Jure Leskovec
Neural Distance Embeddings for Biological Sequences
Oct 11, 2021
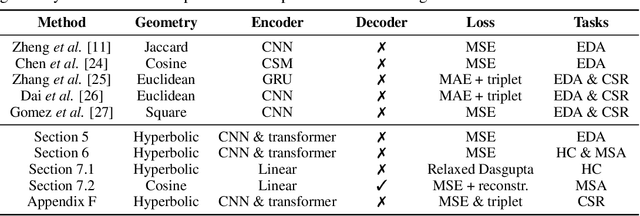
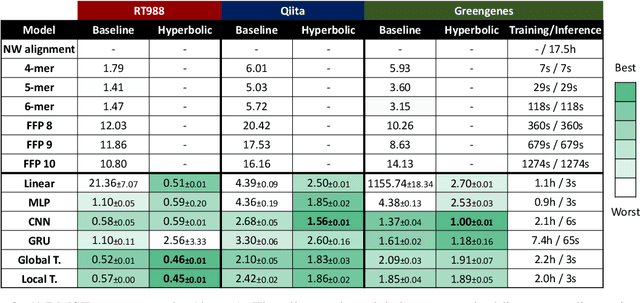
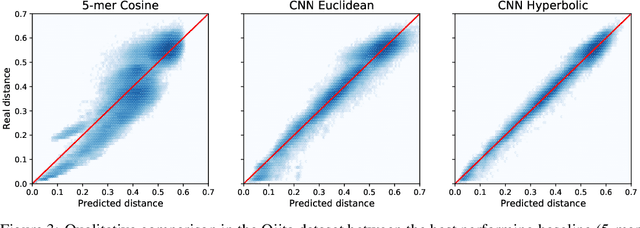
Abstract:The development of data-dependent heuristics and representations for biological sequences that reflect their evolutionary distance is critical for large-scale biological research. However, popular machine learning approaches, based on continuous Euclidean spaces, have struggled with the discrete combinatorial formulation of the edit distance that models evolution and the hierarchical relationship that characterises real-world datasets. We present Neural Distance Embeddings (NeuroSEED), a general framework to embed sequences in geometric vector spaces, and illustrate the effectiveness of the hyperbolic space that captures the hierarchical structure and provides an average 22% reduction in embedding RMSE against the best competing geometry. The capacity of the framework and the significance of these improvements are then demonstrated devising supervised and unsupervised NeuroSEED approaches to multiple core tasks in bioinformatics. Benchmarked with common baselines, the proposed approaches display significant accuracy and/or runtime improvements on real-world datasets. As an example for hierarchical clustering, the proposed pretrained and from-scratch methods match the quality of competing baselines with 30x and 15x runtime reduction, respectively.
LM-Critic: Language Models for Unsupervised Grammatical Error Correction
Sep 14, 2021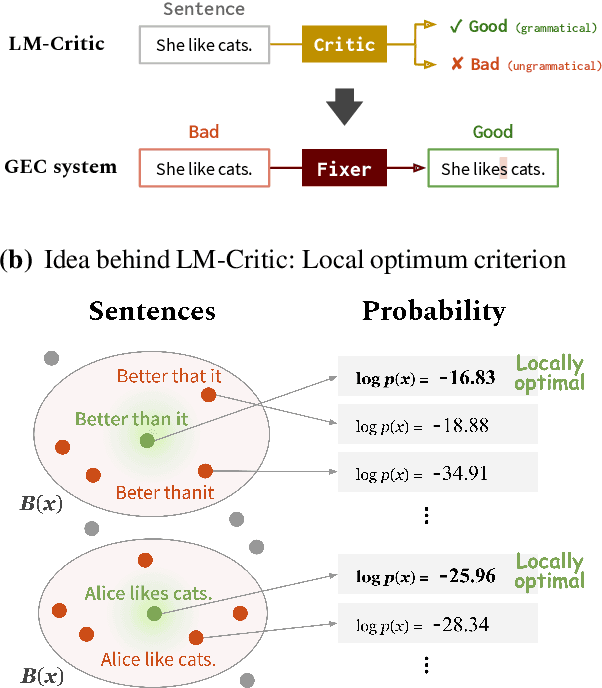
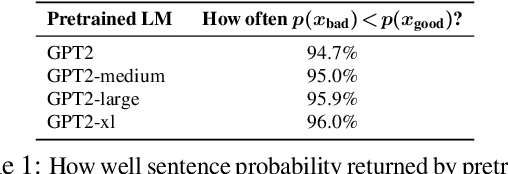
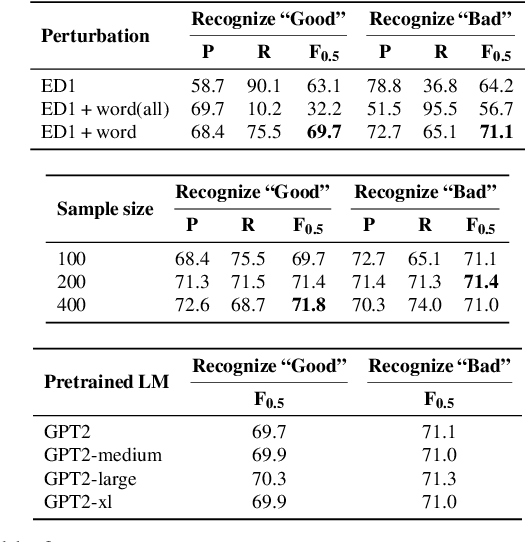
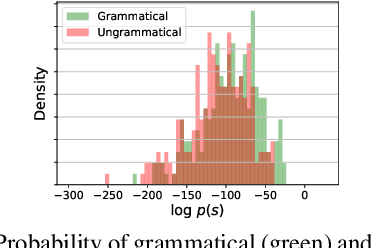
Abstract:Training a model for grammatical error correction (GEC) requires a set of labeled ungrammatical / grammatical sentence pairs, but manually annotating such pairs can be expensive. Recently, the Break-It-Fix-It (BIFI) framework has demonstrated strong results on learning to repair a broken program without any labeled examples, but this relies on a perfect critic (e.g., a compiler) that returns whether an example is valid or not, which does not exist for the GEC task. In this work, we show how to leverage a pretrained language model (LM) in defining an LM-Critic, which judges a sentence to be grammatical if the LM assigns it a higher probability than its local perturbations. We apply this LM-Critic and BIFI along with a large set of unlabeled sentences to bootstrap realistic ungrammatical / grammatical pairs for training a corrector. We evaluate our approach on GEC datasets across multiple domains (CoNLL-2014, BEA-2019, GMEG-wiki and GMEG-yahoo) and show that it outperforms existing methods in both the unsupervised setting (+7.7 F0.5) and the supervised setting (+0.5 F0.5).
On the Opportunities and Risks of Foundation Models
Aug 18, 2021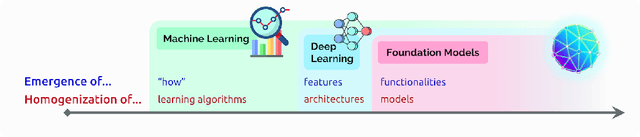
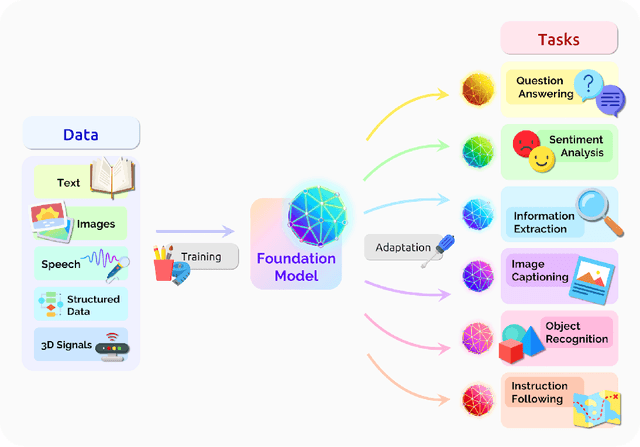
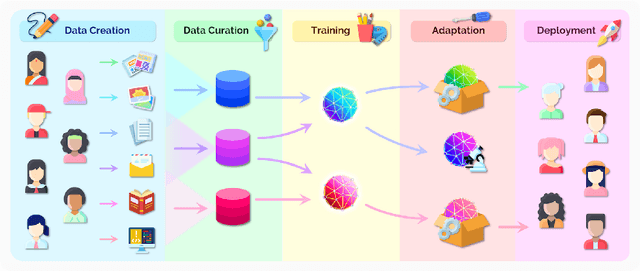
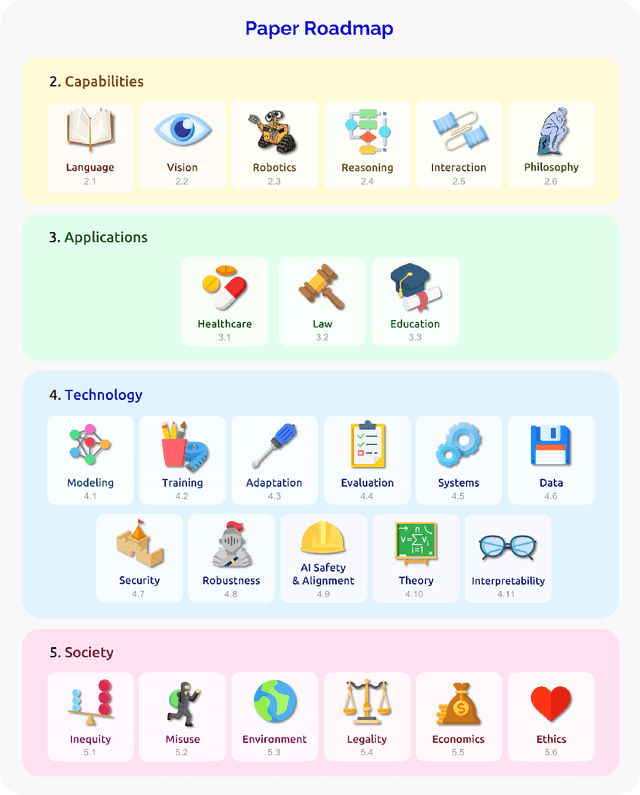
Abstract:AI is undergoing a paradigm shift with the rise of models (e.g., BERT, DALL-E, GPT-3) that are trained on broad data at scale and are adaptable to a wide range of downstream tasks. We call these models foundation models to underscore their critically central yet incomplete character. This report provides a thorough account of the opportunities and risks of foundation models, ranging from their capabilities (e.g., language, vision, robotics, reasoning, human interaction) and technical principles(e.g., model architectures, training procedures, data, systems, security, evaluation, theory) to their applications (e.g., law, healthcare, education) and societal impact (e.g., inequity, misuse, economic and environmental impact, legal and ethical considerations). Though foundation models are based on standard deep learning and transfer learning, their scale results in new emergent capabilities,and their effectiveness across so many tasks incentivizes homogenization. Homogenization provides powerful leverage but demands caution, as the defects of the foundation model are inherited by all the adapted models downstream. Despite the impending widespread deployment of foundation models, we currently lack a clear understanding of how they work, when they fail, and what they are even capable of due to their emergent properties. To tackle these questions, we believe much of the critical research on foundation models will require deep interdisciplinary collaboration commensurate with their fundamentally sociotechnical nature.
Combiner: Full Attention Transformer with Sparse Computation Cost
Jul 12, 2021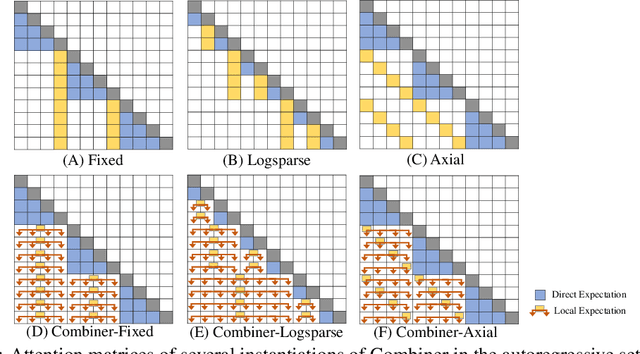
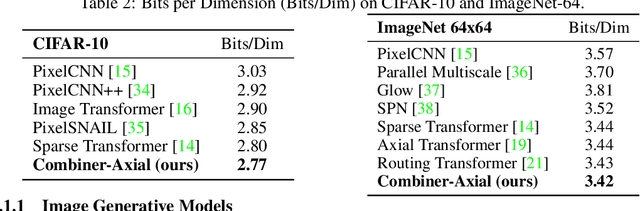
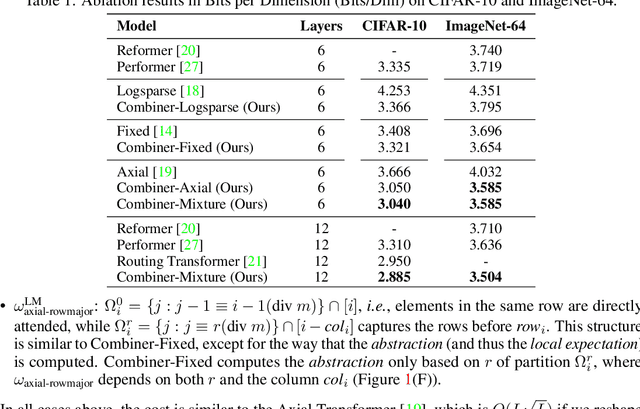
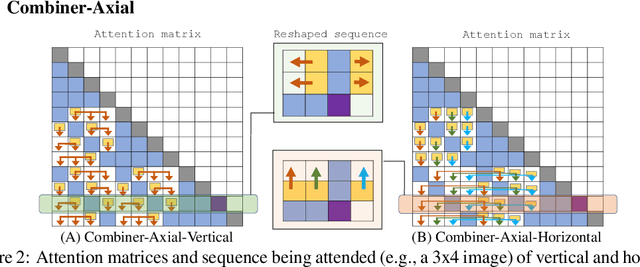

Abstract:Transformers provide a class of expressive architectures that are extremely effective for sequence modeling. However, the key limitation of transformers is their quadratic memory and time complexity $\mathcal{O}(L^2)$ with respect to the sequence length in attention layers, which restricts application in extremely long sequences. Most existing approaches leverage sparsity or low-rank assumptions in the attention matrix to reduce cost, but sacrifice expressiveness. Instead, we propose Combiner, which provides full attention capability in each attention head while maintaining low computation and memory complexity. The key idea is to treat the self-attention mechanism as a conditional expectation over embeddings at each location, and approximate the conditional distribution with a structured factorization. Each location can attend to all other locations, either via direct attention, or through indirect attention to abstractions, which are again conditional expectations of embeddings from corresponding local regions. We show that most sparse attention patterns used in existing sparse transformers are able to inspire the design of such factorization for full attention, resulting in the same sub-quadratic cost ($\mathcal{O}(L\log(L))$ or $\mathcal{O}(L\sqrt{L})$). Combiner is a drop-in replacement for attention layers in existing transformers and can be easily implemented in common frameworks. An experimental evaluation on both autoregressive and bidirectional sequence tasks demonstrates the effectiveness of this approach, yielding state-of-the-art results on several image and text modeling tasks.
GNNAutoScale: Scalable and Expressive Graph Neural Networks via Historical Embeddings
Jun 10, 2021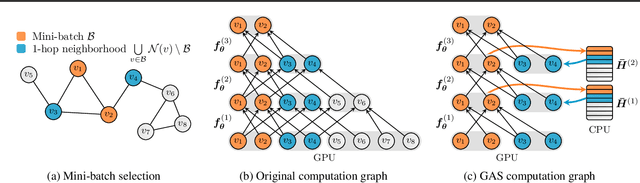
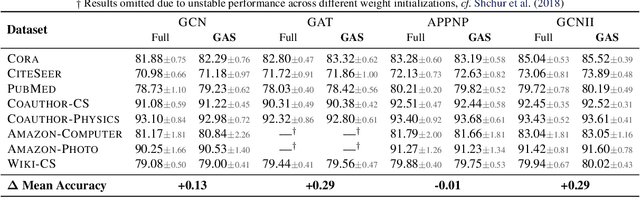
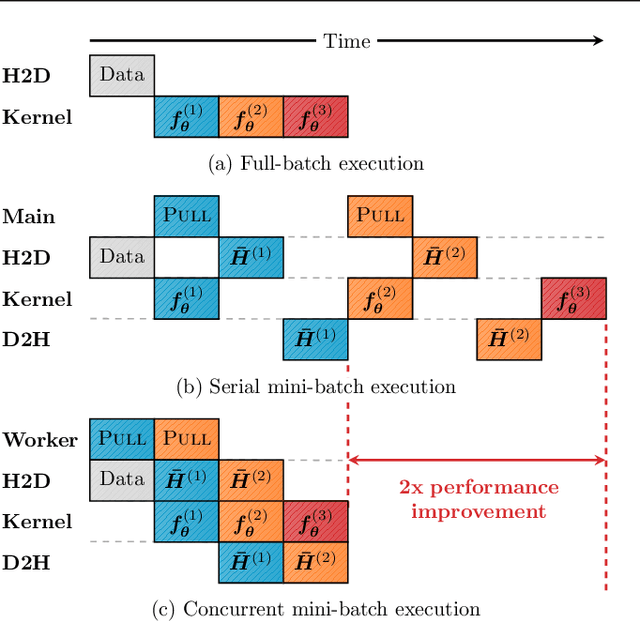
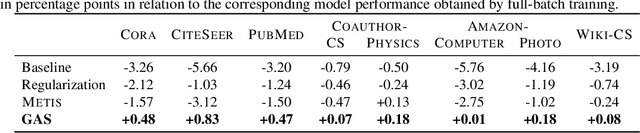
Abstract:We present GNNAutoScale (GAS), a framework for scaling arbitrary message-passing GNNs to large graphs. GAS prunes entire sub-trees of the computation graph by utilizing historical embeddings from prior training iterations, leading to constant GPU memory consumption in respect to input node size without dropping any data. While existing solutions weaken the expressive power of message passing due to sub-sampling of edges or non-trainable propagations, our approach is provably able to maintain the expressive power of the original GNN. We achieve this by providing approximation error bounds of historical embeddings and show how to tighten them in practice. Empirically, we show that the practical realization of our framework, PyGAS, an easy-to-use extension for PyTorch Geometric, is both fast and memory-efficient, learns expressive node representations, closely resembles the performance of their non-scaling counterparts, and reaches state-of-the-art performance on large-scale graphs.
QA-GNN: Reasoning with Language Models and Knowledge Graphs for Question Answering
Apr 13, 2021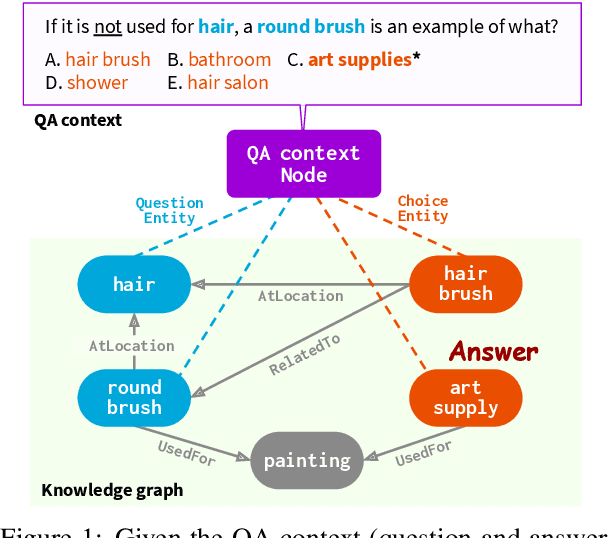
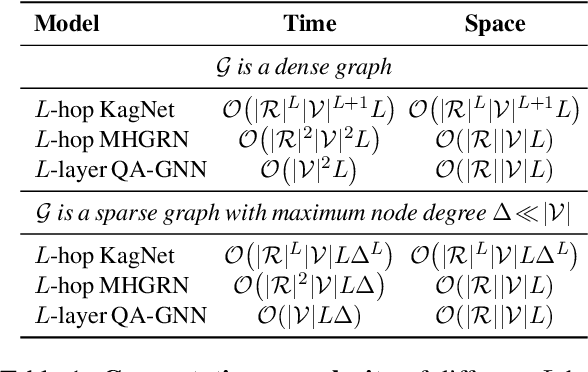

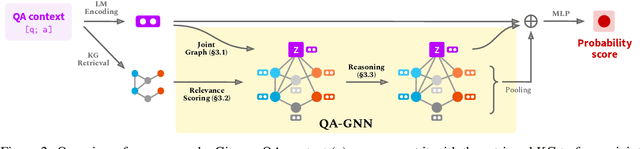
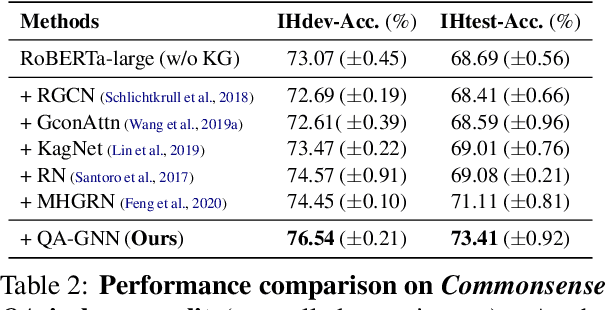
Abstract:The problem of answering questions using knowledge from pre-trained language models (LMs) and knowledge graphs (KGs) presents two challenges: given a QA context (question and answer choice), methods need to (i) identify relevant knowledge from large KGs, and (ii) perform joint reasoning over the QA context and KG. Here we propose a new model, QA-GNN, which addresses the above challenges through two key innovations: (i) relevance scoring, where we use LMs to estimate the importance of KG nodes relative to the given QA context, and (ii) joint reasoning, where we connect the QA context and KG to form a joint graph, and mutually update their representations through graph-based message passing. We evaluate QA-GNN on the CommonsenseQA and OpenBookQA datasets, and show its improvement over existing LM and LM+KG models, as well as its capability to perform interpretable and structured reasoning, e.g., correctly handling negation in questions.
Language-Agnostic Representation Learning of Source Code from Structure and Context
Mar 21, 2021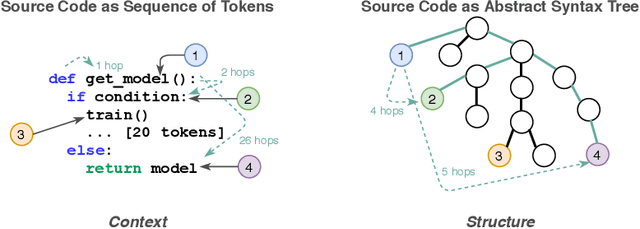
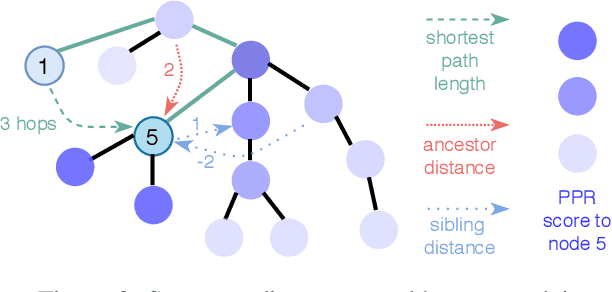
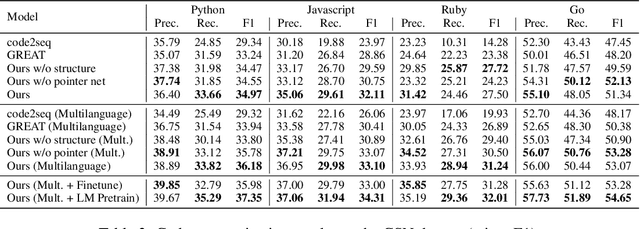
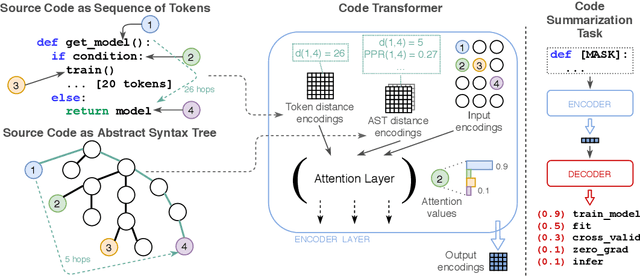
Abstract:Source code (Context) and its parsed abstract syntax tree (AST; Structure) are two complementary representations of the same computer program. Traditionally, designers of machine learning models have relied predominantly either on Structure or Context. We propose a new model, which jointly learns on Context and Structure of source code. In contrast to previous approaches, our model uses only language-agnostic features, i.e., source code and features that can be computed directly from the AST. Besides obtaining state-of-the-art on monolingual code summarization on all five programming languages considered in this work, we propose the first multilingual code summarization model. We show that jointly training on non-parallel data from multiple programming languages improves results on all individual languages, where the strongest gains are on low-resource languages. Remarkably, multilingual training only from Context does not lead to the same improvements, highlighting the benefits of combining Structure and Context for representation learning on code.
OGB-LSC: A Large-Scale Challenge for Machine Learning on Graphs
Mar 17, 2021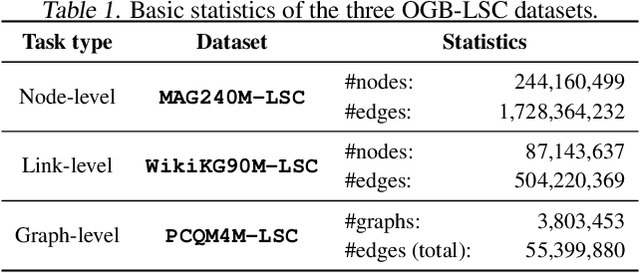
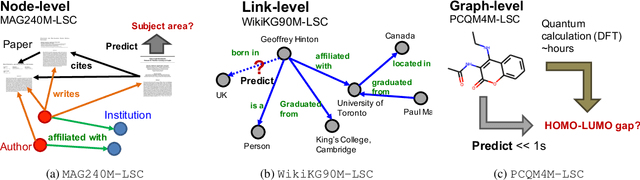
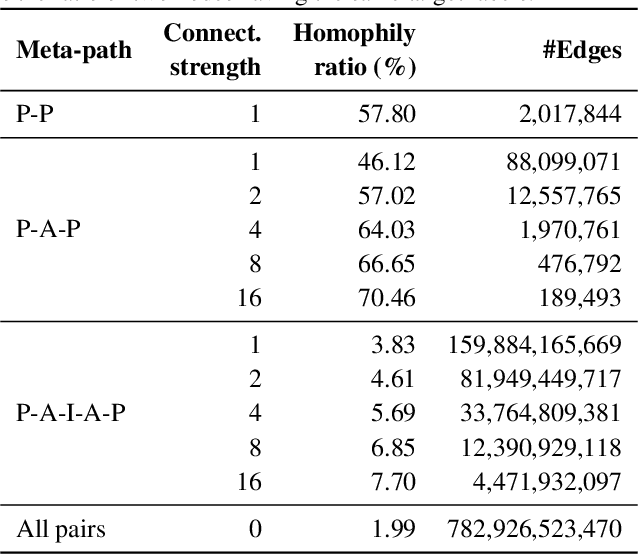
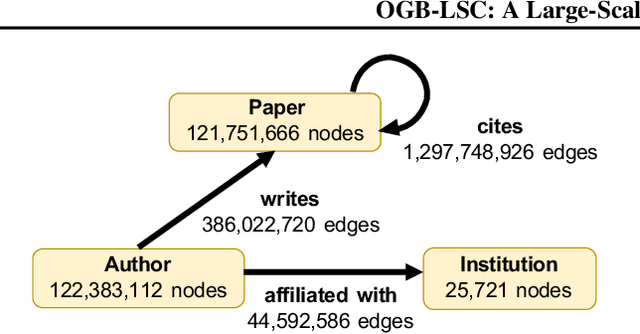
Abstract:Enabling effective and efficient machine learning (ML) over large-scale graph data (e.g., graphs with billions of edges) can have a huge impact on both industrial and scientific applications. However, community efforts to advance large-scale graph ML have been severely limited by the lack of a suitable public benchmark. For KDD Cup 2021, we present OGB Large-Scale Challenge (OGB-LSC), a collection of three real-world datasets for advancing the state-of-the-art in large-scale graph ML. OGB-LSC provides graph datasets that are orders of magnitude larger than existing ones and covers three core graph learning tasks -- link prediction, graph regression, and node classification. Furthermore, OGB-LSC provides dedicated baseline experiments, scaling up expressive graph ML models to the massive datasets. We show that the expressive models significantly outperform simple scalable baselines, indicating an opportunity for dedicated efforts to further improve graph ML at scale. Our datasets and baseline code are released and maintained as part of our OGB initiative (Hu et al., 2020). We hope OGB-LSC at KDD Cup 2021 can empower the community to discover innovative solutions for large-scale graph ML.
ForceNet: A Graph Neural Network for Large-Scale Quantum Calculations
Mar 02, 2021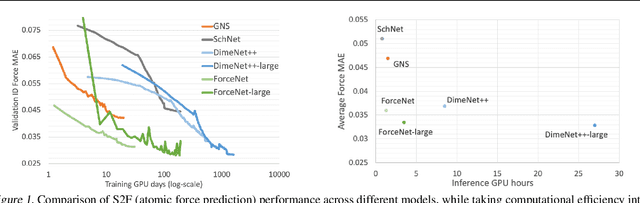
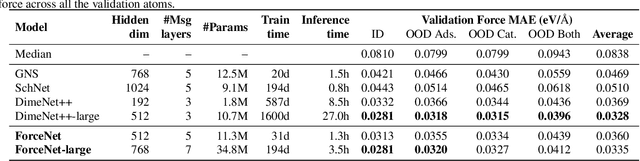
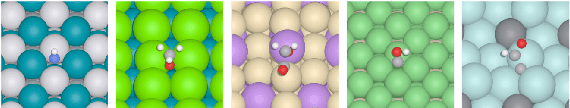
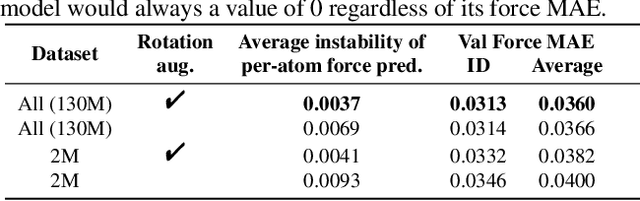
Abstract:With massive amounts of atomic simulation data available, there is a huge opportunity to develop fast and accurate machine learning models to approximate expensive physics-based calculations. The key quantity to estimate is atomic forces, where the state-of-the-art Graph Neural Networks (GNNs) explicitly enforce basic physical constraints such as rotation-covariance. However, to strictly satisfy the physical constraints, existing models have to make tradeoffs between computational efficiency and model expressiveness. Here we explore an alternative approach. By not imposing explicit physical constraints, we can flexibly design expressive models while maintaining their computational efficiency. Physical constraints are implicitly imposed by training the models using physics-based data augmentation. To evaluate the approach, we carefully design a scalable and expressive GNN model, ForceNet, and apply it to OC20 (Chanussot et al., 2020), an unprecedentedly-large dataset of quantum physics calculations. Our proposed ForceNet is able to predict atomic forces more accurately than state-of-the-art physics-based GNNs while being faster both in training and inference. Overall, our promising and counter-intuitive results open up an exciting avenue for future research.
Therapeutics Data Commons: Machine Learning Datasets and Tasks for Therapeutics
Feb 18, 2021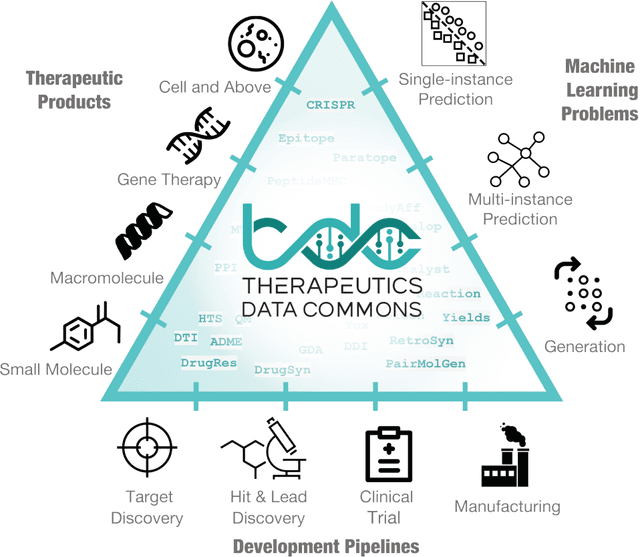
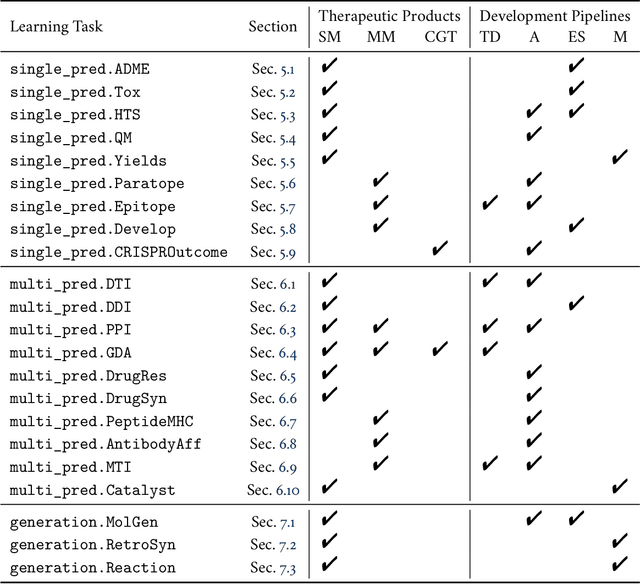
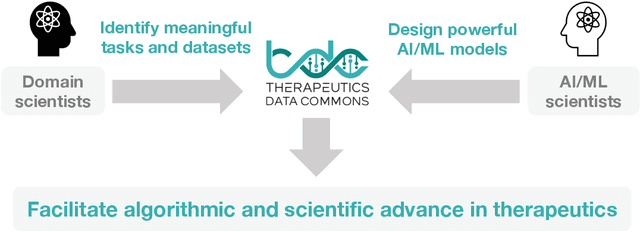
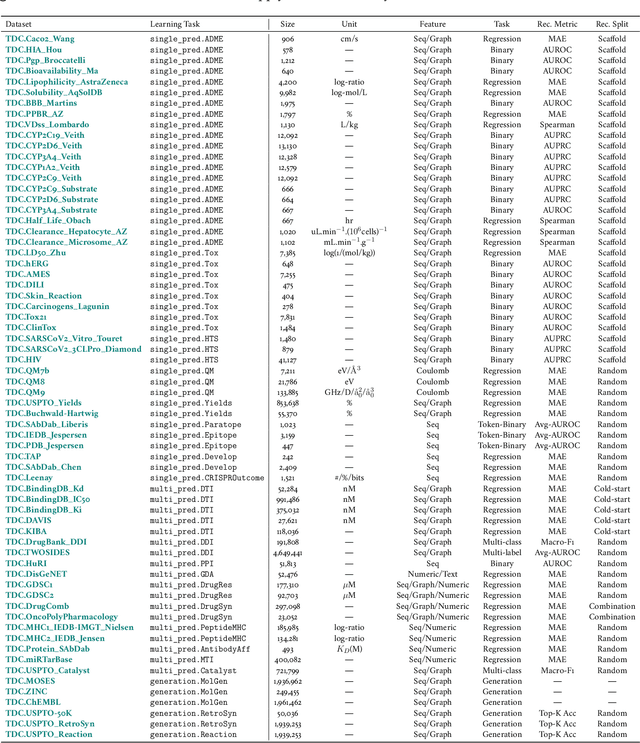
Abstract:Machine learning for therapeutics is an emerging field with incredible opportunities for innovation and expansion. Despite the initial success, many key challenges remain open. Here, we introduce Therapeutics Data Commons (TDC), the first unifying framework to systematically access and evaluate machine learning across the entire range of therapeutics. At its core, TDC is a collection of curated datasets and learning tasks that can translate algorithmic innovation into biomedical and clinical implementation. To date, TDC includes 66 machine learning-ready datasets from 22 learning tasks, spanning the discovery and development of safe and effective medicines. TDC also provides an ecosystem of tools, libraries, leaderboards, and community resources, including data functions, strategies for systematic model evaluation, meaningful data splits, data processors, and molecule generation oracles. All datasets and learning tasks are integrated and accessible via an open-source library. We envision that TDC can facilitate algorithmic and scientific advances and accelerate development, validation, and transition into production and clinical implementation. TDC is a continuous, open-source initiative, and we invite contributions from the research community. TDC is publicly available at https://tdcommons.ai.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge