchers to easily download datasets, reproduce results and re-use the different components for their research. FLamby is available at~\url{www.github.com/owkin/flamby}.
A General Theory for Federated Optimization with Asynchronous and Heterogeneous Clients Updates
Jun 21, 2022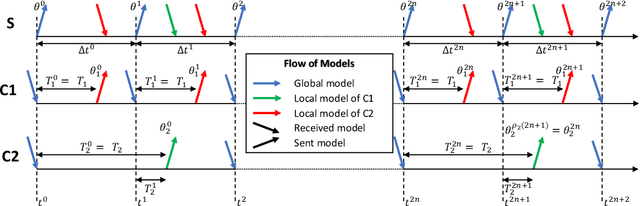
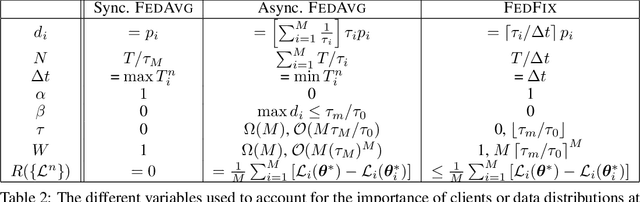


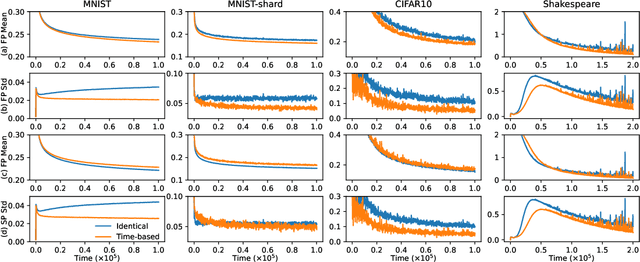
Abstract:We propose a novel framework to study asynchronous federated learning optimization with delays in gradient updates. Our theoretical framework extends the standard FedAvg aggregation scheme by introducing stochastic aggregation weights to represent the variability of the clients update time, due for example to heterogeneous hardware capabilities. Our formalism applies to the general federated setting where clients have heterogeneous datasets and perform at least one step of stochastic gradient descent (SGD). We demonstrate convergence for such a scheme and provide sufficient conditions for the related minimum to be the optimum of the federated problem. We show that our general framework applies to existing optimization schemes including centralized learning, FedAvg, asynchronous FedAvg, and FedBuff. The theory here provided allows drawing meaningful guidelines for designing a federated learning experiment in heterogeneous conditions. In particular, we develop in this work FedFix, a novel extension of FedAvg enabling efficient asynchronous federated training while preserving the convergence stability of synchronous aggregation. We empirically demonstrate our theory on a series of experiments showing that asynchronous FedAvg leads to fast convergence at the expense of stability, and we finally demonstrate the improvements of FedFix over synchronous and asynchronous FedAvg.
Privacy Preserving Image Registration
May 17, 2022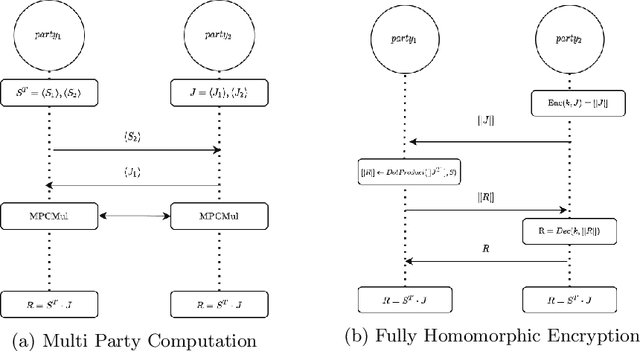
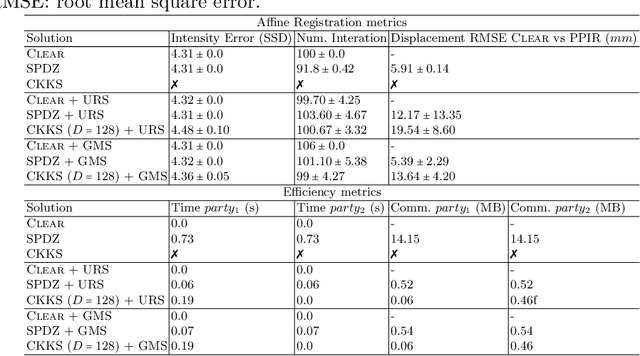
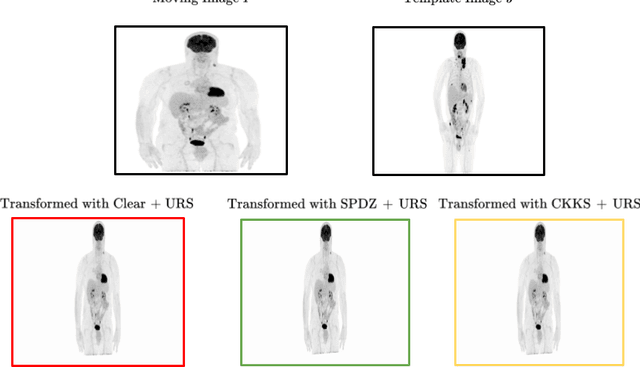
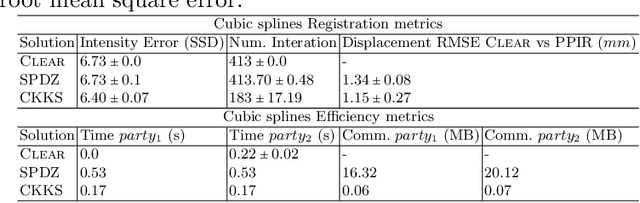
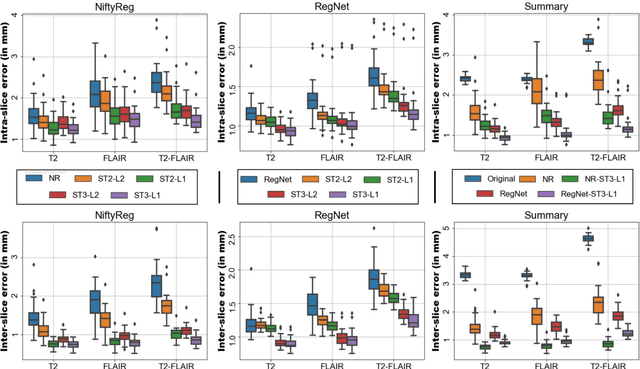
Abstract:Image registration is a key task in medical imaging applications, allowing to represent medical images in a common spatial reference frame. Current literature on image registration is generally based on the assumption that images are usually accessible to the researcher, from which the spatial transformation is subsequently estimated. This common assumption may not be met in current practical applications, since the sensitive nature of medical images may ultimately require their analysis under privacy constraints, preventing to share the image content in clear form. In this work, we formulate the problem of image registration under a privacy preserving regime, where images are assumed to be confidential and cannot be disclosed in clear. We derive our privacy preserving image registration framework by extending classical registration paradigms to account for advanced cryptographic tools, such as secure multi-party computation and homomorphic encryption, that enable the execution of operations without leaking the underlying data. To overcome the problem of performance and scalability of cryptographic tools in high dimensions, we first propose to optimize the underlying image registration operations using gradient approximations. We further revisit the use of homomorphic encryption and use a packing method to allow the encryption and multiplication of large matrices more efficiently. We demonstrate our privacy preserving framework in linear and non-linear registration problems, evaluating its accuracy and scalability with respect to standard image registration. Our results show that privacy preserving image registration is feasible and can be adopted in sensitive medical imaging applications.
A Differentially Private Probabilistic Framework for Modeling the Variability Across Federated Datasets of Heterogeneous Multi-View Observations
Apr 26, 2022
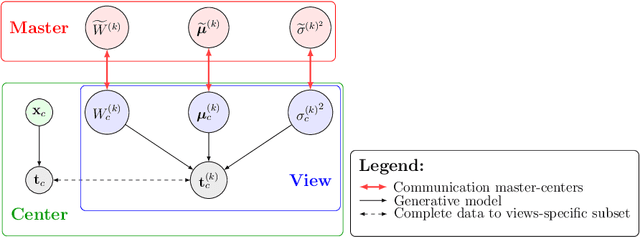
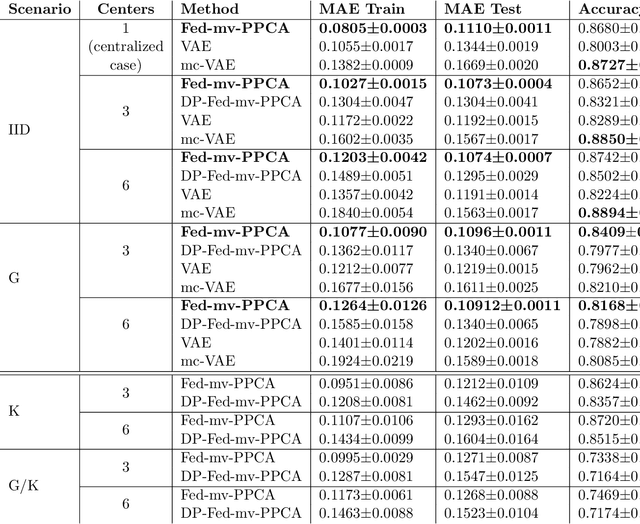

Abstract:We propose a novel federated learning paradigm to model data variability among heterogeneous clients in multi-centric studies. Our method is expressed through a hierarchical Bayesian latent variable model, where client-specific parameters are assumed to be realization from a global distribution at the master level, which is in turn estimated to account for data bias and variability across clients. We show that our framework can be effectively optimized through expectation maximization (EM) over latent master's distribution and clients' parameters. We also introduce formal differential privacy (DP) guarantees compatibly with our EM optimization scheme. We tested our method on the analysis of multi-modal medical imaging data and clinical scores from distributed clinical datasets of patients affected by Alzheimer's disease. We demonstrate that our method is robust when data is distributed either in iid and non-iid manners, even when local parameters perturbation is included to provide DP guarantees. Moreover, the variability of data, views and centers can be quantified in an interpretable manner, while guaranteeing high-quality data reconstruction as compared to state-of-the-art autoencoding models and federated learning schemes. The code is available at https://gitlab.inria.fr/epione/federated-multi-views-ppca.
* Accepted for publication at the Journal of Machine Learning for Biomedical Imaging (MELBA) https://www.melba-journal.org
On The Impact of Client Sampling on Federated Learning Convergence
Jul 26, 2021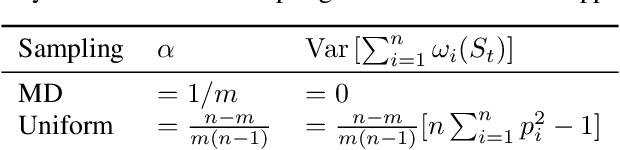
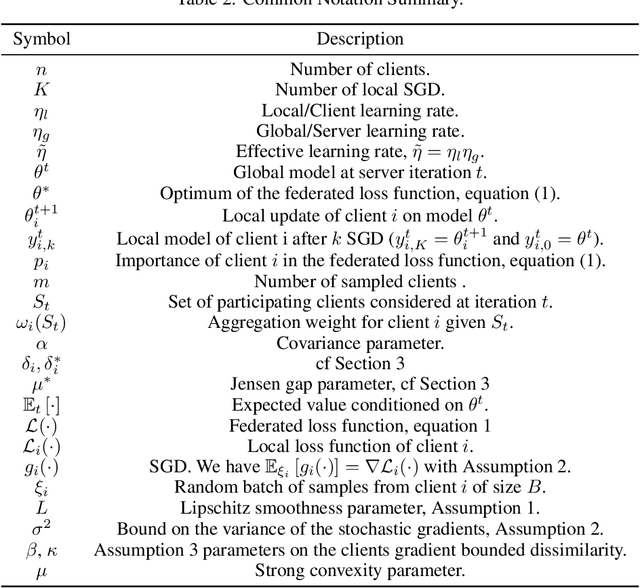
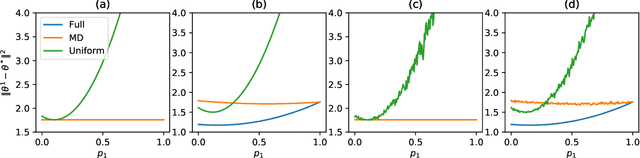
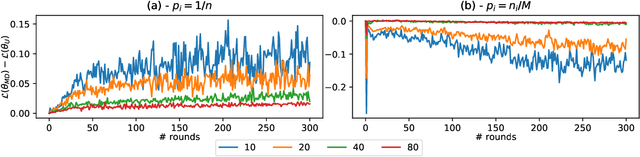

Abstract:While clients' sampling is a central operation of current state-of-the-art federated learning (FL) approaches, the impact of this procedure on the convergence and speed of FL remains to date under-investigated. In this work we introduce a novel decomposition theorem for the convergence of FL, allowing to clearly quantify the impact of client sampling on the global model update. Contrarily to previous convergence analyses, our theorem provides the exact decomposition of a given convergence step, thus enabling accurate considerations about the role of client sampling and heterogeneity. First, we provide a theoretical ground for previously reported results on the relationship between FL convergence and the variance of the aggregation weights. Second, we prove for the first time that the quality of FL convergence is also impacted by the resulting covariance between aggregation weights. Third, we establish that the sum of the aggregation weights is another source of slow-down and should be equal to 1 to improve FL convergence speed. Our theory is general, and is here applied to Multinomial Distribution (MD) and Uniform sampling, the two default client sampling in FL, and demonstrated through a series of experiments in non-iid and unbalanced scenarios. Our results suggest that MD sampling should be used as default sampling scheme, due to the resilience to the changes in data ratio during the learning process, while Uniform sampling is superior only in the special case when clients have the same amount of data.
Clustered Sampling: Low-Variance and Improved Representativity for Clients Selection in Federated Learning
May 21, 2021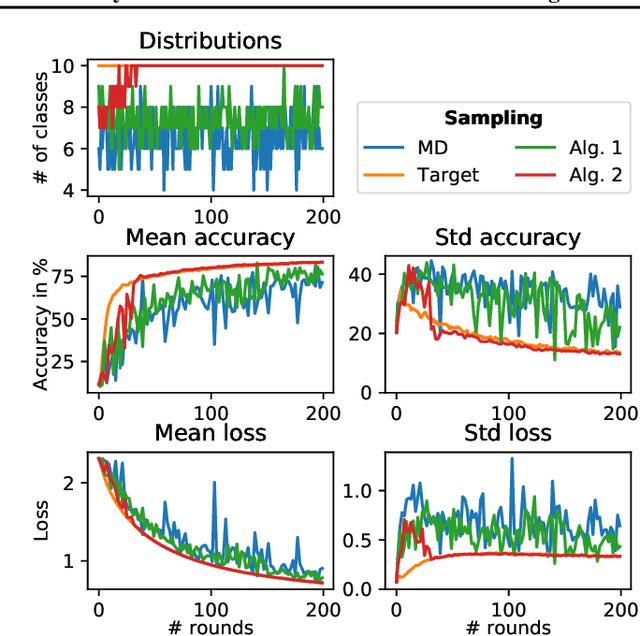
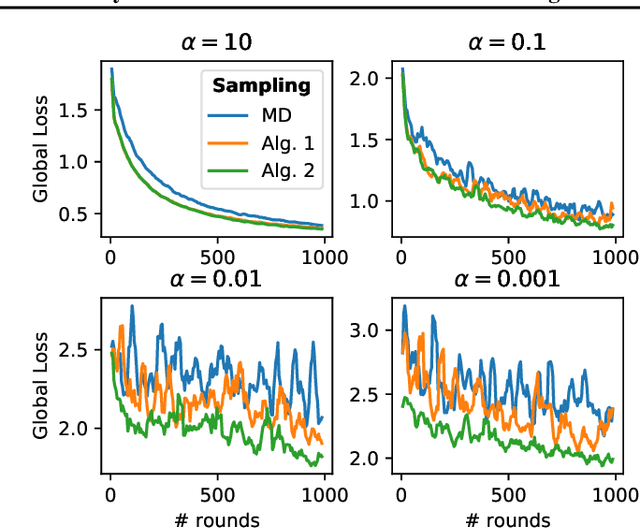
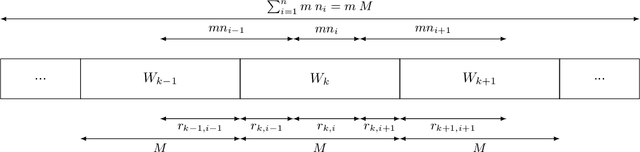
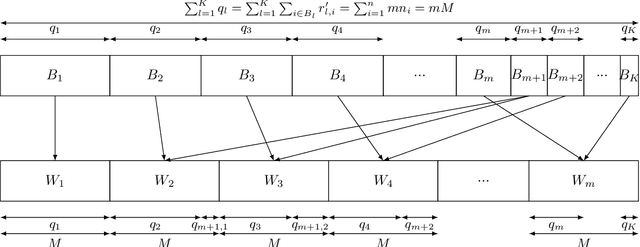
Abstract:This work addresses the problem of optimizing communications between server and clients in federated learning (FL). Current sampling approaches in FL are either biased, or non optimal in terms of server-clients communications and training stability. To overcome this issue, we introduce \textit{clustered sampling} for clients selection. We prove that clustered sampling leads to better clients representatitivity and to reduced variance of the clients stochastic aggregation weights in FL. Compatibly with our theory, we provide two different clustering approaches enabling clients aggregation based on 1) sample size, and 2) models similarity. Through a series of experiments in non-iid and unbalanced scenarios, we demonstrate that model aggregation through clustered sampling consistently leads to better training convergence and variability when compared to standard sampling approaches. Our approach does not require any additional operation on the clients side, and can be seamlessly integrated in standard FL implementations. Finally, clustered sampling is compatible with existing methods and technologies for privacy enhancement, and for communication reduction through model compression.
Robust joint registration of multiple stains and MRI for multimodal 3D histology reconstruction: Application to the Allen human brain atlas
May 04, 2021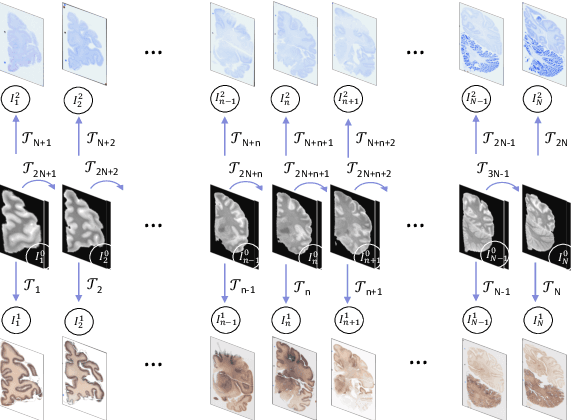
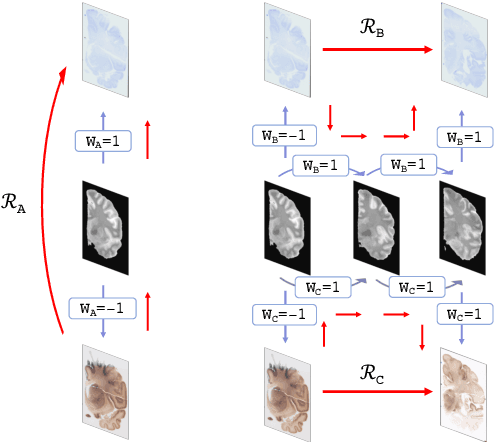
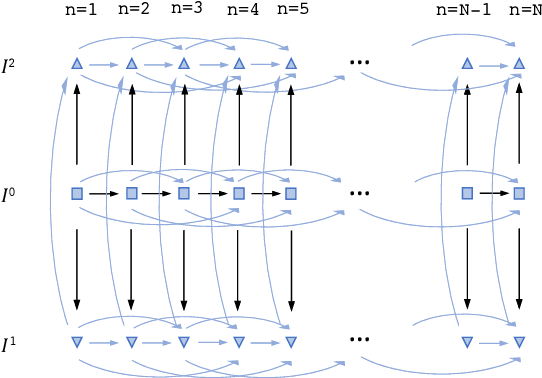
Abstract:Joint registration of a stack of 2D histological sections to recover 3D structure (3D histology reconstruction) finds application in areas such as atlas building and validation of in vivo imaging. Straighforward pairwise registration of neighbouring sections yields smooth reconstructions but has well-known problems such as banana effect (straightening of curved structures) and z-shift (drift). While these problems can be alleviated with an external, linearly aligned reference (e.g., Magnetic Resonance images), registration is often inaccurate due to contrast differences and the strong nonlinear distortion of the tissue, including artefacts such as folds and tears. In this paper, we present a probabilistic model of spatial deformation that yields reconstructions for multiple histological stains that that are jointly smooth, robust to outliers, and follow the reference shape. The model relies on a spanning tree of latent transforms connecting all the sections and slices, and assumes that the registration between any pair of images can be see as a noisy version of the composition of (possibly inverted) latent transforms connecting the two images. Bayesian inference is used to compute the most likely latent transforms given a set of pairwise registrations between image pairs within and across modalities. Results on synthetic deformations on multiple MR modalities, show that our method can accurately and robustly register multiple contrasts even in the presence of outliers. The 3D histology reconstruction of two stains (Nissl and parvalbumin) from the Allen human brain atlas, show its benefits on real data with severe distortions. We also provide the correspondence to MNI space, bridging the gap between two of the most used atlases in histology and MRI. Data is available at https://openneuro.org/datasets/ds003590 and code at https://github.com/acasamitjana/3dhirest.
Vessel-CAPTCHA: an efficient learning framework for vessel annotation and segmentation
Jan 29, 2021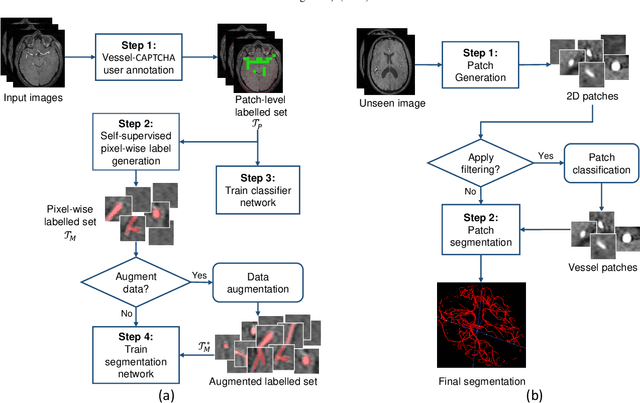
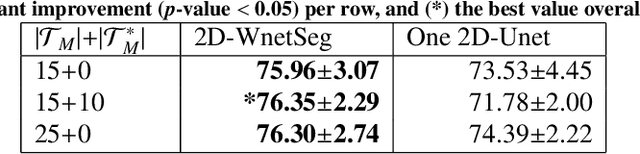
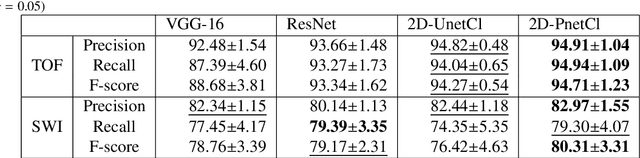
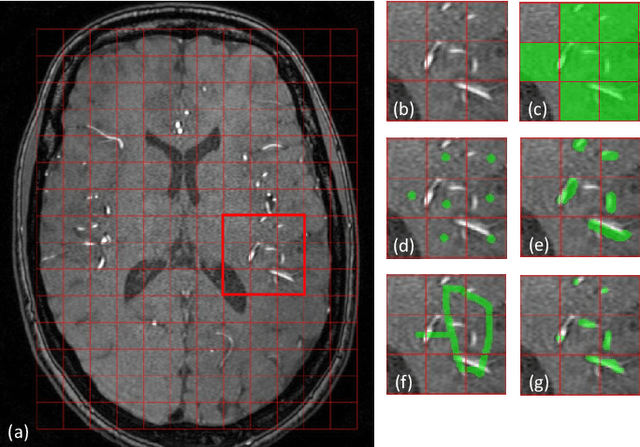

Abstract:The use of deep learning techniques for 3D brain vessel image segmentation has not been as widespread as for the segmentation of other organs and tissues. This can be explained by two factors. First, deep learning techniques tend to show poor performances at the segmentation of relatively small objects compared to the size of the full image. Second, due to the complexity of vascular trees and the small size of vessels, it is challenging to obtain the amount of annotated training data typically needed by deep learning methods. To address these problems, we propose a novel annotation-efficient deep learning vessel segmentation framework. The framework avoids pixel-wise annotations, only requiring patch-level labels to discriminate between vessel and non-vessel 2D patches in the training set, in a setup similar to the CAPTCHAs used to differentiate humans from bots in web applications. The user-provided annotations are used for two tasks: 1) to automatically generate pixel-wise labels for vessels and background in each patch, which are used to train a segmentation network, and 2) to train a classifier network. The classifier network allows to generate additional weak patch labels, further reducing the annotation burden, and it acts as a noise filter for poor quality images. We use this framework for the segmentation of the cerebrovascular tree in Time-of-Flight angiography (TOF) and Susceptibility-Weighted Images (SWI). The results show that the framework achieves state-of-the-art accuracy, while reducing the annotation time by up to 80% with respect to learning-based segmentation methods using pixel-wise labels for training
Joint data imputation and mechanistic modelling for simulating heart-brain interactions in incomplete datasets
Oct 08, 2020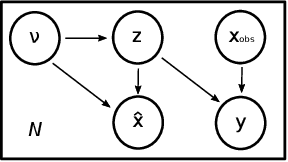
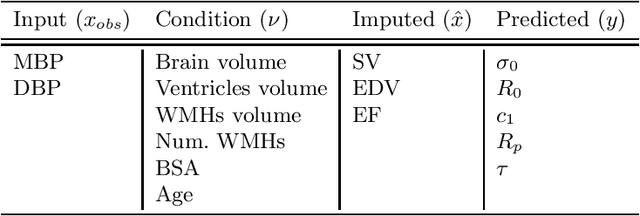
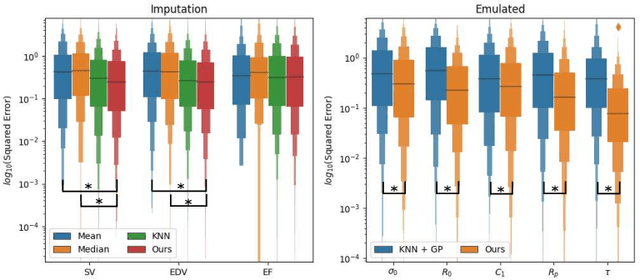
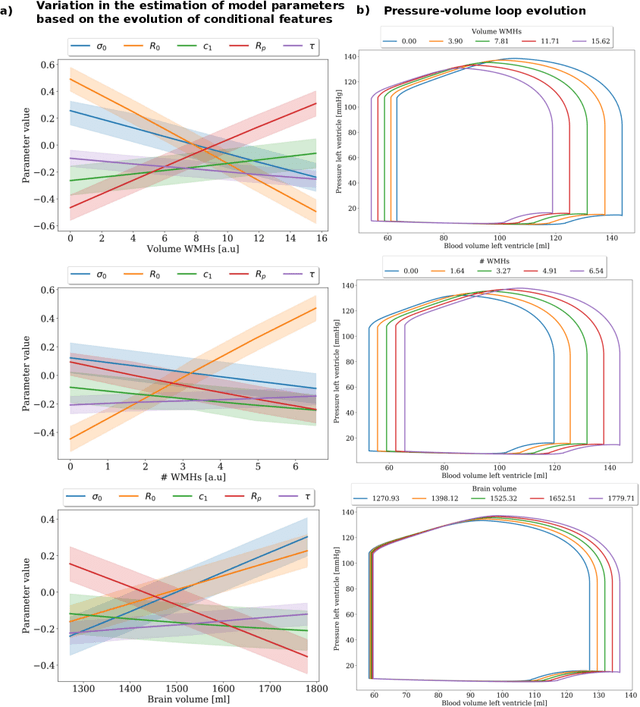
Abstract:The use of mechanistic models in clinical studies is limited by the lack of multi-modal patients data representing different anatomical and physiological processes. For example, neuroimaging datasets do not provide a sufficient representation of heart features for the modeling of cardiovascular factors in brain disorders. To tackle this problem we introduce a probabilistic framework for joint cardiac data imputation and personalisation of cardiovascular mechanistic models, with application to brain studies with incomplete heart data. Our approach is based on a variational framework for the joint inference of an imputation model of cardiac information from the available features, along with a Gaussian Process emulator that can faithfully reproduce personalised cardiovascular dynamics. Experimental results on UK Biobank show that our model allows accurate imputation of missing cardiac features in datasets containing minimal heart information, e.g. systolic and diastolic blood pressures only, while jointly estimating the emulated parameters of the lumped model. This allows a novel exploration of the heart-brain joint relationship through simulation of realistic cardiac dynamics corresponding to different conditions of brain anatomy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge