Kai Ma
School of Electrical Engineering, Yanshan University, Qinhuangdao, China
Revisiting Experience Replay: Continual Learning by Adaptively Tuning Task-wise Relationship
Jan 06, 2022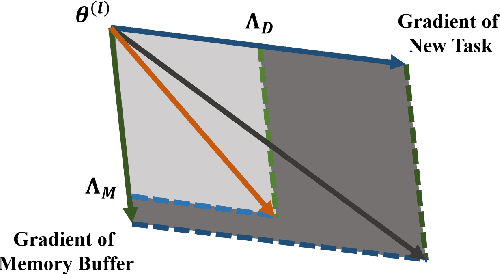
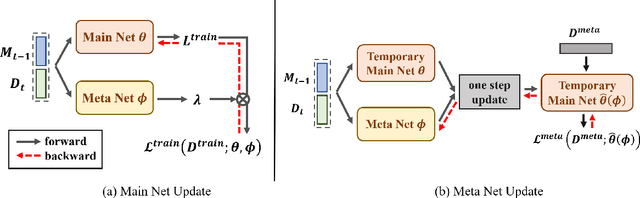
Abstract:Continual learning requires models to learn new tasks while maintaining previously learned knowledge. Various algorithms have been proposed to address this real challenge. Till now, rehearsal-based methods, such as experience replay, have achieved state-of-the-art performance. These approaches save a small part of the data of the past tasks as a memory buffer to prevent models from forgetting previously learned knowledge. However, most of them treat every new task equally, i.e., fixed the hyperparameters of the framework while learning different new tasks. Such a setting lacks the consideration of the relationship/similarity between past and new tasks. For example, the previous knowledge/features learned from dogs are more beneficial for the identification of cats (new task), compared to those learned from buses. In this regard, we propose a meta learning algorithm based on bi-level optimization to adaptively tune the relationship between the knowledge extracted from the past and new tasks. Therefore, the model can find an appropriate direction of gradient during continual learning and avoid the serious overfitting problem on memory buffer. Extensive experiments are conducted on three publicly available datasets (i.e., CIFAR-10, CIFAR-100, and Tiny ImageNet). The experimental results demonstrate that the proposed method can consistently improve the performance of all baselines.
Alleviating Noisy-label Effects in Image Classification via Probability Transition Matrix
Oct 19, 2021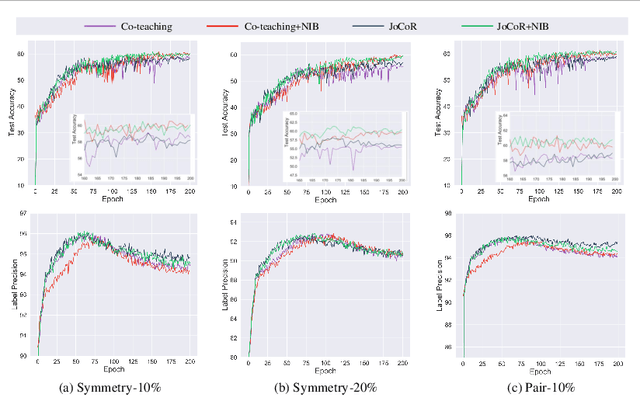
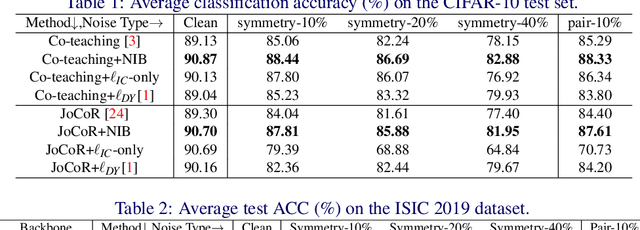
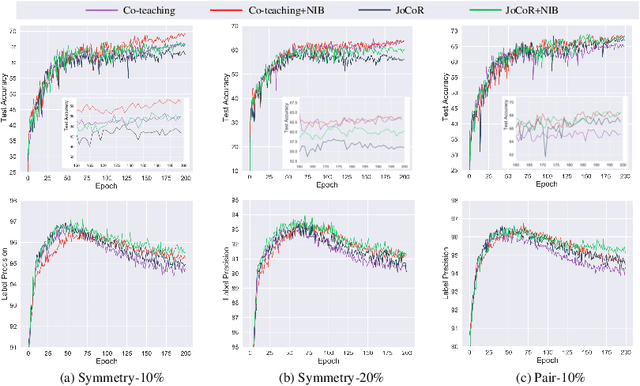
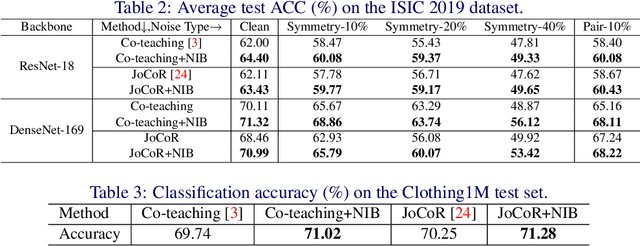
Abstract:Deep-learning-based image classification frameworks often suffer from the noisy label problem caused by the inter-observer variation. Recent studies employed learning-to-learn paradigms (e.g., Co-teaching and JoCoR) to filter the samples with noisy labels from the training set. However, most of them use a simple cross-entropy loss as the criterion for noisy label identification. The hard samples, which are beneficial for classifier learning, are often mistakenly treated as noises in such a setting since both the hard samples and ones with noisy labels lead to a relatively larger loss value than the easy cases. In this paper, we propose a plugin module, namely noise ignoring block (NIB), consisting of a probability transition matrix and an inter-class correlation (IC) loss, to separate the hard samples from the mislabeled ones, and further boost the accuracy of image classification network trained with noisy labels. Concretely, our IC loss is calculated as Kullback-Leibler divergence between the network prediction and the accumulative soft label generated by the probability transition matrix. Such that, with the lower value of IC loss, the hard cases can be easily distinguished from mislabeled cases. Extensive experiments are conducted on natural and medical image datasets (CIFAR-10 and ISIC 2019). The experimental results show that our NIB module consistently improves the performances of the state-of-the-art robust training methods.
A Unified Framework for Generalized Low-Shot Medical Image Segmentation with Scarce Data
Oct 18, 2021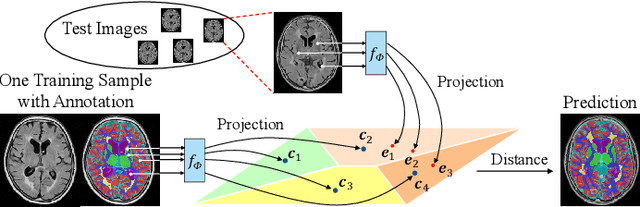
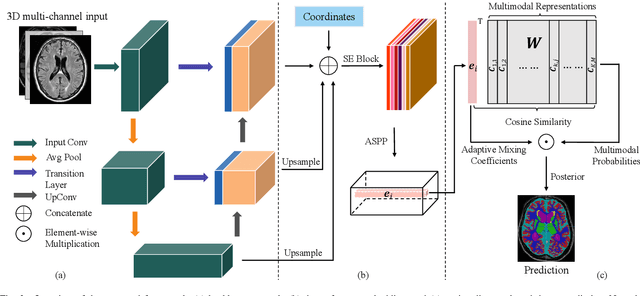
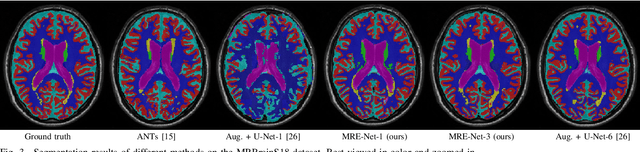
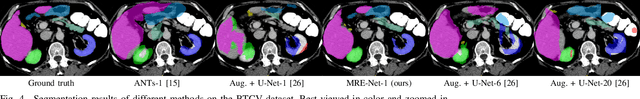
Abstract:Medical image segmentation has achieved remarkable advancements using deep neural networks (DNNs). However, DNNs often need big amounts of data and annotations for training, both of which can be difficult and costly to obtain. In this work, we propose a unified framework for generalized low-shot (one- and few-shot) medical image segmentation based on distance metric learning (DML). Unlike most existing methods which only deal with the lack of annotations while assuming abundance of data, our framework works with extreme scarcity of both, which is ideal for rare diseases. Via DML, the framework learns a multimodal mixture representation for each category, and performs dense predictions based on cosine distances between the pixels' deep embeddings and the category representations. The multimodal representations effectively utilize the inter-subject similarities and intraclass variations to overcome overfitting due to extremely limited data. In addition, we propose adaptive mixing coefficients for the multimodal mixture distributions to adaptively emphasize the modes better suited to the current input. The representations are implicitly embedded as weights of the fc layer, such that the cosine distances can be computed efficiently via forward propagation. In our experiments on brain MRI and abdominal CT datasets, the proposed framework achieves superior performances for low-shot segmentation towards standard DNN-based (3D U-Net) and classical registration-based (ANTs) methods, e.g., achieving mean Dice coefficients of 81%/69% for brain tissue/abdominal multiorgan segmentation using a single training sample, as compared to 52%/31% and 72%/35% by the U-Net and ANTs, respectively.
Unsupervised Representation Learning Meets Pseudo-Label Supervised Self-Distillation: A New Approach to Rare Disease Classification
Oct 09, 2021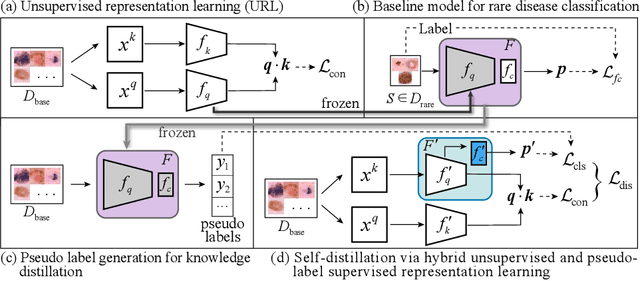
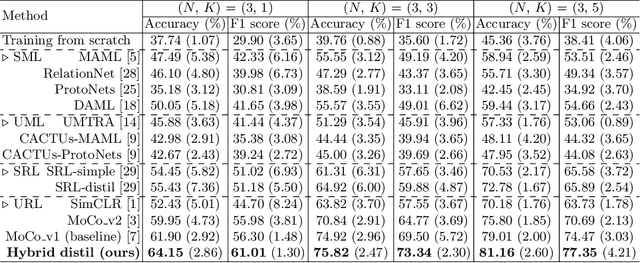
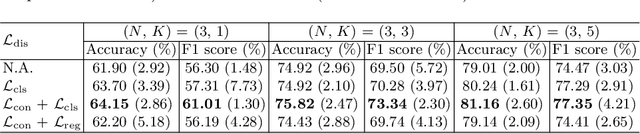
Abstract:Rare diseases are characterized by low prevalence and are often chronically debilitating or life-threatening. Imaging-based classification of rare diseases is challenging due to the severe shortage in training examples. Few-shot learning (FSL) methods tackle this challenge by extracting generalizable prior knowledge from a large base dataset of common diseases and normal controls, and transferring the knowledge to rare diseases. Yet, most existing methods require the base dataset to be labeled and do not make full use of the precious examples of the rare diseases. To this end, we propose in this work a novel hybrid approach to rare disease classification, featuring two key novelties targeted at the above drawbacks. First, we adopt the unsupervised representation learning (URL) based on self-supervising contrastive loss, whereby to eliminate the overhead in labeling the base dataset. Second, we integrate the URL with pseudo-label supervised classification for effective self-distillation of the knowledge about the rare diseases, composing a hybrid approach taking advantages of both unsupervised and (pseudo-) supervised learning on the base dataset. Experimental results on classification of rare skin lesions show that our hybrid approach substantially outperforms existing FSL methods (including those using fully supervised base dataset) for rare disease classification via effective integration of the URL and pseudo-label driven self-distillation, thus establishing a new state of the art.
All-Around Real Label Supervision: Cyclic Prototype Consistency Learning for Semi-supervised Medical Image Segmentation
Sep 28, 2021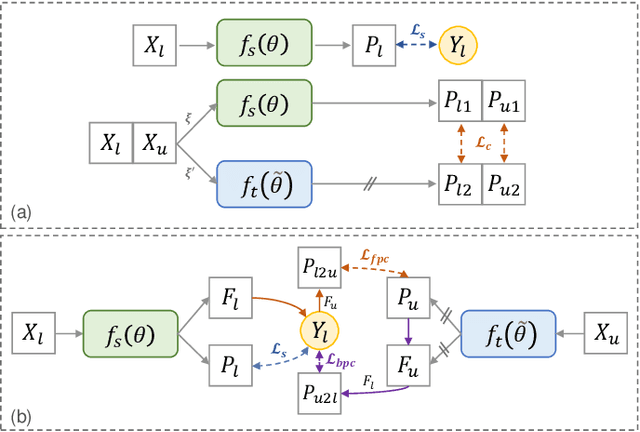
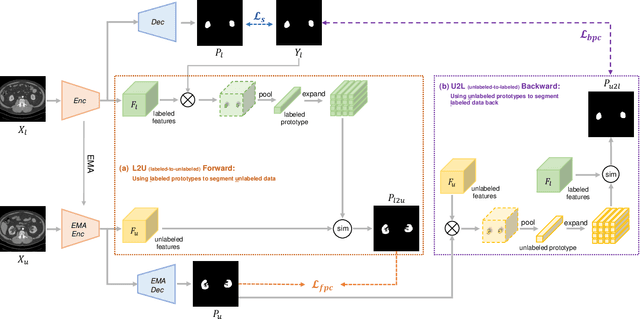
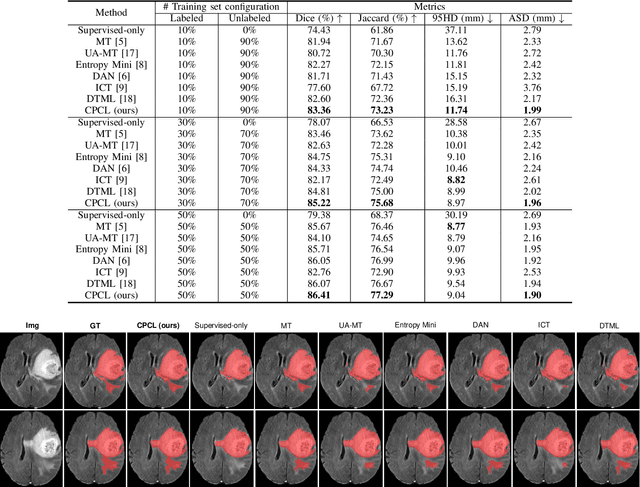
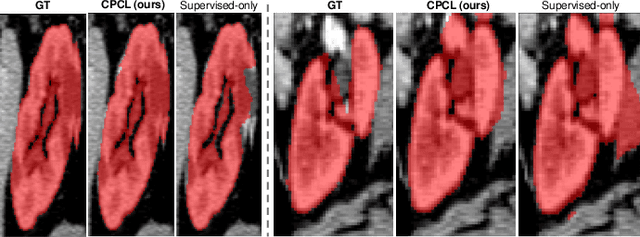
Abstract:Semi-supervised learning has substantially advanced medical image segmentation since it alleviates the heavy burden of acquiring the costly expert-examined annotations. Especially, the consistency-based approaches have attracted more attention for their superior performance, wherein the real labels are only utilized to supervise their paired images via supervised loss while the unlabeled images are exploited by enforcing the perturbation-based \textit{"unsupervised"} consistency without explicit guidance from those real labels. However, intuitively, the expert-examined real labels contain more reliable supervision signals. Observing this, we ask an unexplored but interesting question: can we exploit the unlabeled data via explicit real label supervision for semi-supervised training? To this end, we discard the previous perturbation-based consistency but absorb the essence of non-parametric prototype learning. Based on the prototypical network, we then propose a novel cyclic prototype consistency learning (CPCL) framework, which is constructed by a labeled-to-unlabeled (L2U) prototypical forward process and an unlabeled-to-labeled (U2L) backward process. Such two processes synergistically enhance the segmentation network by encouraging more discriminative and compact features. In this way, our framework turns previous \textit{"unsupervised"} consistency into new \textit{"supervised"} consistency, obtaining the \textit{"all-around real label supervision"} property of our method. Extensive experiments on brain tumor segmentation from MRI and kidney segmentation from CT images show that our CPCL can effectively exploit the unlabeled data and outperform other state-of-the-art semi-supervised medical image segmentation methods.
Training Automatic View Planner for Cardiac MR Imaging via Self-Supervision by Spatial Relationship between Views
Sep 24, 2021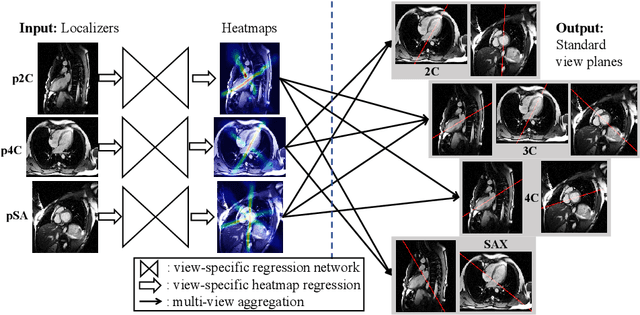
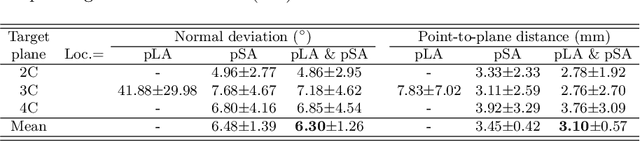
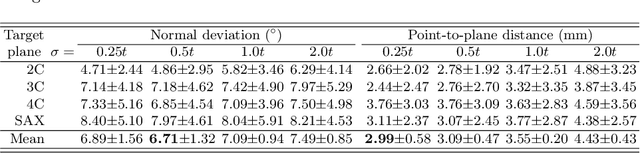
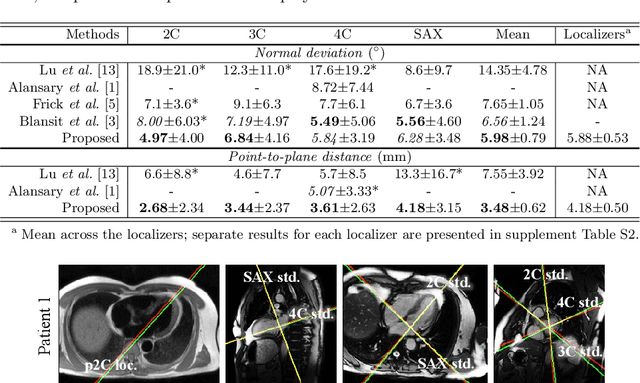
Abstract:View planning for the acquisition of cardiac magnetic resonance imaging (CMR) requires acquaintance with the cardiac anatomy and remains a challenging task in clinical practice. Existing approaches to its automation relied either on an additional volumetric image not typically acquired in clinic routine, or on laborious manual annotations of cardiac structural landmarks. This work presents a clinic-compatible and annotation-free system for automatic CMR view planning. The system mines the spatial relationship -- more specifically, locates and exploits the intersecting lines -- between the source and target views, and trains deep networks to regress heatmaps defined by these intersecting lines. As the spatial relationship is self-contained in properly stored data, e.g., in the DICOM format, the need for manual annotation is eliminated. Then, a multi-view planning strategy is proposed to aggregate information from the predicted heatmaps for all the source views of a target view, for a globally optimal prescription. The multi-view aggregation mimics the similar strategy practiced by skilled human prescribers. Experimental results on 181 clinical CMR exams show that our system achieves superior accuracy to existing approaches including conventional atlas-based and newer deep learning based ones, in prescribing four standard CMR views. The mean angle difference and point-to-plane distance evaluated against the ground truth planes are 5.98 degrees and 3.48 mm, respectively.
InDuDoNet: An Interpretable Dual Domain Network for CT Metal Artifact Reduction
Sep 11, 2021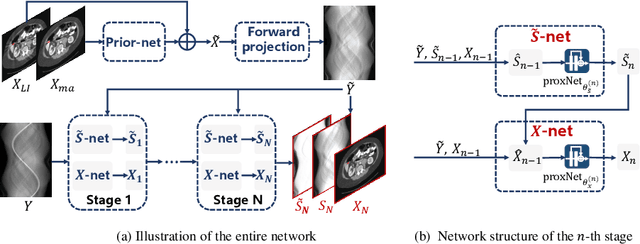
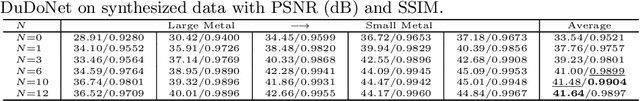
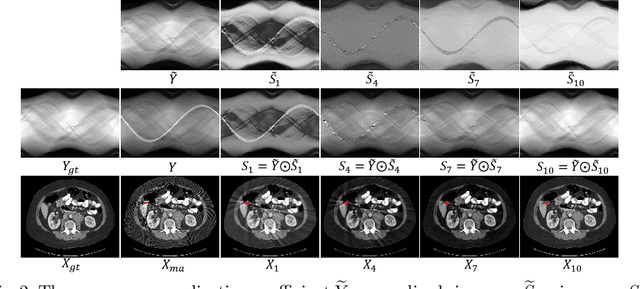
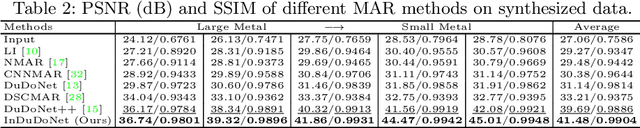
Abstract:For the task of metal artifact reduction (MAR), although deep learning (DL)-based methods have achieved promising performances, most of them suffer from two problems: 1) the CT imaging geometry constraint is not fully embedded into the network during training, leaving room for further performance improvement; 2) the model interpretability is lack of sufficient consideration. Against these issues, we propose a novel interpretable dual domain network, termed as InDuDoNet, which combines the advantages of model-driven and data-driven methodologies. Specifically, we build a joint spatial and Radon domain reconstruction model and utilize the proximal gradient technique to design an iterative algorithm for solving it. The optimization algorithm only consists of simple computational operators, which facilitate us to correspondingly unfold iterative steps into network modules and thus improve the interpretablility of the framework. Extensive experiments on synthesized and clinical data show the superiority of our InDuDoNet. Code is available in \url{https://github.com/hongwang01/InDuDoNet}.%method on the tasks of MAR and downstream multi-class pelvic fracture segmentation.
Multi-Anchor Active Domain Adaptation for Semantic Segmentation
Aug 18, 2021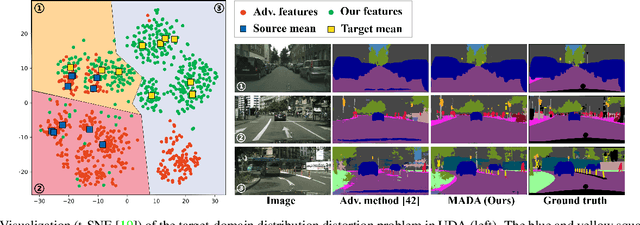
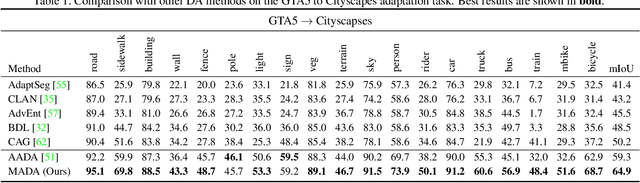
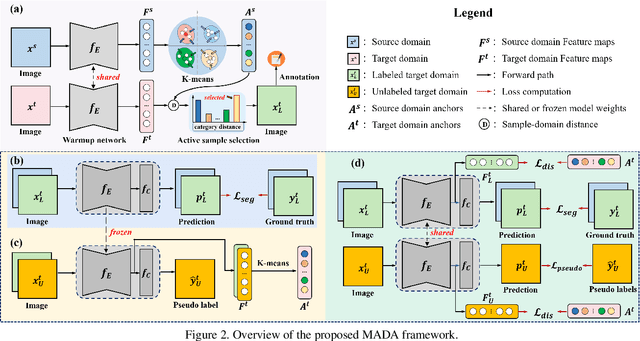
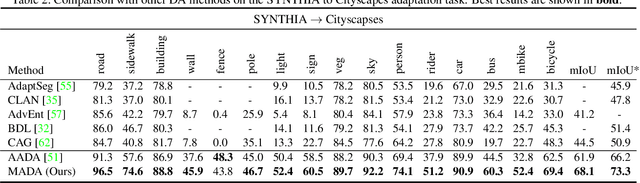
Abstract:Unsupervised domain adaption has proven to be an effective approach for alleviating the intensive workload of manual annotation by aligning the synthetic source-domain data and the real-world target-domain samples. Unfortunately, mapping the target-domain distribution to the source-domain unconditionally may distort the essential structural information of the target-domain data. To this end, we firstly propose to introduce a novel multi-anchor based active learning strategy to assist domain adaptation regarding the semantic segmentation task. By innovatively adopting multiple anchors instead of a single centroid, the source domain can be better characterized as a multimodal distribution, thus more representative and complimentary samples are selected from the target domain. With little workload to manually annotate these active samples, the distortion of the target-domain distribution can be effectively alleviated, resulting in a large performance gain. The multi-anchor strategy is additionally employed to model the target-distribution. By regularizing the latent representation of the target samples compact around multiple anchors through a novel soft alignment loss, more precise segmentation can be achieved. Extensive experiments are conducted on public datasets to demonstrate that the proposed approach outperforms state-of-the-art methods significantly, along with thorough ablation study to verify the effectiveness of each component.
A New Bidirectional Unsupervised Domain Adaptation Segmentation Framework
Aug 18, 2021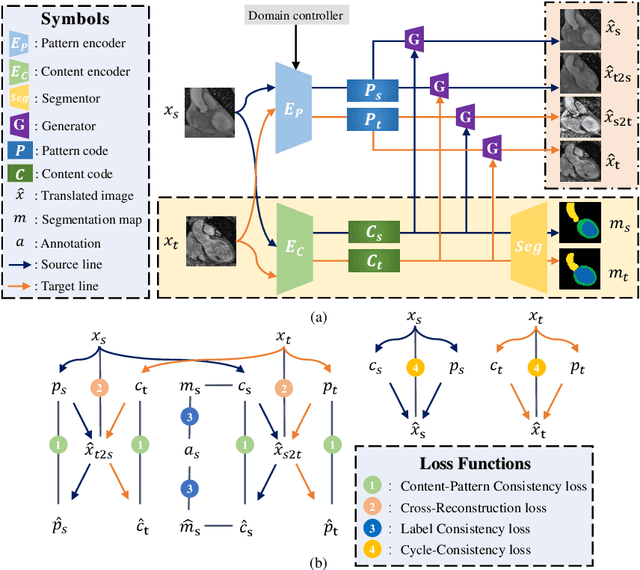
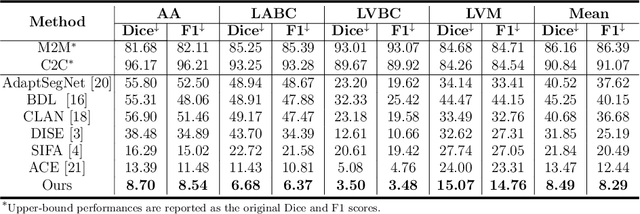
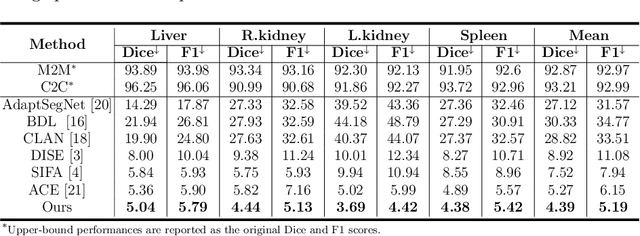
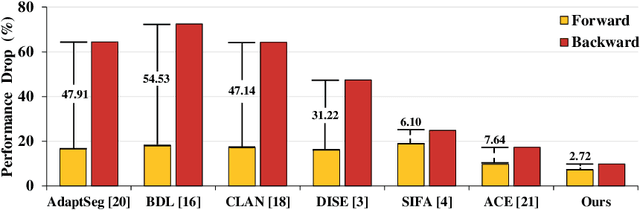
Abstract:Domain shift happens in cross-domain scenarios commonly because of the wide gaps between different domains: when applying a deep learning model well-trained in one domain to another target domain, the model usually performs poorly. To tackle this problem, unsupervised domain adaptation (UDA) techniques are proposed to bridge the gap between different domains, for the purpose of improving model performance without annotation in the target domain. Particularly, UDA has a great value for multimodal medical image analysis, where annotation difficulty is a practical concern. However, most existing UDA methods can only achieve satisfactory improvements in one adaptation direction (e.g., MRI to CT), but often perform poorly in the other (CT to MRI), limiting their practical usage. In this paper, we propose a bidirectional UDA (BiUDA) framework based on disentangled representation learning for equally competent two-way UDA performances. This framework employs a unified domain-aware pattern encoder which not only can adaptively encode images in different domains through a domain controller, but also improve model efficiency by eliminating redundant parameters. Furthermore, to avoid distortion of contents and patterns of input images during the adaptation process, a content-pattern consistency loss is introduced. Additionally, for better UDA segmentation performance, a label consistency strategy is proposed to provide extra supervision by recomposing target-domain-styled images and corresponding source-domain annotations. Comparison experiments and ablation studies conducted on two public datasets demonstrate the superiority of our BiUDA framework to current state-of-the-art UDA methods and the effectiveness of its novel designs. By successfully addressing two-way adaptations, our BiUDA framework offers a flexible solution of UDA techniques to the real-world scenario.
RECIST-Net: Lesion detection via grouping keypoints on RECIST-based annotation
Jul 19, 2021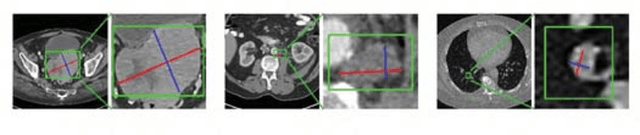
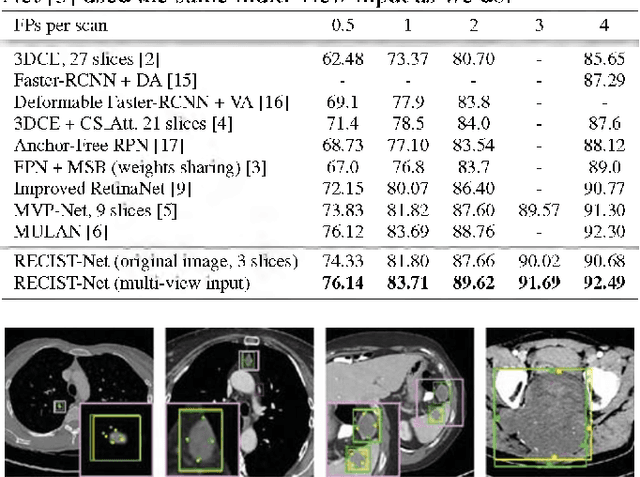
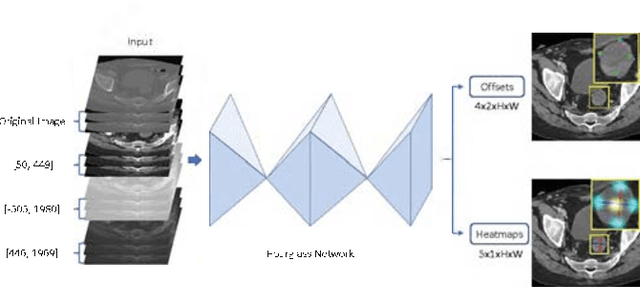
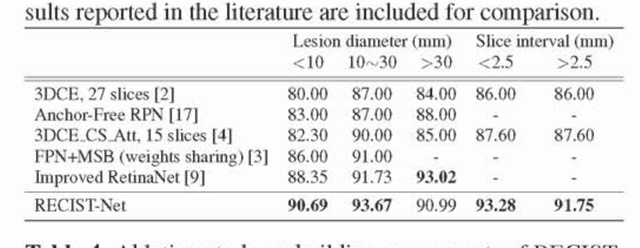
Abstract:Universal lesion detection in computed tomography (CT) images is an important yet challenging task due to the large variations in lesion type, size, shape, and appearance. Considering that data in clinical routine (such as the DeepLesion dataset) are usually annotated with a long and a short diameter according to the standard of Response Evaluation Criteria in Solid Tumors (RECIST) diameters, we propose RECIST-Net, a new approach to lesion detection in which the four extreme points and center point of the RECIST diameters are detected. By detecting a lesion as keypoints, we provide a more conceptually straightforward formulation for detection, and overcome several drawbacks (e.g., requiring extensive effort in designing data-appropriate anchors and losing shape information) of existing bounding-box-based methods while exploring a single-task, one-stage approach compared to other RECIST-based approaches. Experiments show that RECIST-Net achieves a sensitivity of 92.49% at four false positives per image, outperforming other recent methods including those using multi-task learning.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge