Vien Ngoc Dang
Understanding Task Aggregation for Generalizable Ultrasound Foundation Models
Mar 18, 2026Abstract:Foundation models promise to unify multiple clinical tasks within a single framework, but recent ultrasound studies report that unified models can underperform task-specific baselines. We hypothesize that this degradation arises not from model capacity limitations, but from task aggregation strategies that ignore interactions between task heterogeneity and available training data scale. In this work, we systematically analyze when heterogeneous ultrasound tasks can be jointly learned without performance loss, establishing practical criteria for task aggregation in unified clinical imaging models. We introduce M2DINO, a multi-organ, multi-task framework built on DINOv3 with task-conditioned Mixture-of-Experts blocks for adaptive capacity allocation. We systematically evaluate 27 ultrasound tasks spanning segmentation, classification, detection, and regression under three paradigms: task-specific, clinically-grouped, and all-task unified training. Our results show that aggregation effectiveness depends strongly on training data scale. While clinically-grouped training can improve performance in data-rich settings, it may induce substantial negative transfer in low-data settings. In contrast, all-task unified training exhibits more consistent performance across clinical groups. We further observe that task sensitivity varies by task type in our experiments: segmentation shows the largest performance drops compared with regression and classification. These findings provide practical guidance for ultrasound foundation models, emphasizing that aggregation strategies should jointly consider training data availability and task characteristics rather than relying on clinical taxonomy alone.
Fairness and bias correction in machine learning for depression prediction: results from four different study populations
Nov 10, 2022

Abstract:A significant level of stigma and inequality exists in mental healthcare, especially in under-served populations, which spreads through collected data. When not properly accounted for, machine learning (ML) models learned from data can reinforce the structural biases already present in society. Here, we present a systematic study of bias in ML models designed to predict depression in four different case studies covering different countries and populations. We find that standard ML approaches show regularly biased behaviors. However, we show that standard mitigation techniques, and our own post-hoc method, can be effective in reducing the level of unfair bias. We provide practical recommendations to develop ML models for depression risk prediction with increased fairness and trust in the real world. No single best ML model for depression prediction provides equality of outcomes. This emphasizes the importance of analyzing fairness during model selection and transparent reporting about the impact of debiasing interventions.
Vessel-CAPTCHA: an efficient learning framework for vessel annotation and segmentation
Jan 29, 2021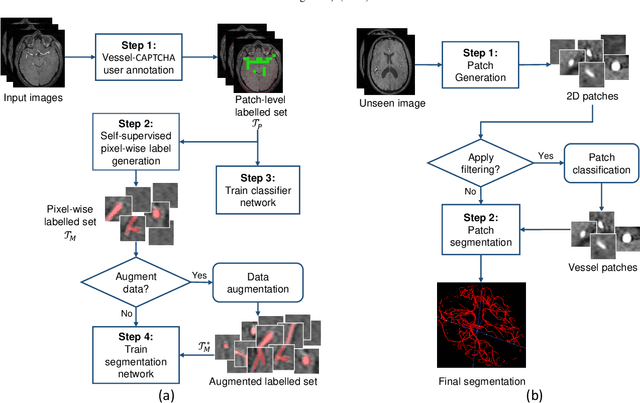
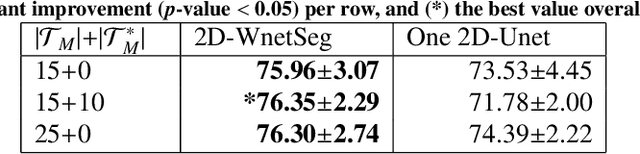
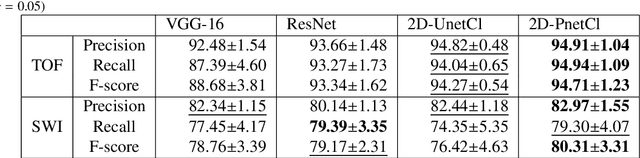
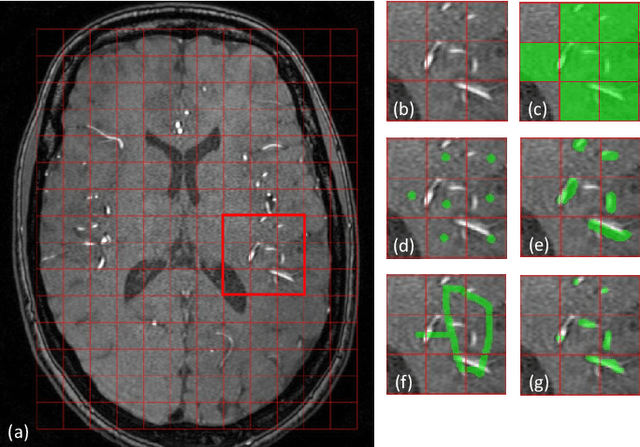

Abstract:The use of deep learning techniques for 3D brain vessel image segmentation has not been as widespread as for the segmentation of other organs and tissues. This can be explained by two factors. First, deep learning techniques tend to show poor performances at the segmentation of relatively small objects compared to the size of the full image. Second, due to the complexity of vascular trees and the small size of vessels, it is challenging to obtain the amount of annotated training data typically needed by deep learning methods. To address these problems, we propose a novel annotation-efficient deep learning vessel segmentation framework. The framework avoids pixel-wise annotations, only requiring patch-level labels to discriminate between vessel and non-vessel 2D patches in the training set, in a setup similar to the CAPTCHAs used to differentiate humans from bots in web applications. The user-provided annotations are used for two tasks: 1) to automatically generate pixel-wise labels for vessels and background in each patch, which are used to train a segmentation network, and 2) to train a classifier network. The classifier network allows to generate additional weak patch labels, further reducing the annotation burden, and it acts as a noise filter for poor quality images. We use this framework for the segmentation of the cerebrovascular tree in Time-of-Flight angiography (TOF) and Susceptibility-Weighted Images (SWI). The results show that the framework achieves state-of-the-art accuracy, while reducing the annotation time by up to 80% with respect to learning-based segmentation methods using pixel-wise labels for training
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge