Greg S. Corrado
Predicting Cardiovascular Disease Risk using Photoplethysmography and Deep Learning
May 09, 2023Abstract:Cardiovascular diseases (CVDs) are responsible for a large proportion of premature deaths in low- and middle-income countries. Early CVD detection and intervention is critical in these populations, yet many existing CVD risk scores require a physical examination or lab measurements, which can be challenging in such health systems due to limited accessibility. Here we investigated the potential to use photoplethysmography (PPG), a sensing technology available on most smartphones that can potentially enable large-scale screening at low cost, for CVD risk prediction. We developed a deep learning PPG-based CVD risk score (DLS) to predict the probability of having major adverse cardiovascular events (MACE: non-fatal myocardial infarction, stroke, and cardiovascular death) within ten years, given only age, sex, smoking status and PPG as predictors. We compared the DLS with the office-based refit-WHO score, which adopts the shared predictors from WHO and Globorisk scores (age, sex, smoking status, height, weight and systolic blood pressure) but refitted on the UK Biobank (UKB) cohort. In UKB cohort, DLS's C-statistic (71.1%, 95% CI 69.9-72.4) was non-inferior to office-based refit-WHO score (70.9%, 95% CI 69.7-72.2; non-inferiority margin of 2.5%, p<0.01). The calibration of the DLS was satisfactory, with a 1.8% mean absolute calibration error. Adding DLS features to the office-based score increased the C-statistic by 1.0% (95% CI 0.6-1.4). DLS predicts ten-year MACE risk comparable with the office-based refit-WHO score. It provides a proof-of-concept and suggests the potential of a PPG-based approach strategies for community-based primary prevention in resource-limited regions.
Large Language Models Encode Clinical Knowledge
Dec 26, 2022Abstract:Large language models (LLMs) have demonstrated impressive capabilities in natural language understanding and generation, but the quality bar for medical and clinical applications is high. Today, attempts to assess models' clinical knowledge typically rely on automated evaluations on limited benchmarks. There is no standard to evaluate model predictions and reasoning across a breadth of tasks. To address this, we present MultiMedQA, a benchmark combining six existing open question answering datasets spanning professional medical exams, research, and consumer queries; and HealthSearchQA, a new free-response dataset of medical questions searched online. We propose a framework for human evaluation of model answers along multiple axes including factuality, precision, possible harm, and bias. In addition, we evaluate PaLM (a 540-billion parameter LLM) and its instruction-tuned variant, Flan-PaLM, on MultiMedQA. Using a combination of prompting strategies, Flan-PaLM achieves state-of-the-art accuracy on every MultiMedQA multiple-choice dataset (MedQA, MedMCQA, PubMedQA, MMLU clinical topics), including 67.6% accuracy on MedQA (US Medical License Exam questions), surpassing prior state-of-the-art by over 17%. However, human evaluation reveals key gaps in Flan-PaLM responses. To resolve this we introduce instruction prompt tuning, a parameter-efficient approach for aligning LLMs to new domains using a few exemplars. The resulting model, Med-PaLM, performs encouragingly, but remains inferior to clinicians. We show that comprehension, recall of knowledge, and medical reasoning improve with model scale and instruction prompt tuning, suggesting the potential utility of LLMs in medicine. Our human evaluations reveal important limitations of today's models, reinforcing the importance of both evaluation frameworks and method development in creating safe, helpful LLM models for clinical applications.
Discovering novel systemic biomarkers in photos of the external eye
Jul 19, 2022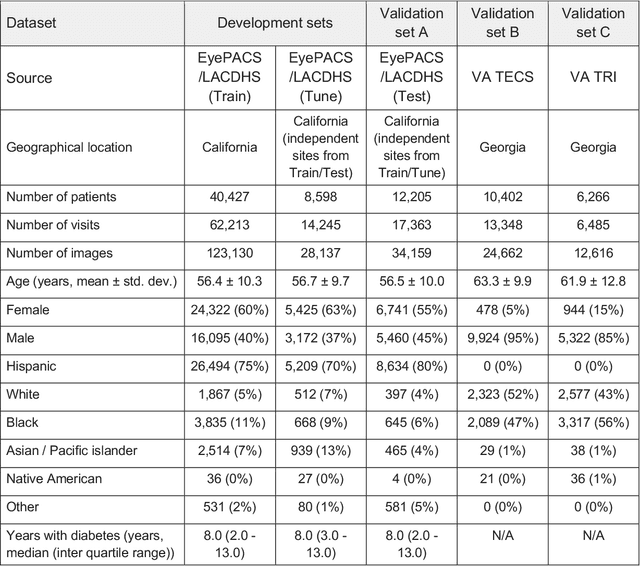
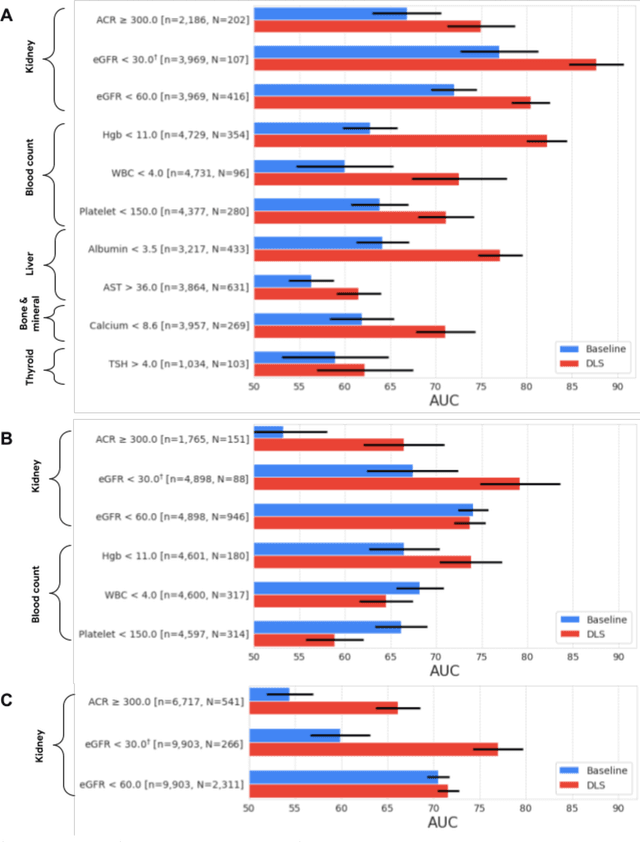
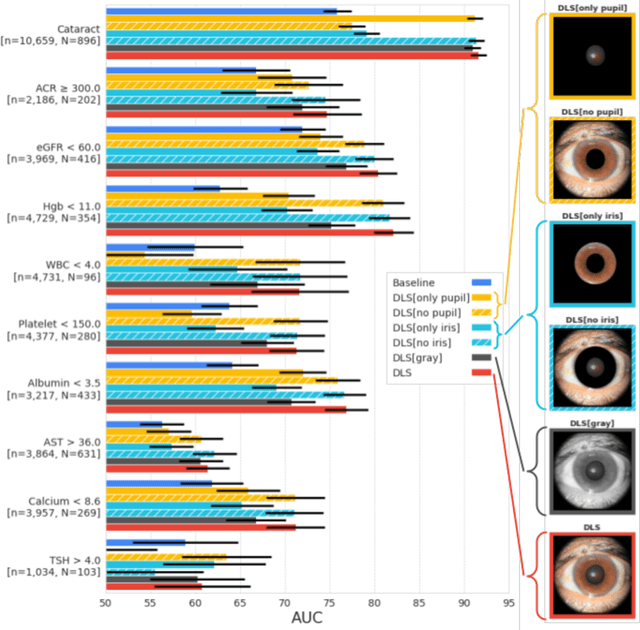
Abstract:External eye photos were recently shown to reveal signs of diabetic retinal disease and elevated HbA1c. In this paper, we evaluate if external eye photos contain information about additional systemic medical conditions. We developed a deep learning system (DLS) that takes external eye photos as input and predicts multiple systemic parameters, such as those related to the liver (albumin, AST); kidney (eGFR estimated using the race-free 2021 CKD-EPI creatinine equation, the urine ACR); bone & mineral (calcium); thyroid (TSH); and blood count (Hgb, WBC, platelets). Development leveraged 151,237 images from 49,015 patients with diabetes undergoing diabetic eye screening in 11 sites across Los Angeles county, CA. Evaluation focused on 9 pre-specified systemic parameters and leveraged 3 validation sets (A, B, C) spanning 28,869 patients with and without diabetes undergoing eye screening in 3 independent sites in Los Angeles County, CA, and the greater Atlanta area, GA. We compared against baseline models incorporating available clinicodemographic variables (e.g. age, sex, race/ethnicity, years with diabetes). Relative to the baseline, the DLS achieved statistically significant superior performance at detecting AST>36, calcium<8.6, eGFR<60, Hgb<11, platelets<150, ACR>=300, and WBC<4 on validation set A (a patient population similar to the development sets), where the AUC of DLS exceeded that of the baseline by 5.2-19.4%. On validation sets B and C, with substantial patient population differences compared to the development sets, the DLS outperformed the baseline for ACR>=300 and Hgb<11 by 7.3-13.2%. Our findings provide further evidence that external eye photos contain important biomarkers of systemic health spanning multiple organ systems. Further work is needed to investigate whether and how these biomarkers can be translated into clinical impact.
Robust and Efficient Medical Imaging with Self-Supervision
May 19, 2022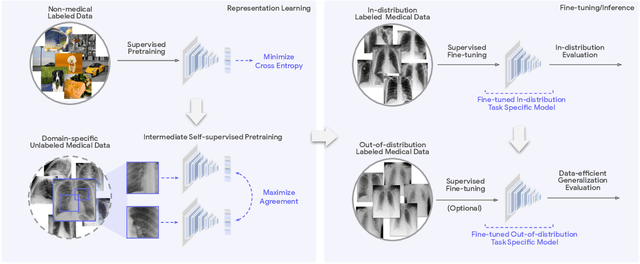
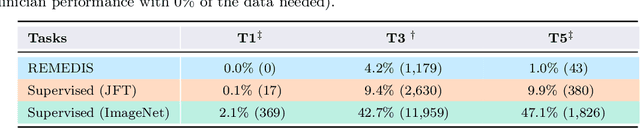
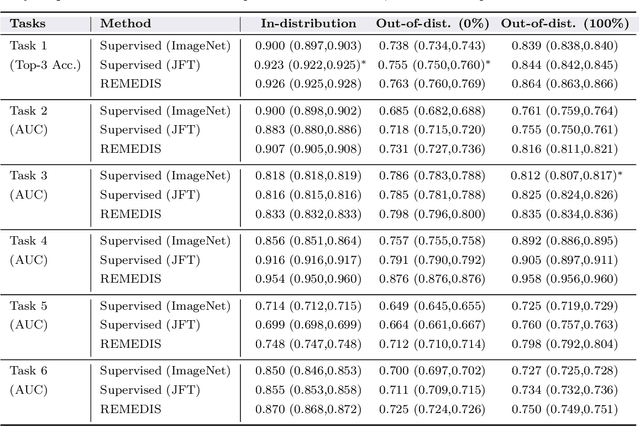
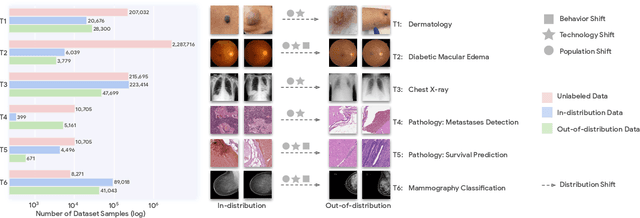

Abstract:Recent progress in Medical Artificial Intelligence (AI) has delivered systems that can reach clinical expert level performance. However, such systems tend to demonstrate sub-optimal "out-of-distribution" performance when evaluated in clinical settings different from the training environment. A common mitigation strategy is to develop separate systems for each clinical setting using site-specific data [1]. However, this quickly becomes impractical as medical data is time-consuming to acquire and expensive to annotate [2]. Thus, the problem of "data-efficient generalization" presents an ongoing difficulty for Medical AI development. Although progress in representation learning shows promise, their benefits have not been rigorously studied, specifically for out-of-distribution settings. To meet these challenges, we present REMEDIS, a unified representation learning strategy to improve robustness and data-efficiency of medical imaging AI. REMEDIS uses a generic combination of large-scale supervised transfer learning with self-supervised learning and requires little task-specific customization. We study a diverse range of medical imaging tasks and simulate three realistic application scenarios using retrospective data. REMEDIS exhibits significantly improved in-distribution performance with up to 11.5% relative improvement in diagnostic accuracy over a strong supervised baseline. More importantly, our strategy leads to strong data-efficient generalization of medical imaging AI, matching strong supervised baselines using between 1% to 33% of retraining data across tasks. These results suggest that REMEDIS can significantly accelerate the life-cycle of medical imaging AI development thereby presenting an important step forward for medical imaging AI to deliver broad impact.
Deep learning for detecting pulmonary tuberculosis via chest radiography: an international study across 10 countries
May 16, 2021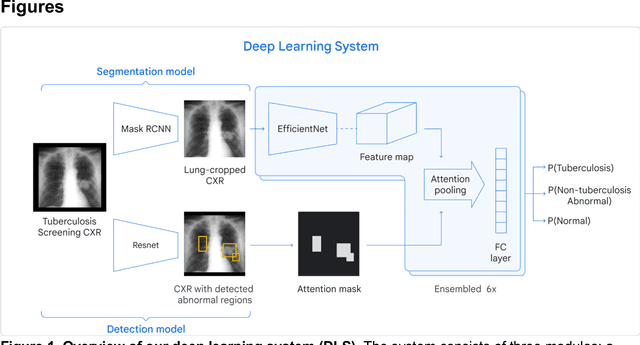
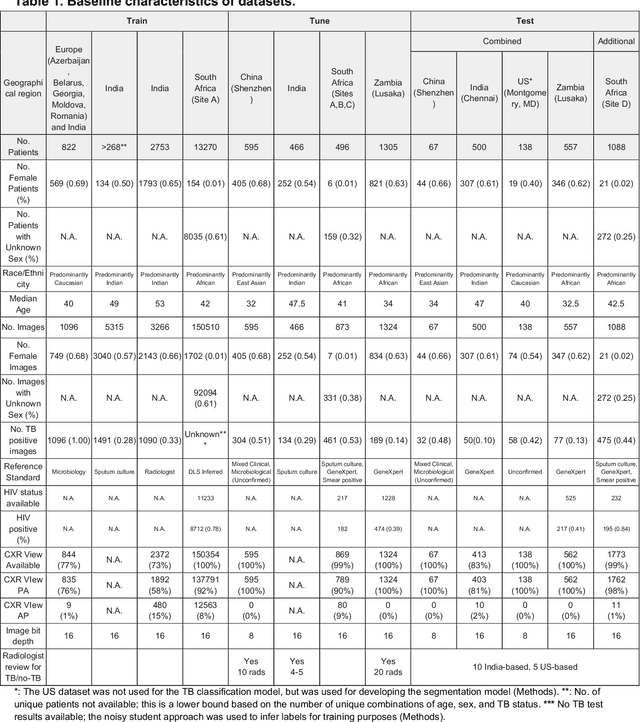
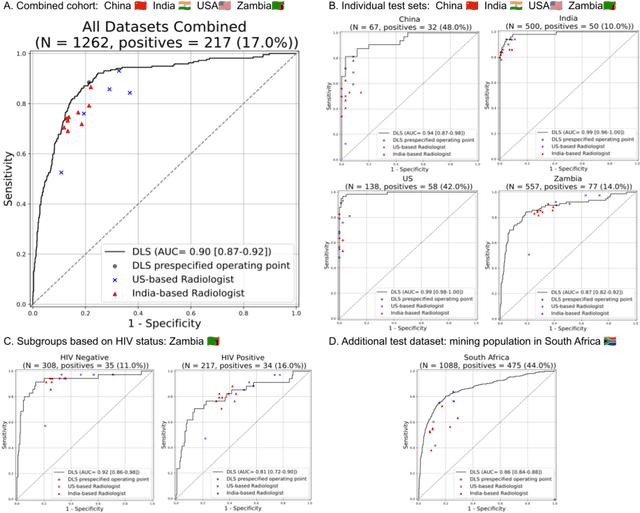
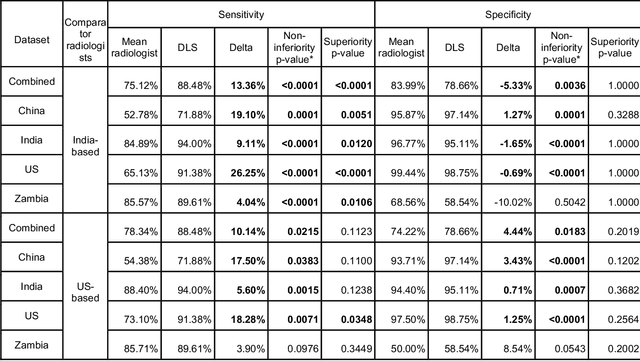
Abstract:Tuberculosis (TB) is a top-10 cause of death worldwide. Though the WHO recommends chest radiographs (CXRs) for TB screening, the limited availability of CXR interpretation is a barrier. We trained a deep learning system (DLS) to detect active pulmonary TB using CXRs from 9 countries across Africa, Asia, and Europe, and utilized large-scale CXR pretraining, attention pooling, and noisy student semi-supervised learning. Evaluation was on (1) a combined test set spanning China, India, US, and Zambia, and (2) an independent mining population in South Africa. Given WHO targets of 90% sensitivity and 70% specificity, the DLS's operating point was prespecified to favor sensitivity over specificity. On the combined test set, the DLS's ROC curve was above all 9 India-based radiologists, with an AUC of 0.90 (95%CI 0.87-0.92). The DLS's sensitivity (88%) was higher than the India-based radiologists (75% mean sensitivity), p<0.001 for superiority; and its specificity (79%) was non-inferior to the radiologists (84% mean specificity), p=0.004. Similar trends were observed within HIV positive and sputum smear positive sub-groups, and in the South Africa test set. We found that 5 US-based radiologists (where TB isn't endemic) were more sensitive and less specific than the India-based radiologists (where TB is endemic). The DLS also remained non-inferior to the US-based radiologists. In simulations, using the DLS as a prioritization tool for confirmatory testing reduced the cost per positive case detected by 40-80% compared to using confirmatory testing alone. To conclude, our DLS generalized to 5 countries, and merits prospective evaluation to assist cost-effective screening efforts in radiologist-limited settings. Operating point flexibility may permit customization of the DLS to account for site-specific factors such as TB prevalence, demographics, clinical resources, and customary practice patterns.
Does Your Dermatology Classifier Know What It Doesn't Know? Detecting the Long-Tail of Unseen Conditions
Apr 08, 2021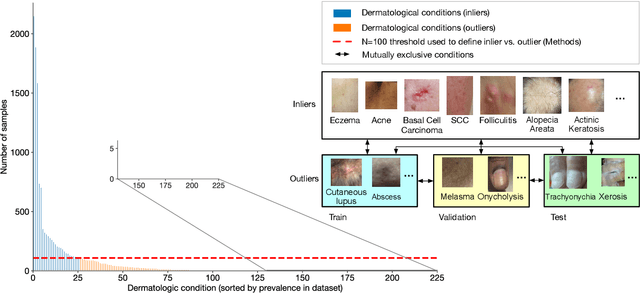
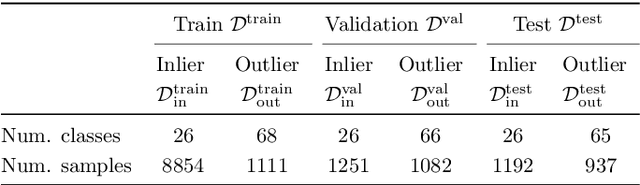
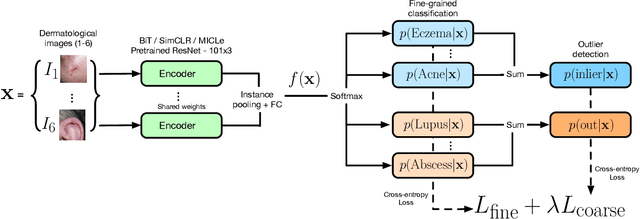
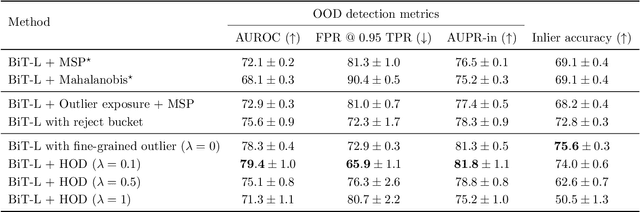
Abstract:We develop and rigorously evaluate a deep learning based system that can accurately classify skin conditions while detecting rare conditions for which there is not enough data available for training a confident classifier. We frame this task as an out-of-distribution (OOD) detection problem. Our novel approach, hierarchical outlier detection (HOD) assigns multiple abstention classes for each training outlier class and jointly performs a coarse classification of inliers vs. outliers, along with fine-grained classification of the individual classes. We demonstrate the effectiveness of the HOD loss in conjunction with modern representation learning approaches (BiT, SimCLR, MICLe) and explore different ensembling strategies for further improving the results. We perform an extensive subgroup analysis over conditions of varying risk levels and different skin types to investigate how the OOD detection performance changes over each subgroup and demonstrate the gains of our framework in comparison to baselines. Finally, we introduce a cost metric to approximate downstream clinical impact. We use this cost metric to compare the proposed method against a baseline system, thereby making a stronger case for the overall system effectiveness in a real-world deployment scenario.
Predicting Prostate Cancer-Specific Mortality with A.I.-based Gleason Grading
Nov 25, 2020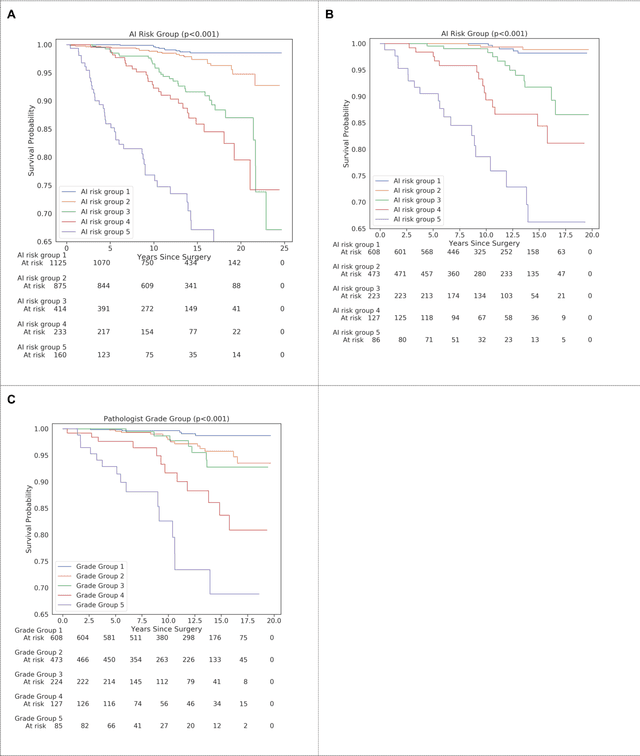
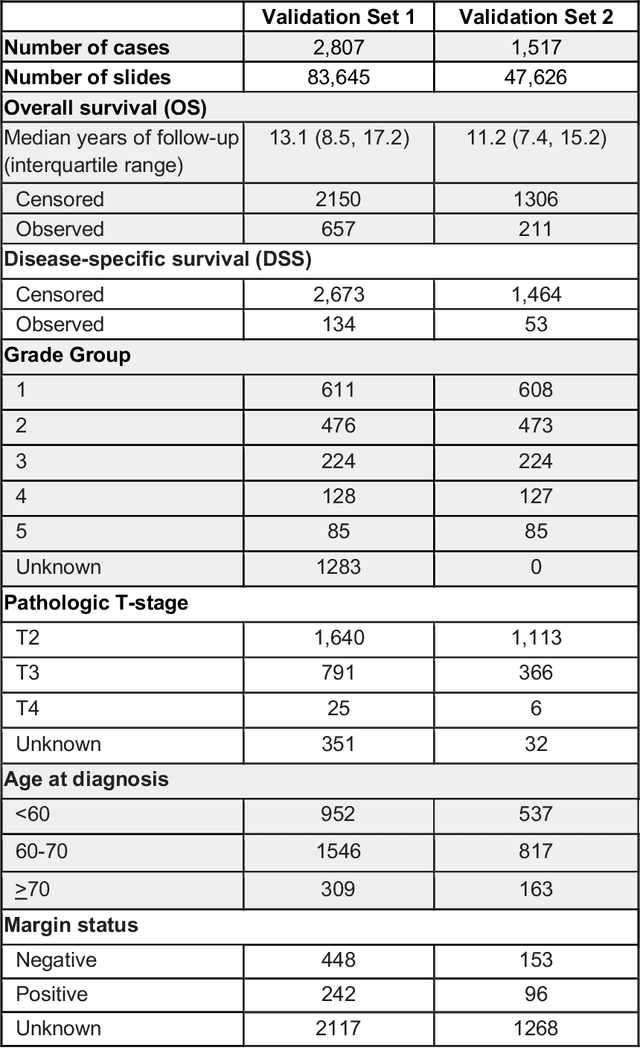
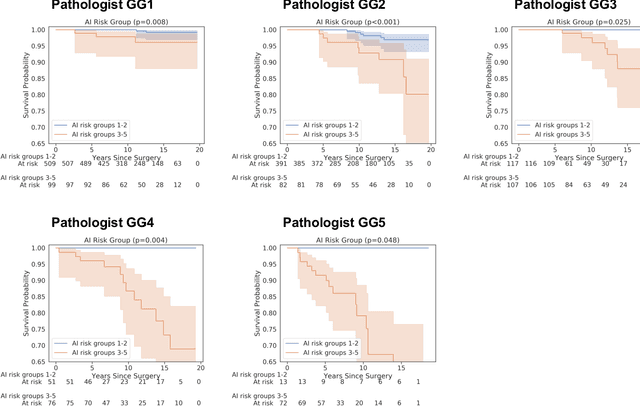
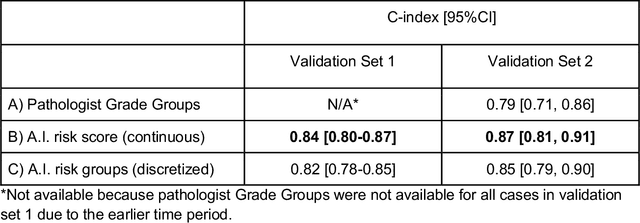
Abstract:Gleason grading of prostate cancer is an important prognostic factor but suffers from poor reproducibility, particularly among non-subspecialist pathologists. Although artificial intelligence (A.I.) tools have demonstrated Gleason grading on-par with expert pathologists, it remains an open question whether A.I. grading translates to better prognostication. In this study, we developed a system to predict prostate-cancer specific mortality via A.I.-based Gleason grading and subsequently evaluated its ability to risk-stratify patients on an independent retrospective cohort of 2,807 prostatectomy cases from a single European center with 5-25 years of follow-up (median: 13, interquartile range 9-17). The A.I.'s risk scores produced a C-index of 0.84 (95%CI 0.80-0.87) for prostate cancer-specific mortality. Upon discretizing these risk scores into risk groups analogous to pathologist Grade Groups (GG), the A.I. had a C-index of 0.82 (95%CI 0.78-0.85). On the subset of cases with a GG in the original pathology report (n=1,517), the A.I.'s C-indices were 0.87 and 0.85 for continuous and discrete grading, respectively, compared to 0.79 (95%CI 0.71-0.86) for GG obtained from the reports. These represent improvements of 0.08 (95%CI 0.01-0.15) and 0.07 (95%CI 0.00-0.14) respectively. Our results suggest that A.I.-based Gleason grading can lead to effective risk-stratification and warrants further evaluation for improving disease management.
Detecting hidden signs of diabetes in external eye photographs
Nov 23, 2020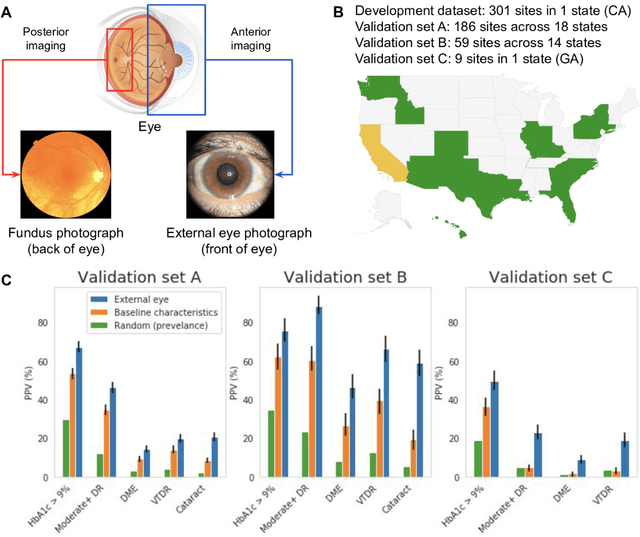
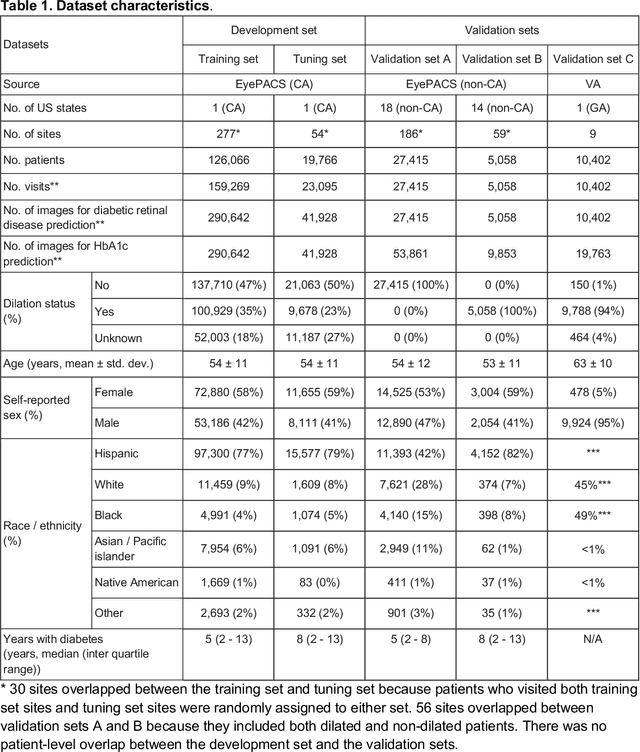
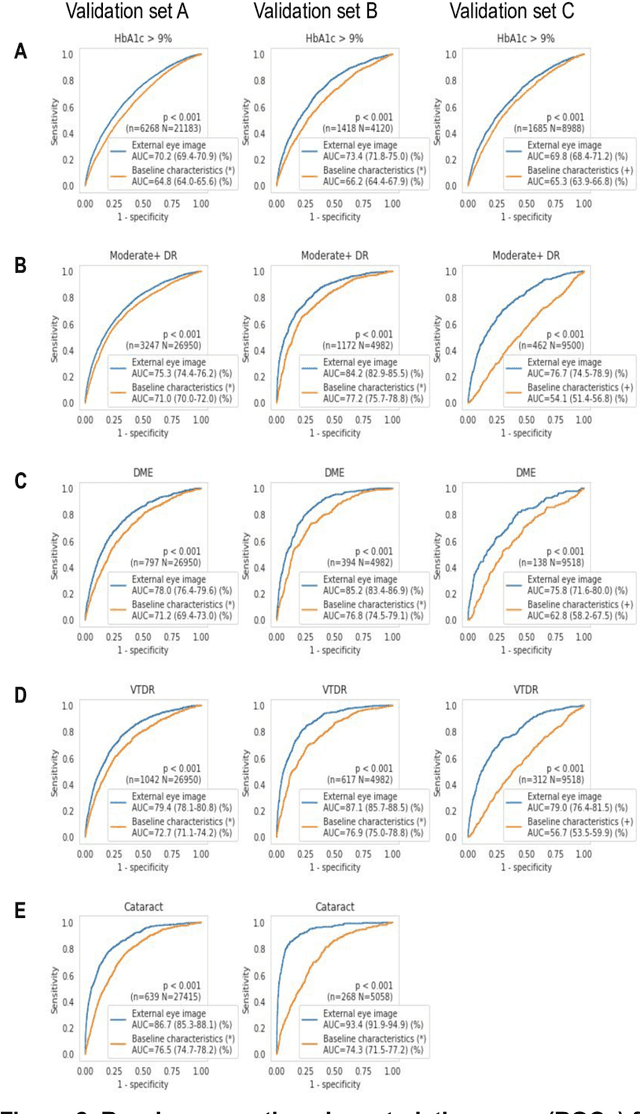
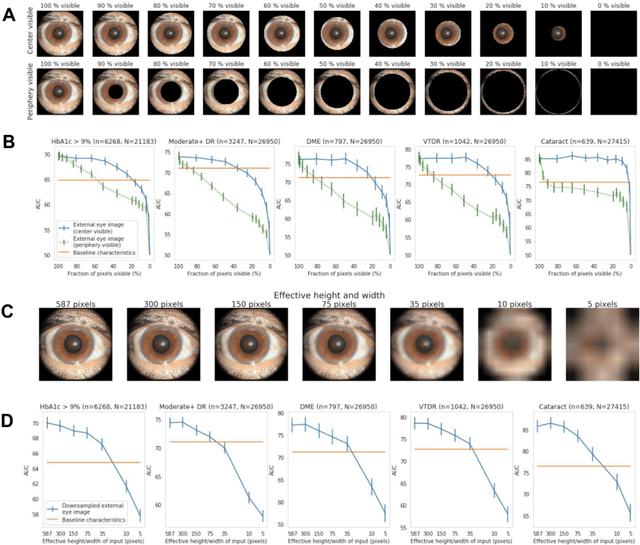
Abstract:Diabetes-related retinal conditions can be detected by examining the posterior of the eye. By contrast, examining the anterior of the eye can reveal conditions affecting the front of the eye, such as changes to the eyelids, cornea, or crystalline lens. In this work, we studied whether external photographs of the front of the eye can reveal insights into both diabetic retinal diseases and blood glucose control. We developed a deep learning system (DLS) using external eye photographs of 145,832 patients with diabetes from 301 diabetic retinopathy (DR) screening sites in one US state, and evaluated the DLS on three validation sets containing images from 198 sites in 18 other US states. In validation set A (n=27,415 patients, all undilated), the DLS detected poor blood glucose control (HbA1c > 9%) with an area under receiver operating characteristic curve (AUC) of 70.2; moderate-or-worse DR with an AUC of 75.3; diabetic macular edema with an AUC of 78.0; and vision-threatening DR with an AUC of 79.4. For all 4 prediction tasks, the DLS's AUC was higher (p<0.001) than using available self-reported baseline characteristics (age, sex, race/ethnicity, years with diabetes). In terms of positive predictive value, the predicted top 5% of patients had a 67% chance of having HbA1c > 9%, and a 20% chance of having vision threatening diabetic retinopathy. The results generalized to dilated pupils (validation set B, 5,058 patients) and to a different screening service (validation set C, 10,402 patients). Our results indicate that external eye photographs contain information useful for healthcare providers managing patients with diabetes, and may help prioritize patients for in-person screening. Further work is needed to validate these findings on different devices and patient populations (those without diabetes) to evaluate its utility for remote diagnosis and management.
Interpretable Survival Prediction for Colorectal Cancer using Deep Learning
Nov 17, 2020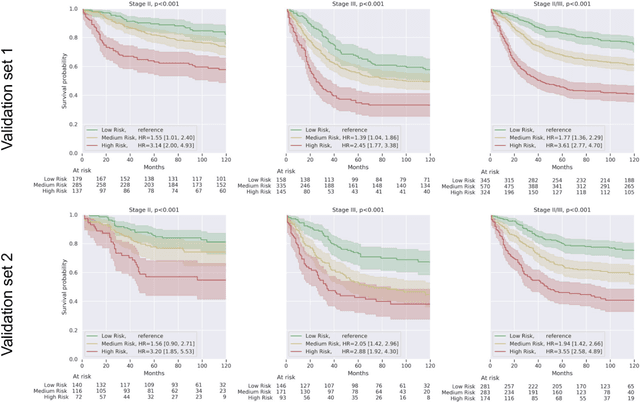
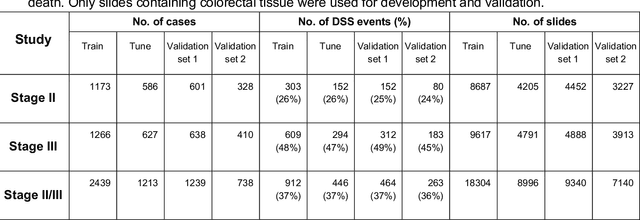
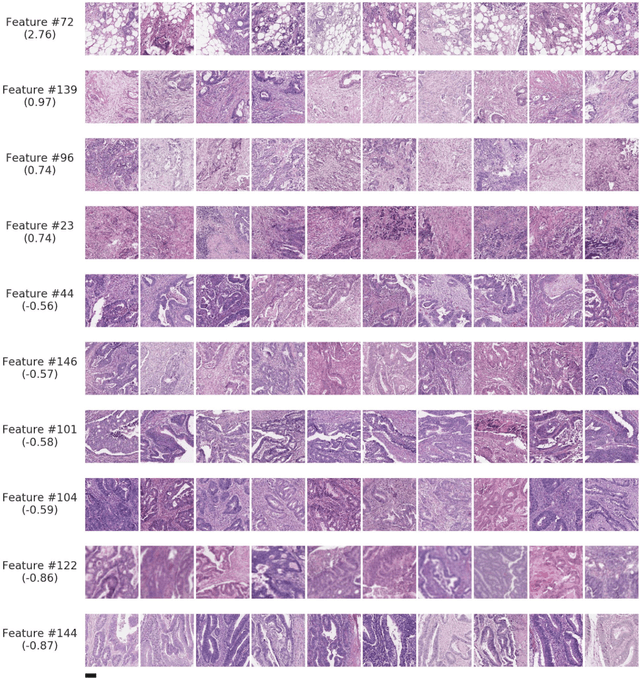
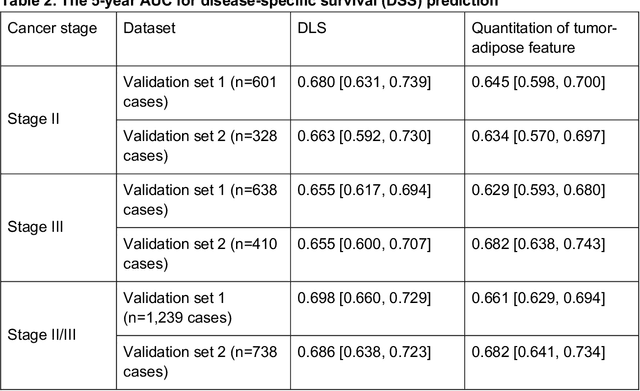
Abstract:Deriving interpretable prognostic features from deep-learning-based prognostic histopathology models remains a challenge. In this study, we developed a deep learning system (DLS) for predicting disease specific survival for stage II and III colorectal cancer using 3,652 cases (27,300 slides). When evaluated on two validation datasets containing 1,239 cases (9,340 slides) and 738 cases (7,140 slides) respectively, the DLS achieved a 5-year disease-specific survival AUC of 0.70 (95%CI 0.66-0.73) and 0.69 (95%CI 0.64-0.72), and added significant predictive value to a set of 9 clinicopathologic features. To interpret the DLS, we explored the ability of different human-interpretable features to explain the variance in DLS scores. We observed that clinicopathologic features such as T-category, N-category, and grade explained a small fraction of the variance in DLS scores (R2=18% in both validation sets). Next, we generated human-interpretable histologic features by clustering embeddings from a deep-learning based image-similarity model and showed that they explain the majority of the variance (R2 of 73% to 80%). Furthermore, the clustering-derived feature most strongly associated with high DLS scores was also highly prognostic in isolation. With a distinct visual appearance (poorly differentiated tumor cell clusters adjacent to adipose tissue), this feature was identified by annotators with 87.0-95.5% accuracy. Our approach can be used to explain predictions from a prognostic deep learning model and uncover potentially-novel prognostic features that can be reliably identified by people for future validation studies.
Deep Learning for Distinguishing Normal versus Abnormal Chest Radiographs and Generalization to Unseen Diseases
Oct 22, 2020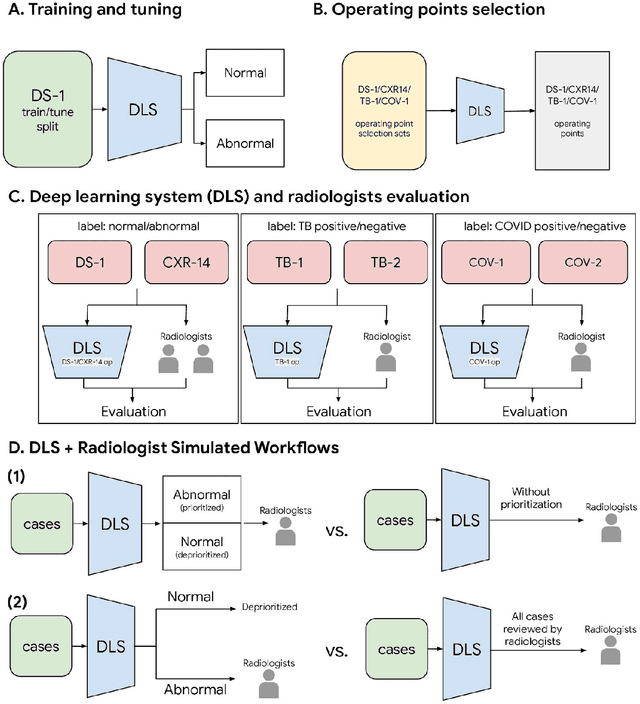
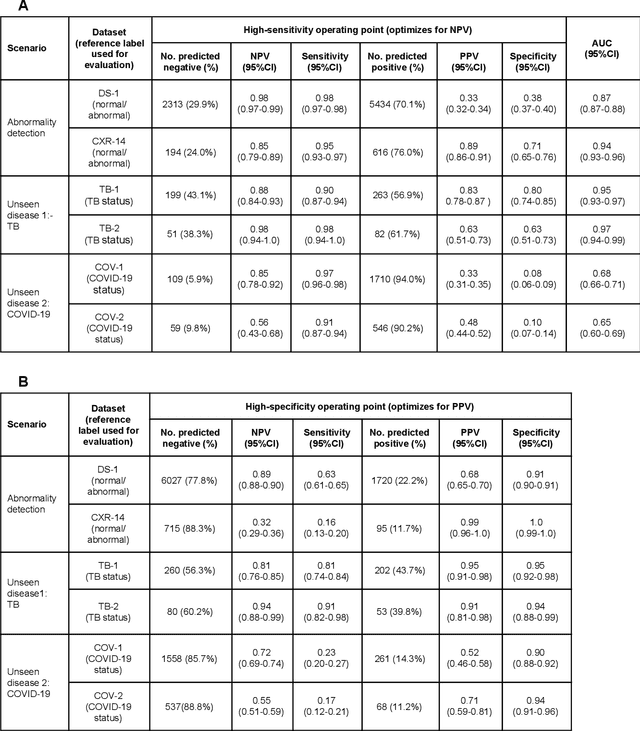
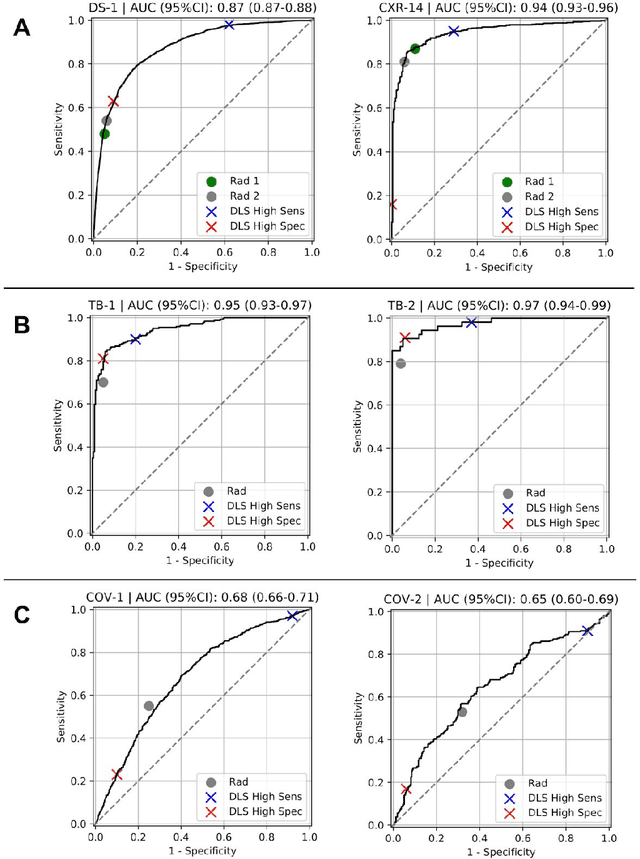
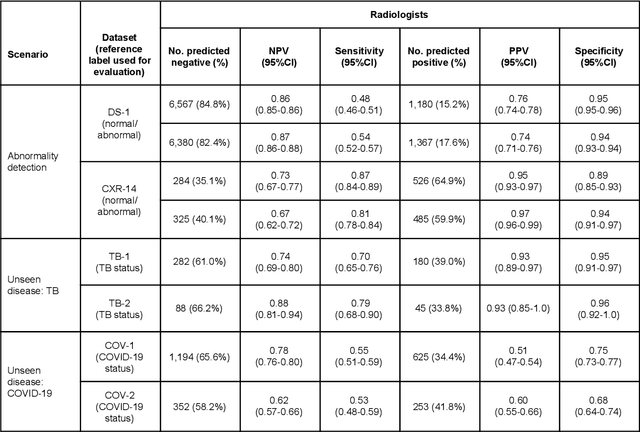
Abstract:Chest radiography (CXR) is the most widely-used thoracic clinical imaging modality and is crucial for guiding the management of cardiothoracic conditions. The detection of specific CXR findings has been the main focus of several artificial intelligence (AI) systems. However, the wide range of possible CXR abnormalities makes it impractical to build specific systems to detect every possible condition. In this work, we developed and evaluated an AI system to classify CXRs as normal or abnormal. For development, we used a de-identified dataset of 248,445 patients from a multi-city hospital network in India. To assess generalizability, we evaluated our system using 6 international datasets from India, China, and the United States. Of these datasets, 4 focused on diseases that the AI was not trained to detect: 2 datasets with tuberculosis and 2 datasets with coronavirus disease 2019. Our results suggest that the AI system generalizes to new patient populations and abnormalities. In a simulated workflow where the AI system prioritized abnormal cases, the turnaround time for abnormal cases reduced by 7-28%. These results represent an important step towards evaluating whether AI can be safely used to flag cases in a general setting where previously unseen abnormalities exist.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge