Georges El Fakhri
UniMRSeg: Unified Modality-Relax Segmentation via Hierarchical Self-Supervised Compensation
Sep 19, 2025Abstract:Multi-modal image segmentation faces real-world deployment challenges from incomplete/corrupted modalities degrading performance. While existing methods address training-inference modality gaps via specialized per-combination models, they introduce high deployment costs by requiring exhaustive model subsets and model-modality matching. In this work, we propose a unified modality-relax segmentation network (UniMRSeg) through hierarchical self-supervised compensation (HSSC). Our approach hierarchically bridges representation gaps between complete and incomplete modalities across input, feature and output levels. % First, we adopt modality reconstruction with the hybrid shuffled-masking augmentation, encouraging the model to learn the intrinsic modality characteristics and generate meaningful representations for missing modalities through cross-modal fusion. % Next, modality-invariant contrastive learning implicitly compensates the feature space distance among incomplete-complete modality pairs. Furthermore, the proposed lightweight reverse attention adapter explicitly compensates for the weak perceptual semantics in the frozen encoder. Last, UniMRSeg is fine-tuned under the hybrid consistency constraint to ensure stable prediction under all modality combinations without large performance fluctuations. Without bells and whistles, UniMRSeg significantly outperforms the state-of-the-art methods under diverse missing modality scenarios on MRI-based brain tumor segmentation, RGB-D semantic segmentation, RGB-D/T salient object segmentation. The code will be released at https://github.com/Xiaoqi-Zhao-DLUT/UniMRSeg.
Power Battery Detection
Aug 11, 2025



Abstract:Power batteries are essential components in electric vehicles, where internal structural defects can pose serious safety risks. We conduct a comprehensive study on a new task, power battery detection (PBD), which aims to localize the dense endpoints of cathode and anode plates from industrial X-ray images for quality inspection. Manual inspection is inefficient and error-prone, while traditional vision algorithms struggle with densely packed plates, low contrast, scale variation, and imaging artifacts. To address this issue and drive more attention into this meaningful task, we present PBD5K, the first large-scale benchmark for this task, consisting of 5,000 X-ray images from nine battery types with fine-grained annotations and eight types of real-world visual interference. To support scalable and consistent labeling, we develop an intelligent annotation pipeline that combines image filtering, model-assisted pre-labeling, cross-verification, and layered quality evaluation. We formulate PBD as a point-level segmentation problem and propose MDCNeXt, a model designed to extract and integrate multi-dimensional structure clues including point, line, and count information from the plate itself. To improve discrimination between plates and suppress visual interference, MDCNeXt incorporates two state space modules. The first is a prompt-filtered module that learns contrastive relationships guided by task-specific prompts. The second is a density-aware reordering module that refines segmentation in regions with high plate density. In addition, we propose a distance-adaptive mask generation strategy to provide robust supervision under varying spatial distributions of anode and cathode positions. The source code and datasets will be publicly available at \href{https://github.com/Xiaoqi-Zhao-DLUT/X-ray-PBD}{PBD5K}.
Dual Prompting for Diverse Count-level PET Denoising
May 05, 2025Abstract:The to-be-denoised positron emission tomography (PET) volumes are inherent with diverse count levels, which imposes challenges for a unified model to tackle varied cases. In this work, we resort to the recently flourished prompt learning to achieve generalizable PET denoising with different count levels. Specifically, we propose dual prompts to guide the PET denoising in a divide-and-conquer manner, i.e., an explicitly count-level prompt to provide the specific prior information and an implicitly general denoising prompt to encode the essential PET denoising knowledge. Then, a novel prompt fusion module is developed to unify the heterogeneous prompts, followed by a prompt-feature interaction module to inject prompts into the features. The prompts are able to dynamically guide the noise-conditioned denoising process. Therefore, we are able to efficiently train a unified denoising model for various count levels, and deploy it to different cases with personalized prompts. We evaluated on 1940 low-count PET 3D volumes with uniformly randomly selected 13-22\% fractions of events from 97 $^{18}$F-MK6240 tau PET studies. It shows our dual prompting can largely improve the performance with informed count-level and outperform the count-conditional model.
Mixture-of-Shape-Experts (MoSE): End-to-End Shape Dictionary Framework to Prompt SAM for Generalizable Medical Segmentation
Apr 13, 2025



Abstract:Single domain generalization (SDG) has recently attracted growing attention in medical image segmentation. One promising strategy for SDG is to leverage consistent semantic shape priors across different imaging protocols, scanner vendors, and clinical sites. However, existing dictionary learning methods that encode shape priors often suffer from limited representational power with a small set of offline computed shape elements, or overfitting when the dictionary size grows. Moreover, they are not readily compatible with large foundation models such as the Segment Anything Model (SAM). In this paper, we propose a novel Mixture-of-Shape-Experts (MoSE) framework that seamlessly integrates the idea of mixture-of-experts (MoE) training into dictionary learning to efficiently capture diverse and robust shape priors. Our method conceptualizes each dictionary atom as a shape expert, which specializes in encoding distinct semantic shape information. A gating network dynamically fuses these shape experts into a robust shape map, with sparse activation guided by SAM encoding to prevent overfitting. We further provide this shape map as a prompt to SAM, utilizing the powerful generalization capability of SAM through bidirectional integration. All modules, including the shape dictionary, are trained in an end-to-end manner. Extensive experiments on multiple public datasets demonstrate its effectiveness.
Anatomically and Metabolically Informed Diffusion for Unified Denoising and Segmentation in Low-Count PET Imaging
Mar 17, 2025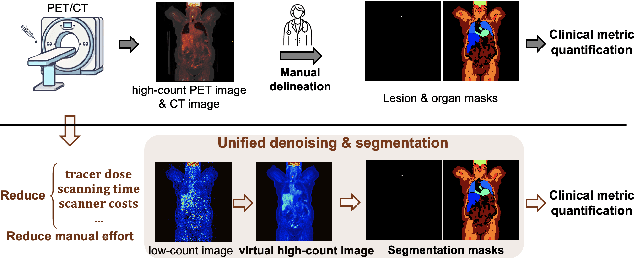
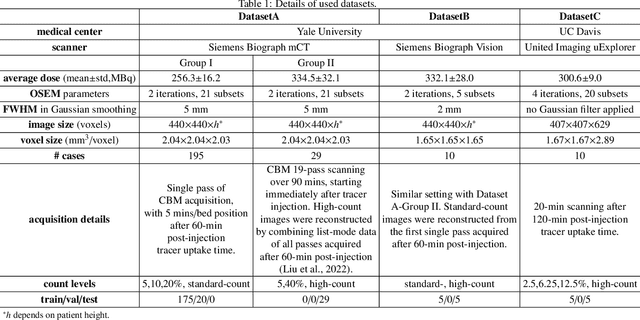
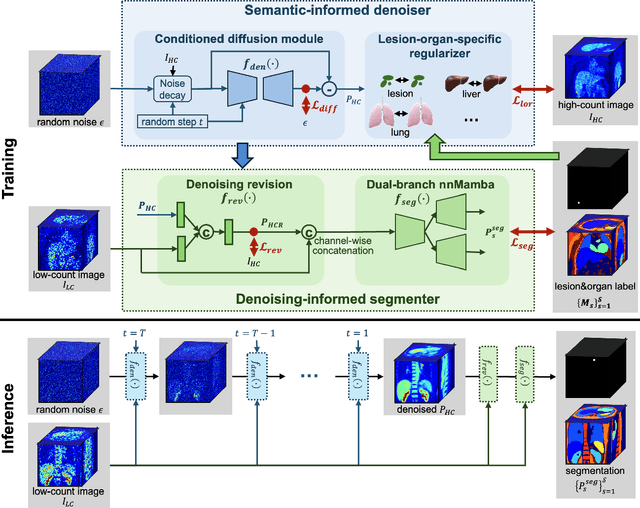
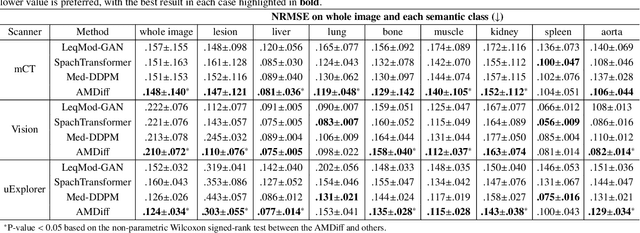
Abstract:Positron emission tomography (PET) image denoising, along with lesion and organ segmentation, are critical steps in PET-aided diagnosis. However, existing methods typically treat these tasks independently, overlooking inherent synergies between them as correlated steps in the analysis pipeline. In this work, we present the anatomically and metabolically informed diffusion (AMDiff) model, a unified framework for denoising and lesion/organ segmentation in low-count PET imaging. By integrating multi-task functionality and exploiting the mutual benefits of these tasks, AMDiff enables direct quantification of clinical metrics, such as total lesion glycolysis (TLG), from low-count inputs. The AMDiff model incorporates a semantic-informed denoiser based on diffusion strategy and a denoising-informed segmenter utilizing nnMamba architecture. The segmenter constrains denoised outputs via a lesion-organ-specific regularizer, while the denoiser enhances the segmenter by providing enriched image information through a denoising revision module. These components are connected via a warming-up mechanism to optimize multitask interactions. Experiments on multi-vendor, multi-center, and multi-noise-level datasets demonstrate the superior performance of AMDiff. For test cases below 20% of the clinical count levels from participating sites, AMDiff achieves TLG quantification biases of -26.98%, outperforming its ablated versions which yield biases of -35.85% (without the lesion-organ-specific regularizer) and -40.79% (without the denoising revision module).
Inspiring the Next Generation of Segment Anything Models: Comprehensively Evaluate SAM and SAM 2 with Diverse Prompts Towards Context-Dependent Concepts under Different Scenes
Dec 02, 2024Abstract:As a foundational model, SAM has significantly influenced multiple fields within computer vision, and its upgraded version, SAM 2, enhances capabilities in video segmentation, poised to make a substantial impact once again. While SAMs (SAM and SAM 2) have demonstrated excellent performance in segmenting context-independent concepts like people, cars, and roads, they overlook more challenging context-dependent (CD) concepts, such as visual saliency, camouflage, product defects, and medical lesions. CD concepts rely heavily on global and local contextual information, making them susceptible to shifts in different contexts, which requires strong discriminative capabilities from the model. The lack of comprehensive evaluation of SAMs limits understanding of their performance boundaries, which may hinder the design of future models. In this paper, we conduct a thorough quantitative evaluation of SAMs on 11 CD concepts across 2D and 3D images and videos in various visual modalities within natural, medical, and industrial scenes. We develop a unified evaluation framework for SAM and SAM 2 that supports manual, automatic, and intermediate self-prompting, aided by our specific prompt generation and interaction strategies. We further explore the potential of SAM 2 for in-context learning and introduce prompt robustness testing to simulate real-world imperfect prompts. Finally, we analyze the benefits and limitations of SAMs in understanding CD concepts and discuss their future development in segmentation tasks. This work aims to provide valuable insights to guide future research in both context-independent and context-dependent concepts segmentation, potentially informing the development of the next version - SAM 3.
Free-breathing 3D cardiac extracellular volume (ECV) mapping using a linear tangent space alignment (LTSA) model
Aug 22, 2024



Abstract:$\textbf{Purpose:}$ To develop a new method for free-breathing 3D extracellular volume (ECV) mapping of the whole heart at 3T. $\textbf{Methods:}$ A free-breathing 3D cardiac ECV mapping method was developed at 3T. T1 mapping was performed before and after contrast agent injection using a free-breathing ECG-gated inversion-recovery sequence with spoiled gradient echo readout. A linear tangent space alignment (LTSA) model-based method was used to reconstruct high-frame-rate dynamic images from (k,t)-space data sparsely sampled along a random stack-of-stars trajectory. Joint T1 and transmit B1 estimation was performed voxel-by-voxel for pre- and post-contrast T1 mapping. To account for the time-varying T1 after contrast agent injection, a linearly time-varying T1 model was introduced for post-contrast T1 mapping. ECV maps were generated by aligning pre- and post-contrast T1 maps through affine transformation. $\textbf{Results:}$ The feasibility of the proposed method was demonstrated using in vivo studies with six healthy volunteers at 3T. We obtained 3D ECV maps at a spatial resolution of 1.9$\times$1.9$\times$4.5 $mm^{3}$ and a FOV of 308$\times$308$\times$144 $mm^{3}$, with a scan time of 10.1$\pm$1.4 and 10.6$\pm$1.6 min before and after contrast agent injection, respectively. The ECV maps and the pre- and post-contrast T1 maps obtained by the proposed method were in good agreement with the 2D MOLLI method both qualitatively and quantitatively. $\textbf{Conclusion:}$ The proposed method allows for free-breathing 3D ECV mapping of the whole heart within a practically feasible imaging time. The estimated ECV values from the proposed method were comparable to those from the existing method. $\textbf{Keywords:}$ cardiac extracellular volume (ECV) mapping, cardiac T1 mapping, linear tangent space alignment (LTSA), manifold learning
Point-supervised Brain Tumor Segmentation with Box-prompted MedSAM
Aug 01, 2024

Abstract:Delineating lesions and anatomical structure is important for image-guided interventions. Point-supervised medical image segmentation (PSS) has great potential to alleviate costly expert delineation labeling. However, due to the lack of precise size and boundary guidance, the effectiveness of PSS often falls short of expectations. Although recent vision foundational models, such as the medical segment anything model (MedSAM), have made significant advancements in bounding-box-prompted segmentation, it is not straightforward to utilize point annotation, and is prone to semantic ambiguity. In this preliminary study, we introduce an iterative framework to facilitate semantic-aware point-supervised MedSAM. Specifically, the semantic box-prompt generator (SBPG) module has the capacity to convert the point input into potential pseudo bounding box suggestions, which are explicitly refined by the prototype-based semantic similarity. This is then succeeded by a prompt-guided spatial refinement (PGSR) module that harnesses the exceptional generalizability of MedSAM to infer the segmentation mask, which also updates the box proposal seed in SBPG. Performance can be progressively improved with adequate iterations. We conducted an evaluation on BraTS2018 for the segmentation of whole brain tumors and demonstrated its superior performance compared to traditional PSS methods and on par with box-supervised methods.
Nuclear Medicine Artificial Intelligence in Action: The Bethesda Report (AI Summit 2024)
Jun 03, 2024
Abstract:The 2nd SNMMI Artificial Intelligence (AI) Summit, organized by the SNMMI AI Task Force, took place in Bethesda, MD, on February 29 - March 1, 2024. Bringing together various community members and stakeholders, and following up on a prior successful 2022 AI Summit, the summit theme was: AI in Action. Six key topics included (i) an overview of prior and ongoing efforts by the AI task force, (ii) emerging needs and tools for computational nuclear oncology, (iii) new frontiers in large language and generative models, (iv) defining the value proposition for the use of AI in nuclear medicine, (v) open science including efforts for data and model repositories, and (vi) issues of reimbursement and funding. The primary efforts, findings, challenges, and next steps are summarized in this manuscript.
Treatment-wise Glioblastoma Survival Inference with Multi-parametric Preoperative MRI
Feb 10, 2024Abstract:In this work, we aim to predict the survival time (ST) of glioblastoma (GBM) patients undergoing different treatments based on preoperative magnetic resonance (MR) scans. The personalized and precise treatment planning can be achieved by comparing the ST of different treatments. It is well established that both the current status of the patient (as represented by the MR scans) and the choice of treatment are the cause of ST. While previous related MR-based glioblastoma ST studies have focused only on the direct mapping of MR scans to ST, they have not included the underlying causal relationship between treatments and ST. To address this limitation, we propose a treatment-conditioned regression model for glioblastoma ST that incorporates treatment information in addition to MR scans. Our approach allows us to effectively utilize the data from all of the treatments in a unified manner, rather than having to train separate models for each of the treatments. Furthermore, treatment can be effectively injected into each convolutional layer through the adaptive instance normalization we employ. We evaluate our framework on the BraTS20 ST prediction task. Three treatment options are considered: Gross Total Resection (GTR), Subtotal Resection (STR), and no resection. The evaluation results demonstrate the effectiveness of injecting the treatment for estimating GBM survival.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge