Rosa Cortese
Multi-Domain Brain Vessel Segmentation Through Feature Disentanglement
Oct 02, 2025


Abstract:The intricate morphology of brain vessels poses significant challenges for automatic segmentation models, which usually focus on a single imaging modality. However, accurately treating brain-related conditions requires a comprehensive understanding of the cerebrovascular tree, regardless of the specific acquisition procedure. Our framework effectively segments brain arteries and veins in various datasets through image-to-image translation while avoiding domain-specific model design and data harmonization between the source and the target domain. This is accomplished by employing disentanglement techniques to independently manipulate different image properties, allowing them to move from one domain to another in a label-preserving manner. Specifically, we focus on manipulating vessel appearances during adaptation while preserving spatial information, such as shapes and locations, which are crucial for correct segmentation. Our evaluation effectively bridges large and varied domain gaps across medical centers, image modalities, and vessel types. Additionally, we conduct ablation studies on the optimal number of required annotations and other architectural choices. The results highlight our framework's robustness and versatility, demonstrating the potential of domain adaptation methodologies to perform cerebrovascular image segmentation in multiple scenarios accurately. Our code is available at https://github.com/i-vesseg/MultiVesSeg.
* 19 pages, 7 figures, 3 tables. Joint first authors: Francesco Galati and Daniele Falcetta. Accepted for publication at the Journal of Machine Learning for Biomedical Imaging (MELBA) https://melba-journal.org/2025:021. Code available at https://github.com/i-vesseg/MultiVesSeg
A2V: A Semi-Supervised Domain Adaptation Framework for Brain Vessel Segmentation via Two-Phase Training Angiography-to-Venography Translation
Sep 12, 2023



Abstract:We present a semi-supervised domain adaptation framework for brain vessel segmentation from different image modalities. Existing state-of-the-art methods focus on a single modality, despite the wide range of available cerebrovascular imaging techniques. This can lead to significant distribution shifts that negatively impact the generalization across modalities. By relying on annotated angiographies and a limited number of annotated venographies, our framework accomplishes image-to-image translation and semantic segmentation, leveraging a disentangled and semantically rich latent space to represent heterogeneous data and perform image-level adaptation from source to target domains. Moreover, we reduce the typical complexity of cycle-based architectures and minimize the use of adversarial training, which allows us to build an efficient and intuitive model with stable training. We evaluate our method on magnetic resonance angiographies and venographies. While achieving state-of-the-art performance in the source domain, our method attains a Dice score coefficient in the target domain that is only 8.9% lower, highlighting its promising potential for robust cerebrovascular image segmentation across different modalities.
Vessel-CAPTCHA: an efficient learning framework for vessel annotation and segmentation
Jan 29, 2021
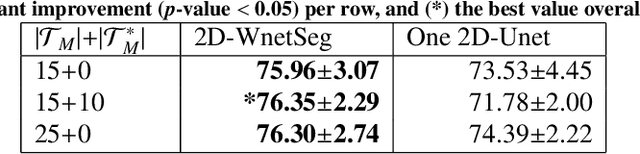
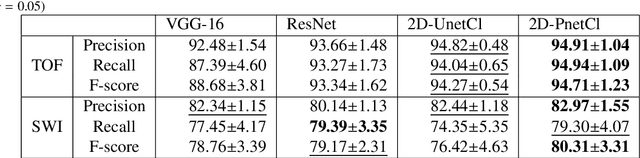
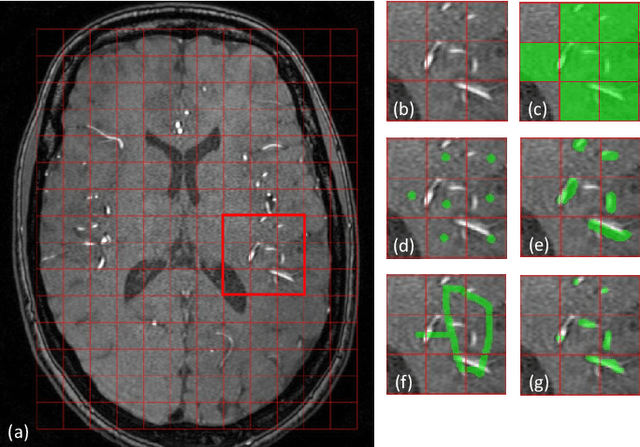

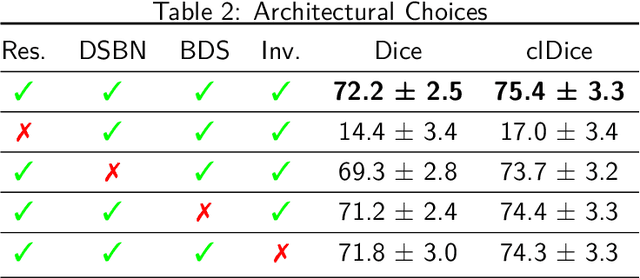
Abstract:The use of deep learning techniques for 3D brain vessel image segmentation has not been as widespread as for the segmentation of other organs and tissues. This can be explained by two factors. First, deep learning techniques tend to show poor performances at the segmentation of relatively small objects compared to the size of the full image. Second, due to the complexity of vascular trees and the small size of vessels, it is challenging to obtain the amount of annotated training data typically needed by deep learning methods. To address these problems, we propose a novel annotation-efficient deep learning vessel segmentation framework. The framework avoids pixel-wise annotations, only requiring patch-level labels to discriminate between vessel and non-vessel 2D patches in the training set, in a setup similar to the CAPTCHAs used to differentiate humans from bots in web applications. The user-provided annotations are used for two tasks: 1) to automatically generate pixel-wise labels for vessels and background in each patch, which are used to train a segmentation network, and 2) to train a classifier network. The classifier network allows to generate additional weak patch labels, further reducing the annotation burden, and it acts as a noise filter for poor quality images. We use this framework for the segmentation of the cerebrovascular tree in Time-of-Flight angiography (TOF) and Susceptibility-Weighted Images (SWI). The results show that the framework achieves state-of-the-art accuracy, while reducing the annotation time by up to 80% with respect to learning-based segmentation methods using pixel-wise labels for training
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge