Jaume Banus
Towards Brain MRI Foundation Models for the Clinic: Findings from the FOMO25 Challenge
Apr 13, 2026Abstract:Clinical deployment of automated brain MRI analysis faces a fundamental challenge: clinical data is heterogeneous and noisy, and high-quality labels are prohibitively costly to obtain. Self-supervised learning (SSL) can address this by leveraging the vast amounts of unlabeled data produced in clinical workflows to train robust \textit{foundation models} that adapt out-of-domain with minimal supervision. However, the development of foundation models for brain MRI has been limited by small pretraining datasets and in-domain benchmarking focused on high-quality, research-grade data. To address this gap, we organized the FOMO25 challenge as a satellite event at MICCAI 2025. FOMO25 provided participants with a large pretraining dataset, FOMO60K, and evaluated models on data sourced directly from clinical workflows in few-shot and out-of-domain settings. Tasks covered infarct classification, meningioma segmentation, and brain age regression, and considered both models trained on FOMO60K (method track) and any data (open track). Nineteen foundation models from sixteen teams were evaluated using a standardized containerized pipeline. Results show that (a) self-supervised pretraining improves generalization on clinical data under domain shift, with the strongest models trained \textit{out-of-domain} surpassing supervised baselines trained \textit{in-domain}. (b) No single pretraining objective benefits all tasks: MAE favors segmentation, hybrid reconstruction-contrastive objectives favor classification, and (c) strong performance was achieved by small pretrained models, and improvements from scaling model size and training duration did not yield reliable benefits.
From 100,000+ images to winning the first brain MRI foundation model challenges: Sharing lessons and models
Jan 19, 2026Abstract:Developing Foundation Models for medical image analysis is essential to overcome the unique challenges of radiological tasks. The first challenges of this kind for 3D brain MRI, SSL3D and FOMO25, were held at MICCAI 2025. Our solution ranked first in tracks of both contests. It relies on a U-Net CNN architecture combined with strategies leveraging anatomical priors and neuroimaging domain knowledge. Notably, our models trained 1-2 orders of magnitude faster and were 10 times smaller than competing transformer-based approaches. Models are available here: https://github.com/jbanusco/BrainFM4Challenges.
Causal Attribution of Model Performance Gaps in Medical Imaging Under Distribution Shifts
Dec 09, 2025Abstract:Deep learning models for medical image segmentation suffer significant performance drops due to distribution shifts, but the causal mechanisms behind these drops remain poorly understood. We extend causal attribution frameworks to high-dimensional segmentation tasks, quantifying how acquisition protocols and annotation variability independently contribute to performance degradation. We model the data-generating process through a causal graph and employ Shapley values to fairly attribute performance changes to individual mechanisms. Our framework addresses unique challenges in medical imaging: high-dimensional outputs, limited samples, and complex mechanism interactions. Validation on multiple sclerosis (MS) lesion segmentation across 4 centers and 7 annotators reveals context-dependent failure modes: annotation protocol shifts dominate when crossing annotators (7.4% $\pm$ 8.9% DSC attribution), while acquisition shifts dominate when crossing imaging centers (6.5% $\pm$ 9.1%). This mechanism-specific quantification enables practitioners to prioritize targeted interventions based on deployment context.
Spatiotemporal graph neural process for reconstruction, extrapolation, and classification of cardiac trajectories
Sep 16, 2025



Abstract:We present a probabilistic framework for modeling structured spatiotemporal dynamics from sparse observations, focusing on cardiac motion. Our approach integrates neural ordinary differential equations (NODEs), graph neural networks (GNNs), and neural processes into a unified model that captures uncertainty, temporal continuity, and anatomical structure. We represent dynamic systems as spatiotemporal multiplex graphs and model their latent trajectories using a GNN-parameterized vector field. Given the sparse context observations at node and edge levels, the model infers a distribution over latent initial states and control variables, enabling both interpolation and extrapolation of trajectories. We validate the method on three synthetic dynamical systems (coupled pendulum, Lorenz attractor, and Kuramoto oscillators) and two real-world cardiac imaging datasets - ACDC (N=150) and UK Biobank (N=526) - demonstrating accurate reconstruction, extrapolation, and disease classification capabilities. The model accurately reconstructs trajectories and extrapolates future cardiac cycles from a single observed cycle. It achieves state-of-the-art results on the ACDC classification task (up to 99% accuracy), and detects atrial fibrillation in UK Biobank subjects with competitive performance (up to 67% accuracy). This work introduces a flexible approach for analyzing cardiac motion and offers a foundation for graph-based learning in structured biomedical spatiotemporal time-series data.
Joint data imputation and mechanistic modelling for simulating heart-brain interactions in incomplete datasets
Oct 08, 2020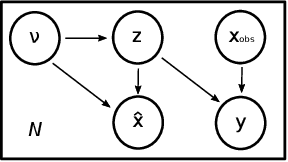
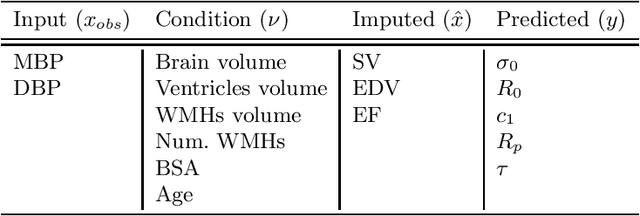
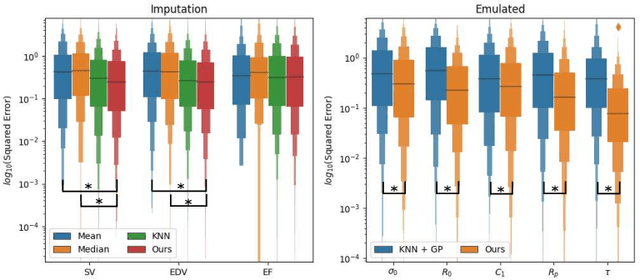
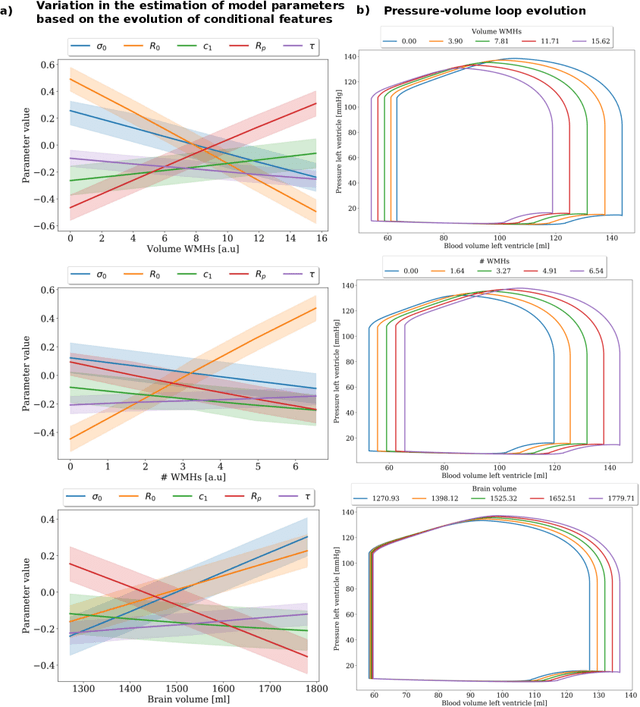
Abstract:The use of mechanistic models in clinical studies is limited by the lack of multi-modal patients data representing different anatomical and physiological processes. For example, neuroimaging datasets do not provide a sufficient representation of heart features for the modeling of cardiovascular factors in brain disorders. To tackle this problem we introduce a probabilistic framework for joint cardiac data imputation and personalisation of cardiovascular mechanistic models, with application to brain studies with incomplete heart data. Our approach is based on a variational framework for the joint inference of an imputation model of cardiac information from the available features, along with a Gaussian Process emulator that can faithfully reproduce personalised cardiovascular dynamics. Experimental results on UK Biobank show that our model allows accurate imputation of missing cardiac features in datasets containing minimal heart information, e.g. systolic and diastolic blood pressures only, while jointly estimating the emulated parameters of the lumped model. This allows a novel exploration of the heart-brain joint relationship through simulation of realistic cardiac dynamics corresponding to different conditions of brain anatomy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge