Lei Xing
Solving Inverse Problems in Medical Imaging with Score-Based Generative Models
Nov 15, 2021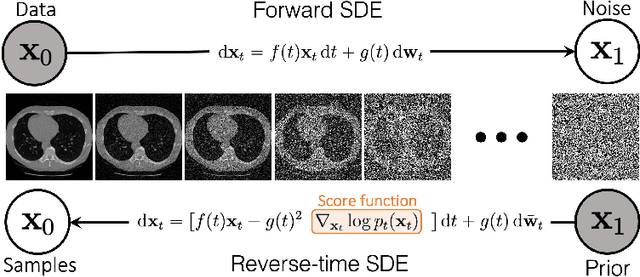
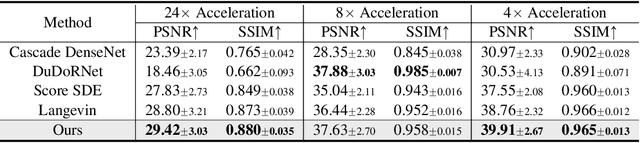

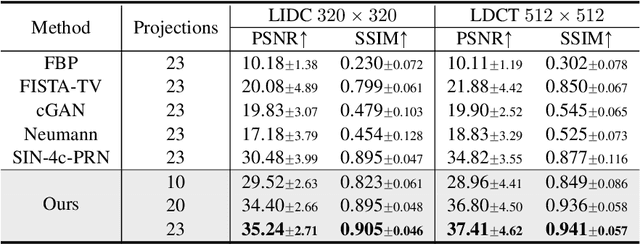
Abstract:Reconstructing medical images from partial measurements is an important inverse problem in Computed Tomography (CT) and Magnetic Resonance Imaging (MRI). Existing solutions based on machine learning typically train a model to directly map measurements to medical images, leveraging a training dataset of paired images and measurements. These measurements are typically synthesized from images using a fixed physical model of the measurement process, which hinders the generalization capability of models to unknown measurement processes. To address this issue, we propose a fully unsupervised technique for inverse problem solving, leveraging the recently introduced score-based generative models. Specifically, we first train a score-based generative model on medical images to capture their prior distribution. Given measurements and a physical model of the measurement process at test time, we introduce a sampling method to reconstruct an image consistent with both the prior and the observed measurements. Our method does not assume a fixed measurement process during training, and can thus be flexibly adapted to different measurement processes at test time. Empirically, we observe comparable or better performance to supervised learning techniques in several medical imaging tasks in CT and MRI, while demonstrating significantly better generalization to unknown measurement processes.
MHFC: Multi-Head Feature Collaboration for Few-Shot Learning
Oct 10, 2021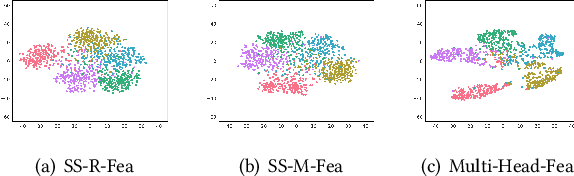
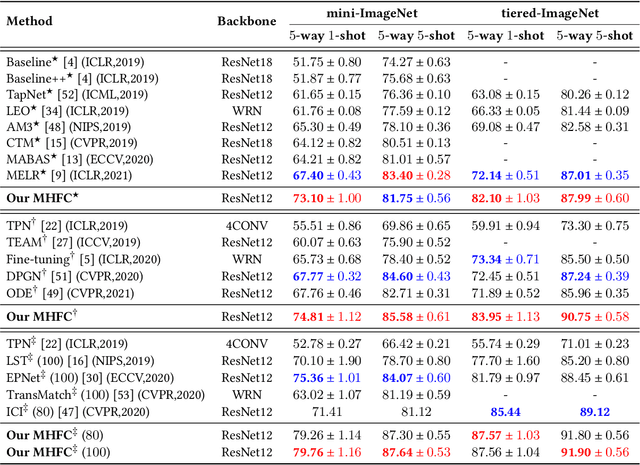
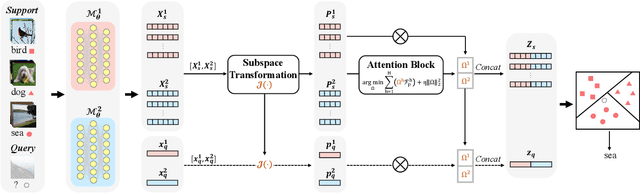
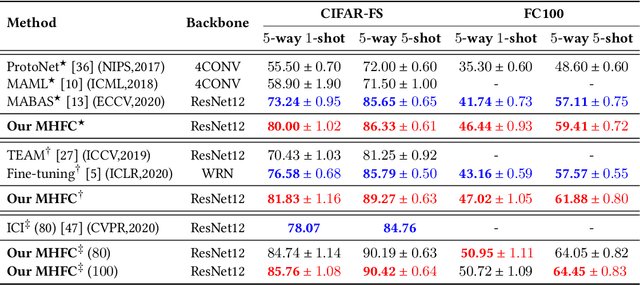
Abstract:Few-shot learning (FSL) aims to address the data-scarce problem. A standard FSL framework is composed of two components: (1) Pre-train. Employ the base data to generate a CNN-based feature extraction model (FEM). (2) Meta-test. Apply the trained FEM to acquire the novel data's features and recognize them. FSL relies heavily on the design of the FEM. However, various FEMs have distinct emphases. For example, several may focus more attention on the contour information, whereas others may lay particular emphasis on the texture information. The single-head feature is only a one-sided representation of the sample. Besides the negative influence of cross-domain (e.g., the trained FEM can not adapt to the novel class flawlessly), the distribution of novel data may have a certain degree of deviation compared with the ground truth distribution, which is dubbed as distribution-shift-problem (DSP). To address the DSP, we propose Multi-Head Feature Collaboration (MHFC) algorithm, which attempts to project the multi-head features (e.g., multiple features extracted from a variety of FEMs) to a unified space and fuse them to capture more discriminative information. Typically, first, we introduce a subspace learning method to transform the multi-head features to aligned low-dimensional representations. It corrects the DSP via learning the feature with more powerful discrimination and overcomes the problem of inconsistent measurement scales from different head features. Then, we design an attention block to update combination weights for each head feature automatically. It comprehensively considers the contribution of various perspectives and further improves the discrimination of features. We evaluate the proposed method on five benchmark datasets (including cross-domain experiments) and achieve significant improvements of 2.1%-7.8% compared with state-of-the-arts.
Metal Artifact Reduction in 2D CT Images with Self-supervised Cross-domain Learning
Sep 28, 2021Abstract:The presence of metallic implants often introduces severe metal artifacts in the X-ray CT images, which could adversely influence clinical diagnosis or dose calculation in radiation therapy. In this work, we present a novel deep-learning-based approach for metal artifact reduction (MAR). In order to alleviate the need for anatomically identical CT image pairs (i.e., metal artifact-corrupted CT image and metal artifact-free CT image) for network learning, we propose a self-supervised cross-domain learning framework. Specifically, we train a neural network to restore the metal trace region values in the given metal-free sinogram, where the metal trace is identified by the forward projection of metal masks. We then design a novel FBP reconstruction loss to encourage the network to generate more perfect completion results and a residual-learning-based image refinement module to reduce the secondary artifacts in the reconstructed CT images. To preserve the fine structure details and fidelity of the final MAR image, instead of directly adopting CNN-refined images as output, we incorporate the metal trace replacement into our framework and replace the metal-affected projections of the original sinogram with the prior sinogram generated by the forward projection of the CNN output. We then use the filtered backward projection (FBP) algorithms for final MAR image reconstruction. We conduct an extensive evaluation on simulated and real artifact data to show the effectiveness of our design. Our method produces superior MAR results and outperforms other compelling methods. We also demonstrate the potential of our framework for other organ sites.
CIM: Class-Irrelevant Mapping for Few-Shot Classification
Sep 07, 2021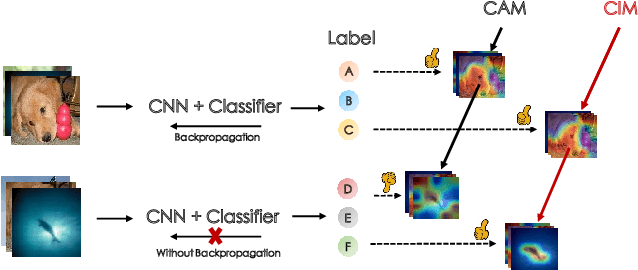
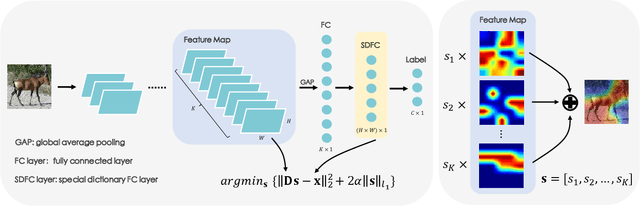
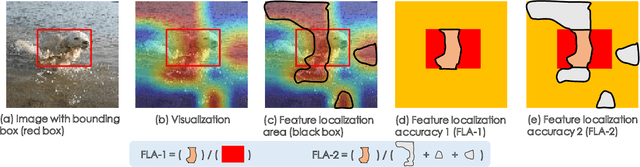
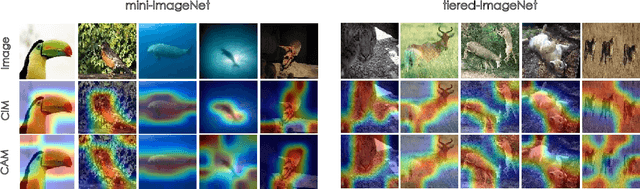
Abstract:Few-shot classification (FSC) is one of the most concerned hot issues in recent years. The general setting consists of two phases: (1) Pre-train a feature extraction model (FEM) with base data (has large amounts of labeled samples). (2) Use the FEM to extract the features of novel data (with few labeled samples and totally different categories from base data), then classify them with the to-be-designed classifier. The adaptability of pre-trained FEM to novel data determines the accuracy of novel features, thereby affecting the final classification performances. To this end, how to appraise the pre-trained FEM is the most crucial focus in the FSC community. It sounds like traditional Class Activate Mapping (CAM) based methods can achieve this by overlaying weighted feature maps. However, due to the particularity of FSC (e.g., there is no backpropagation when using the pre-trained FEM to extract novel features), we cannot activate the feature map with the novel classes. To address this challenge, we propose a simple, flexible method, dubbed as Class-Irrelevant Mapping (CIM). Specifically, first, we introduce dictionary learning theory and view the channels of the feature map as the bases in a dictionary. Then we utilize the feature map to fit the feature vector of an image to achieve the corresponding channel weights. Finally, we overlap the weighted feature map for visualization to appraise the ability of pre-trained FEM on novel data. For fair use of CIM in evaluating different models, we propose a new measurement index, called Feature Localization Accuracy (FLA). In experiments, we first compare our CIM with CAM in regular tasks and achieve outstanding performances. Next, we use our CIM to appraise several classical FSC frameworks without considering the classification results and discuss them.
NeRP: Implicit Neural Representation Learning with Prior Embedding for Sparsely Sampled Image Reconstruction
Aug 24, 2021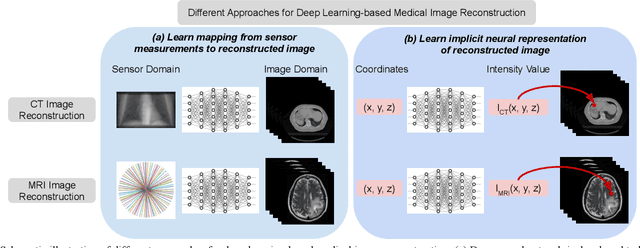
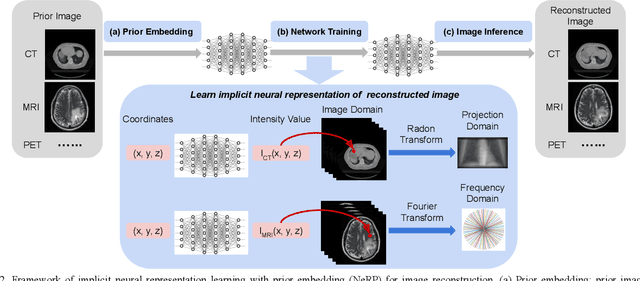
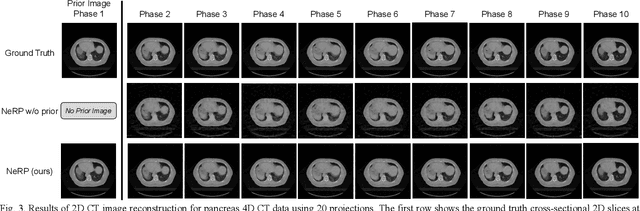
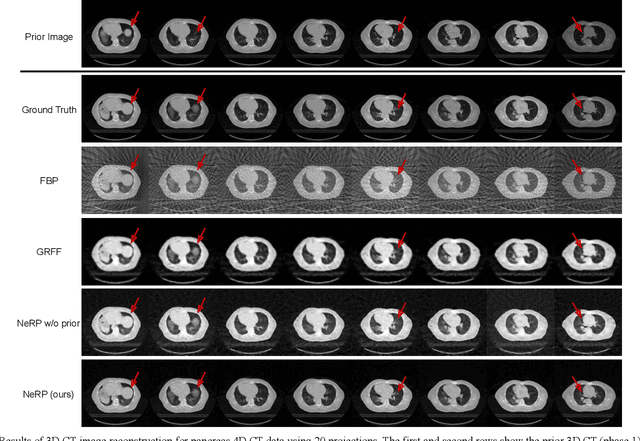
Abstract:Image reconstruction is an inverse problem that solves for a computational image based on sampled sensor measurement. Sparsely sampled image reconstruction poses addition challenges due to limited measurements. In this work, we propose an implicit Neural Representation learning methodology with Prior embedding (NeRP) to reconstruct a computational image from sparsely sampled measurements. The method differs fundamentally from previous deep learning-based image reconstruction approaches in that NeRP exploits the internal information in an image prior, and the physics of the sparsely sampled measurements to produce a representation of the unknown subject. No large-scale data is required to train the NeRP except for a prior image and sparsely sampled measurements. In addition, we demonstrate that NeRP is a general methodology that generalizes to different imaging modalities such as CT and MRI. We also show that NeRP can robustly capture the subtle yet significant image changes required for assessing tumor progression.
A Geometry-Informed Deep Learning Framework for Ultra-Sparse 3D Tomographic Image Reconstruction
May 25, 2021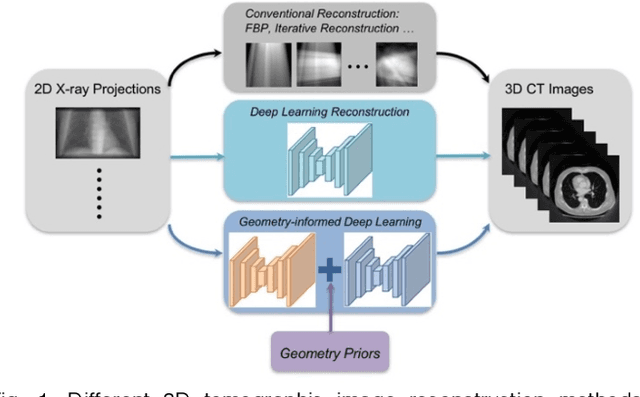
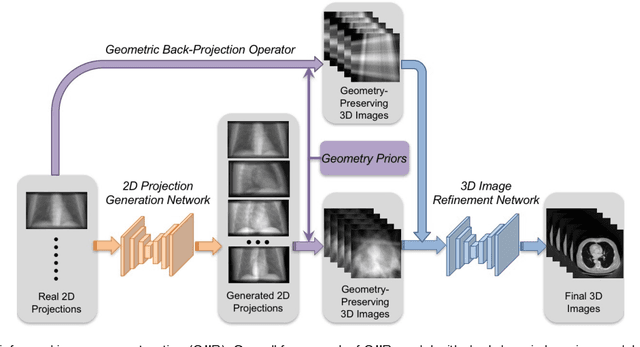
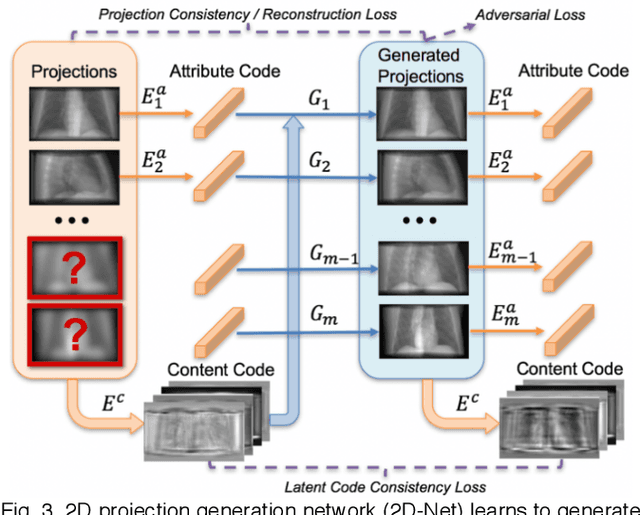
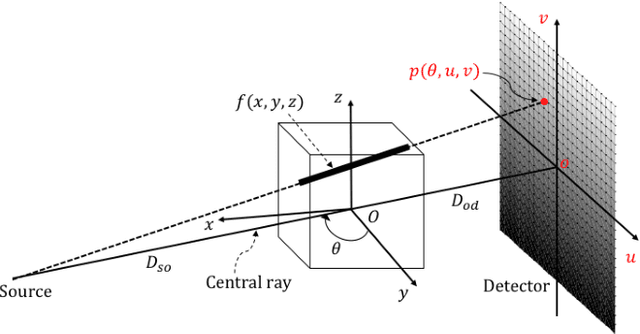
Abstract:Deep learning affords enormous opportunities to augment the armamentarium of biomedical imaging, albeit its design and implementation have potential flaws. Fundamentally, most deep learning models are driven entirely by data without consideration of any prior knowledge, which dramatically increases the complexity of neural networks and limits the application scope and model generalizability. Here we establish a geometry-informed deep learning framework for ultra-sparse 3D tomographic image reconstruction. We introduce a novel mechanism for integrating geometric priors of the imaging system. We demonstrate that the seamless inclusion of known priors is essential to enhance the performance of 3D volumetric computed tomography imaging with ultra-sparse sampling. The study opens new avenues for data-driven biomedical imaging and promises to provide substantially improved imaging tools for various clinical imaging and image-guided interventions.
DualNorm-UNet: Incorporating Global and Local Statistics for Robust Medical Image Segmentation
Mar 29, 2021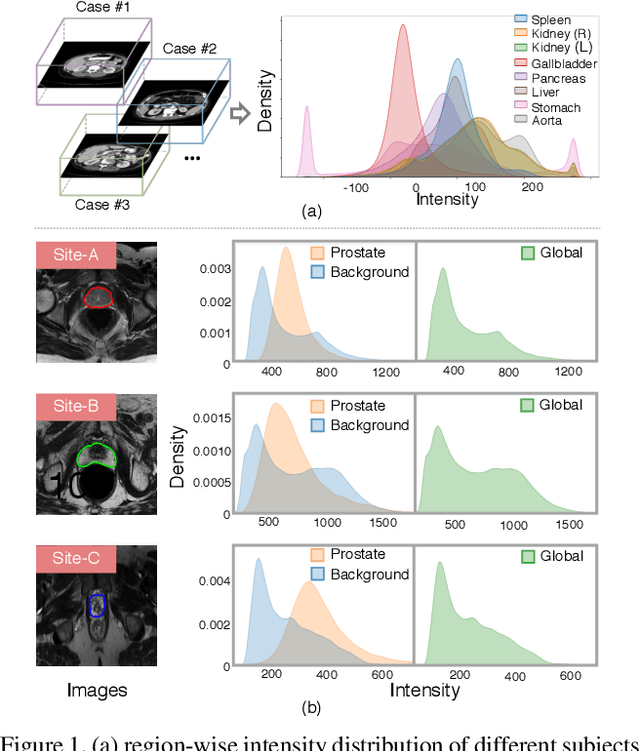
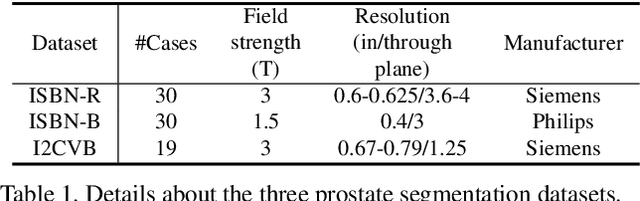
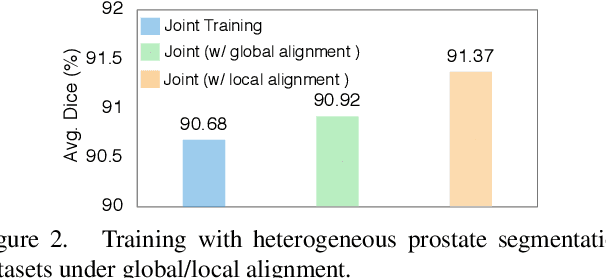
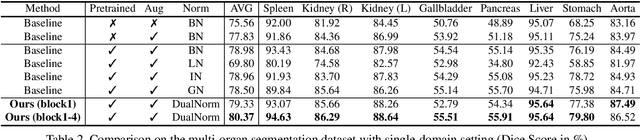
Abstract:Batch Normalization (BN) is one of the key components for accelerating network training, and has been widely adopted in the medical image analysis field. However, BN only calculates the global statistics at the batch level, and applies the same affine transformation uniformly across all spatial coordinates, which would suppress the image contrast of different semantic structures. In this paper, we propose to incorporate the semantic class information into normalization layers, so that the activations corresponding to different regions (i.e., classes) can be modulated differently. We thus develop a novel DualNorm-UNet, to concurrently incorporate both global image-level statistics and local region-wise statistics for network normalization. Specifically, the local statistics are integrated by adaptively modulating the activations along different class regions via the learned semantic masks in the normalization layer. Compared with existing methods, our approach exploits semantic knowledge at normalization and yields more discriminative features for robust segmentation results. More importantly, our network demonstrates superior abilities in capturing domain-invariant information from multiple domains (institutions) of medical data. Extensive experiments show that our proposed DualNorm-UNet consistently improves the performance on various segmentation tasks, even in the face of more complex and variable data distributions. Code is available at https://github.com/lambert-x/DualNorm-Unet.
TransCT: Dual-path Transformer for Low Dose Computed Tomography
Mar 03, 2021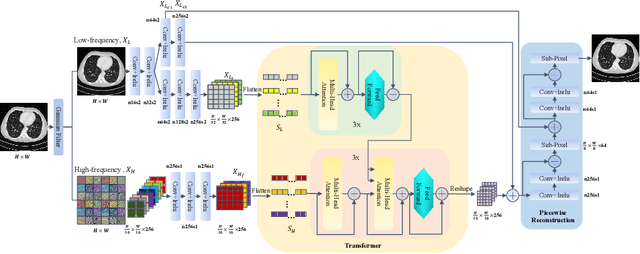

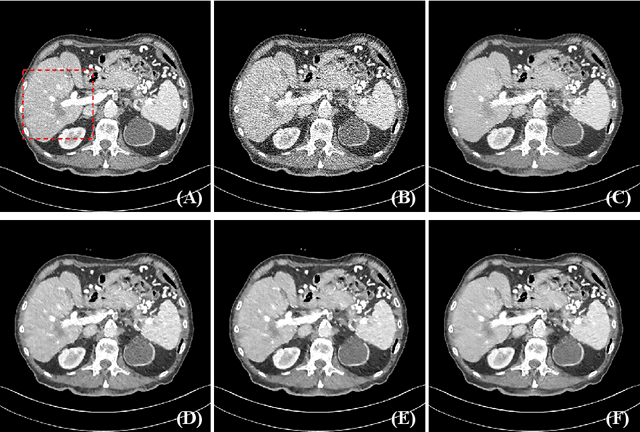
Abstract:Low dose computed tomography (LDCT) has attracted more and more attention in routine clinical diagnosis assessment, therapy planning, etc., which can reduce the dose of X-ray radiation to patients. However, the noise caused by low X-ray exposure degrades the CT image quality and then affects clinical diagnosis accuracy. In this paper, we train a transformer-based neural network to enhance the final CT image quality. To be specific, we first decompose the noisy LDCT image into two parts: high-frequency (HF) and low-frequency (LF) compositions. Then, we extract content features (X_{L_c}) and latent texture features (X_{L_t}) from the LF part, as well as HF embeddings (X_{H_f}) from the HF part. Further, we feed X_{L_t} and X_{H_f} into a modified transformer with three encoders and decoders to obtain well-refined HF texture features. After that, we combine these well-refined HF texture features with the pre-extracted X_{L_c} to encourage the restoration of high-quality LDCT images with the assistance of piecewise reconstruction. Extensive experiments on Mayo LDCT dataset show that our method produces superior results and outperforms other methods.
Region-specific Dictionary Learning-based Low-dose Thoracic CT Reconstruction
Oct 20, 2020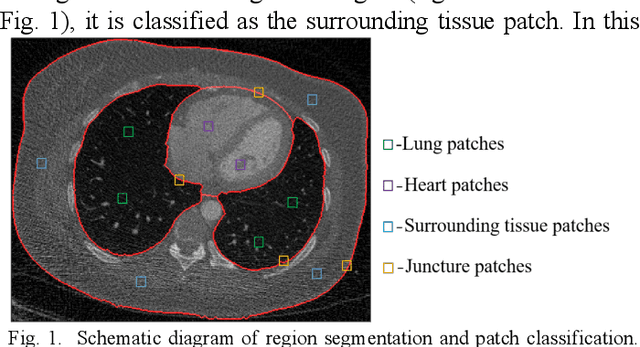
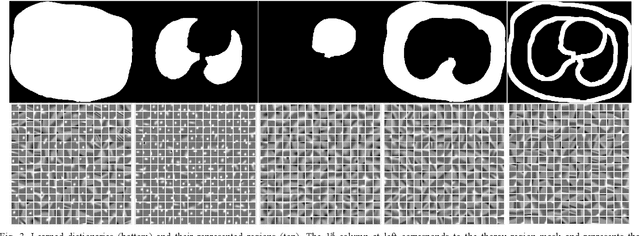
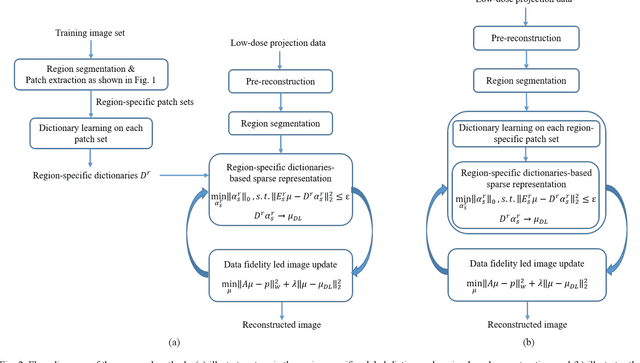

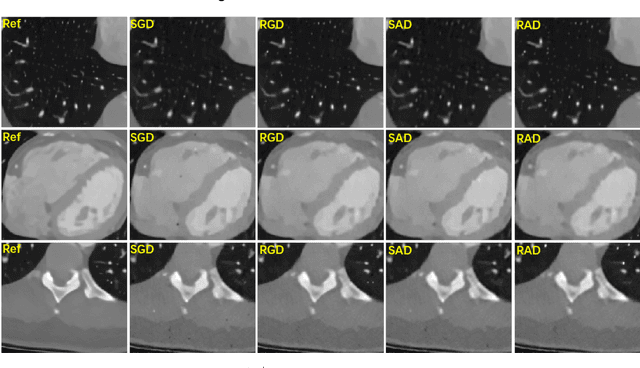
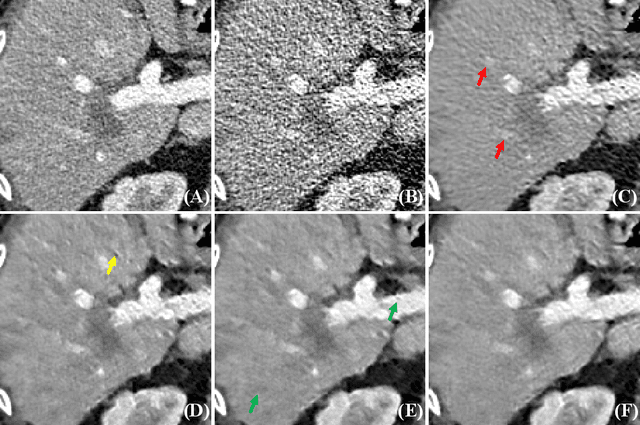
Abstract:This paper presents a dictionary learning-based method with region-specific image patches to maximize the utility of the powerful sparse data processing technique for CT image reconstruction. Considering heterogeneous distributions of image features and noise in CT, region-specific customization of dictionaries is utilized in iterative reconstruction. Thoracic CT images are partitioned into several regions according to their structural and noise characteristics. Dictionaries specific to each region are then learned from the segmented thoracic CT images and applied to subsequent image reconstruction of the region. Parameters for dictionary learning and sparse representation are determined according to the structural and noise properties of each region. The proposed method results in better performance than the conventional reconstruction based on a single dictionary in recovering structures and suppressing noise in both simulation and human CT imaging. Quantitatively, the simulation study shows maximum improvement of image quality for the whole thorax can achieve 4.88% and 11.1% in terms of the Structure-SIMilarity (SSIM) and Root-Mean-Square Error (RMSE) indices, respectively. For human imaging data, it is found that the structures in the lungs and heart can be better recovered, while simultaneously decreasing noise around the vertebra effectively. The proposed strategy takes into account inherent regional differences inside of the reconstructed object and leads to improved images. The method can be readily extended to CT imaging of other anatomical regions and other applications.
Deep Sinogram Completion with Image Prior for Metal Artifact Reduction in CT Images
Sep 16, 2020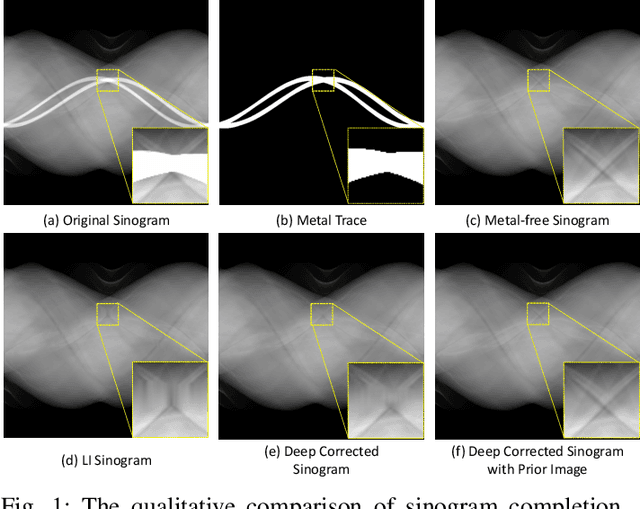
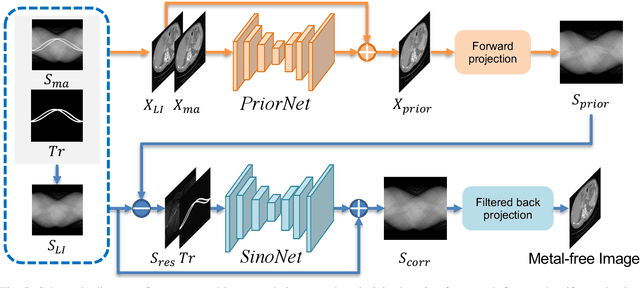
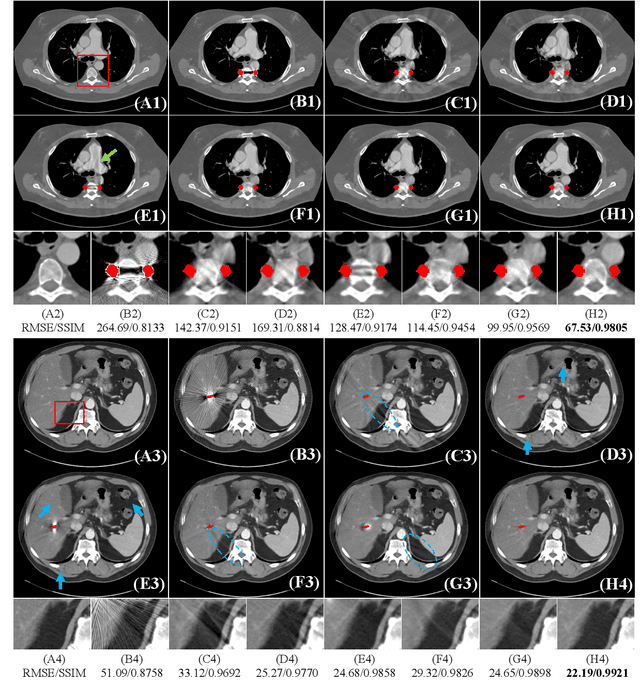
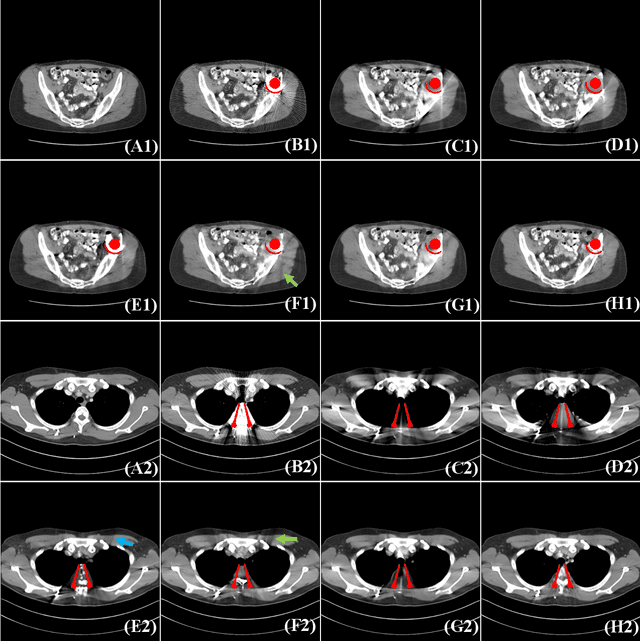
Abstract:Computed tomography (CT) has been widely used for medical diagnosis, assessment, and therapy planning and guidance. In reality, CT images may be affected adversely in the presence of metallic objects, which could lead to severe metal artifacts and influence clinical diagnosis or dose calculation in radiation therapy. In this paper, we propose a generalizable framework for metal artifact reduction (MAR) by simultaneously leveraging the advantages of image domain and sinogram domain-based MAR techniques. We formulate our framework as a sinogram completion problem and train a neural network (SinoNet) to restore the metal-affected projections. To improve the continuity of the completed projections at the boundary of metal trace and thus alleviate new artifacts in the reconstructed CT images, we train another neural network (PriorNet) to generate a good prior image to guide sinogram learning, and further design a novel residual sinogram learning strategy to effectively utilize the prior image information for better sinogram completion. The two networks are jointly trained in an end-to-end fashion with a differentiable forward projection (FP) operation so that the prior image generation and deep sinogram completion procedures can benefit from each other. Finally, the artifact-reduced CT images are reconstructed using the filtered backward projection (FBP) from the completed sinogram. Extensive experiments on simulated and real artifacts data demonstrate that our method produces superior artifact-reduced results while preserving the anatomical structures and outperforms other MAR methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge