Fenglin Liu
Key Laboratory of Optoelectronic Technology and Systems, Ministry of Education, Chongqing University, Chongqing 400044, China
Competence-based Multimodal Curriculum Learning for Medical Report Generation
Jun 24, 2022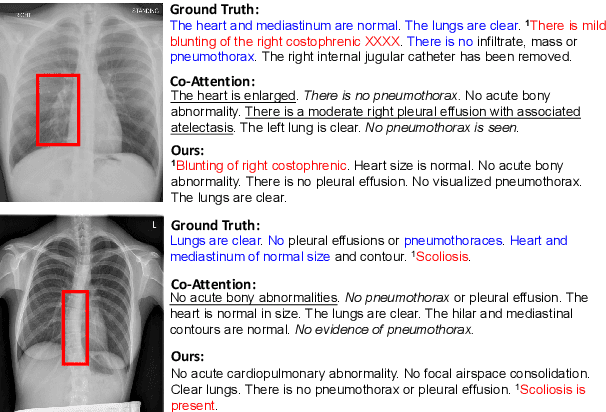
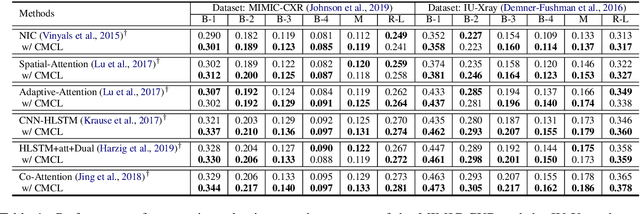
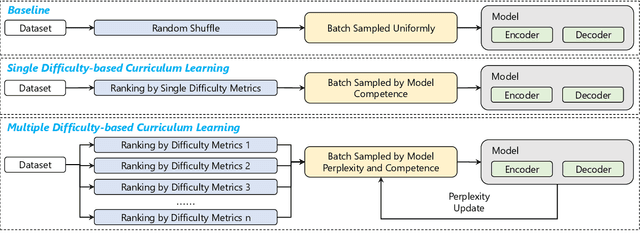
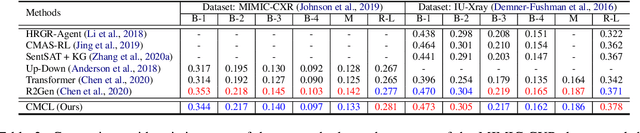
Abstract:Medical report generation task, which targets to produce long and coherent descriptions of medical images, has attracted growing research interests recently. Different from the general image captioning tasks, medical report generation is more challenging for data-driven neural models. This is mainly due to 1) the serious data bias and 2) the limited medical data. To alleviate the data bias and make best use of available data, we propose a Competence-based Multimodal Curriculum Learning framework (CMCL). Specifically, CMCL simulates the learning process of radiologists and optimizes the model in a step by step manner. Firstly, CMCL estimates the difficulty of each training instance and evaluates the competence of current model; Secondly, CMCL selects the most suitable batch of training instances considering current model competence. By iterating above two steps, CMCL can gradually improve the model's performance. The experiments on the public IU-Xray and MIMIC-CXR datasets show that CMCL can be incorporated into existing models to improve their performance.
Graph-in-Graph Network for Automatic Gene Ontology Description Generation
Jun 10, 2022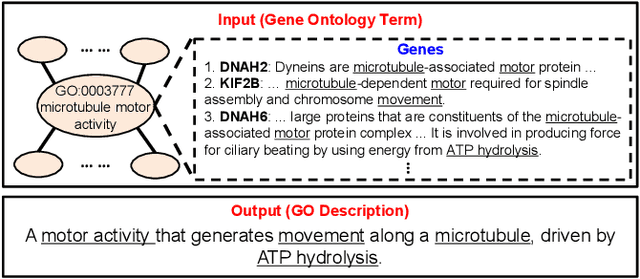
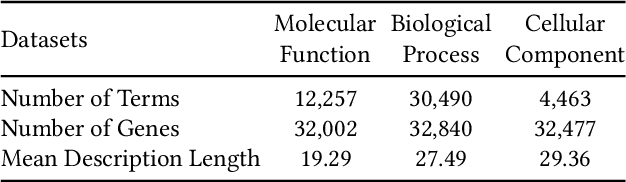
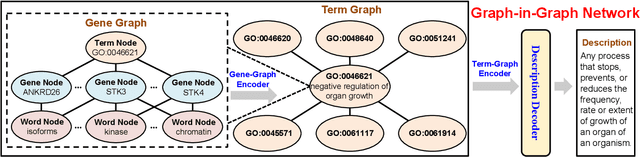
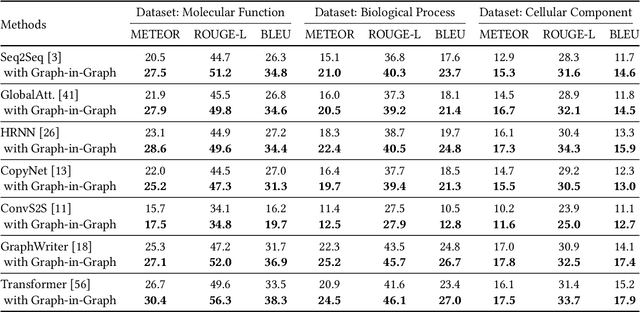
Abstract:Gene Ontology (GO) is the primary gene function knowledge base that enables computational tasks in biomedicine. The basic element of GO is a term, which includes a set of genes with the same function. Existing research efforts of GO mainly focus on predicting gene term associations. Other tasks, such as generating descriptions of new terms, are rarely pursued. In this paper, we propose a novel task: GO term description generation. This task aims to automatically generate a sentence that describes the function of a GO term belonging to one of the three categories, i.e., molecular function, biological process, and cellular component. To address this task, we propose a Graph-in-Graph network that can efficiently leverage the structural information of GO. The proposed network introduces a two-layer graph: the first layer is a graph of GO terms where each node is also a graph (gene graph). Such a Graph-in-Graph network can derive the biological functions of GO terms and generate proper descriptions. To validate the effectiveness of the proposed network, we build three large-scale benchmark datasets. By incorporating the proposed Graph-in-Graph network, the performances of seven different sequence-to-sequence models can be substantially boosted across all evaluation metrics, with up to 34.7%, 14.5%, and 39.1% relative improvements in BLEU, ROUGE-L, and METEOR, respectively.
End-to-end Spoken Conversational Question Answering: Task, Dataset and Model
Apr 29, 2022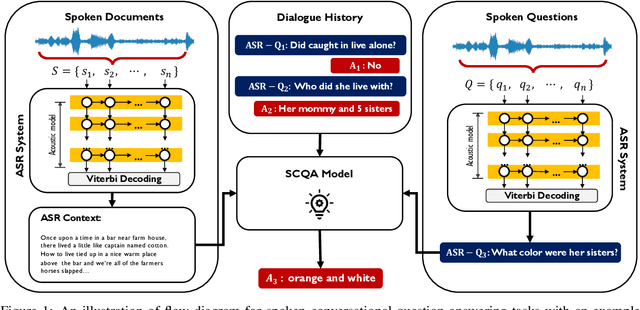
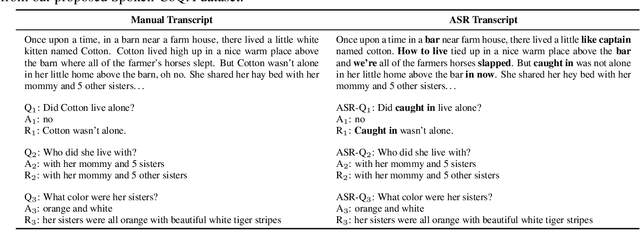
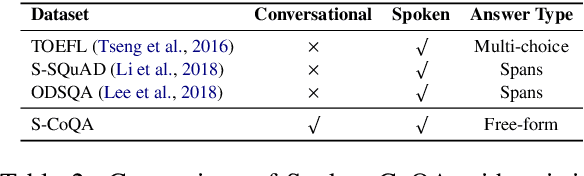

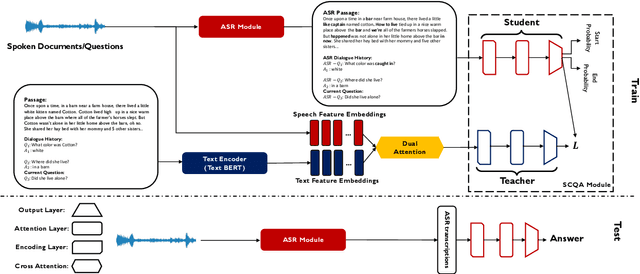

Abstract:In spoken question answering, the systems are designed to answer questions from contiguous text spans within the related speech transcripts. However, the most natural way that human seek or test their knowledge is via human conversations. Therefore, we propose a new Spoken Conversational Question Answering task (SCQA), aiming at enabling the systems to model complex dialogue flows given the speech documents. In this task, our main objective is to build the system to deal with conversational questions based on the audio recordings, and to explore the plausibility of providing more cues from different modalities with systems in information gathering. To this end, instead of directly adopting automatically generated speech transcripts with highly noisy data, we propose a novel unified data distillation approach, DDNet, which effectively ingests cross-modal information to achieve fine-grained representations of the speech and language modalities. Moreover, we propose a simple and novel mechanism, termed Dual Attention, by encouraging better alignments between audio and text to ease the process of knowledge transfer. To evaluate the capacity of SCQA systems in a dialogue-style interaction, we assemble a Spoken Conversational Question Answering (Spoken-CoQA) dataset with more than 40k question-answer pairs from 4k conversations. The performance of the existing state-of-the-art methods significantly degrade on our dataset, hence demonstrating the necessity of cross-modal information integration. Our experimental results demonstrate that our proposed method achieves superior performance in spoken conversational question answering tasks.
AlignTransformer: Hierarchical Alignment of Visual Regions and Disease Tags for Medical Report Generation
Mar 18, 2022
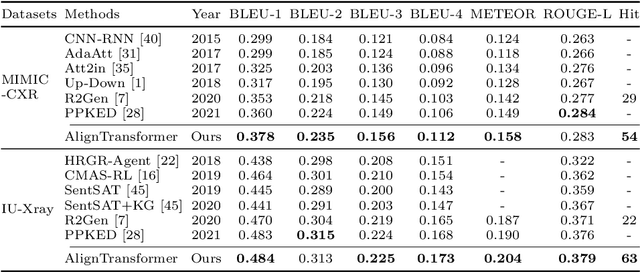
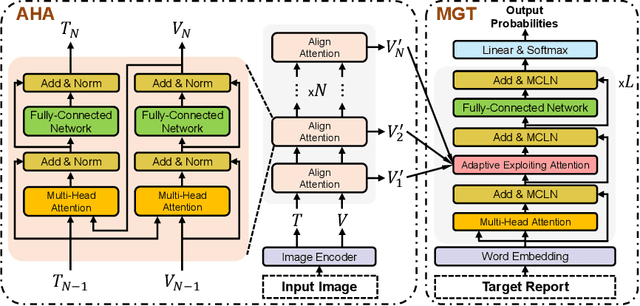
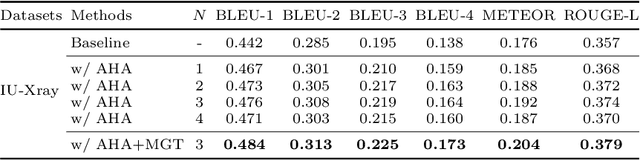
Abstract:Recently, medical report generation, which aims to automatically generate a long and coherent descriptive paragraph of a given medical image, has received growing research interests. Different from the general image captioning tasks, medical report generation is more challenging for data-driven neural models. This is mainly due to 1) the serious data bias: the normal visual regions dominate the dataset over the abnormal visual regions, and 2) the very long sequence. To alleviate above two problems, we propose an AlignTransformer framework, which includes the Align Hierarchical Attention (AHA) and the Multi-Grained Transformer (MGT) modules: 1) AHA module first predicts the disease tags from the input image and then learns the multi-grained visual features by hierarchically aligning the visual regions and disease tags. The acquired disease-grounded visual features can better represent the abnormal regions of the input image, which could alleviate data bias problem; 2) MGT module effectively uses the multi-grained features and Transformer framework to generate the long medical report. The experiments on the public IU-Xray and MIMIC-CXR datasets show that the AlignTransformer can achieve results competitive with state-of-the-art methods on the two datasets. Moreover, the human evaluation conducted by professional radiologists further proves the effectiveness of our approach.
Class-Aware Generative Adversarial Transformers for Medical Image Segmentation
Jan 28, 2022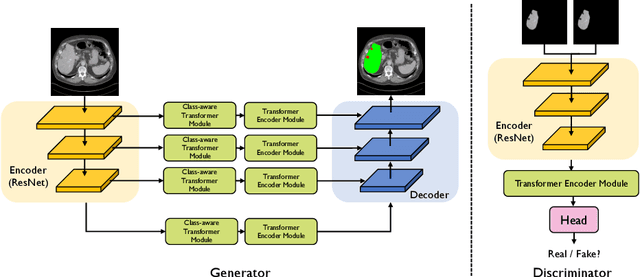
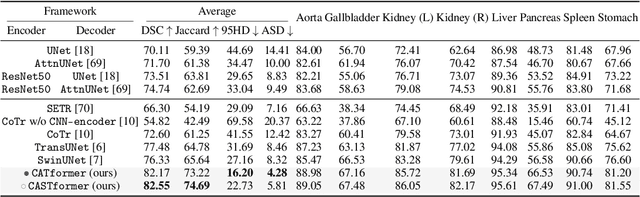
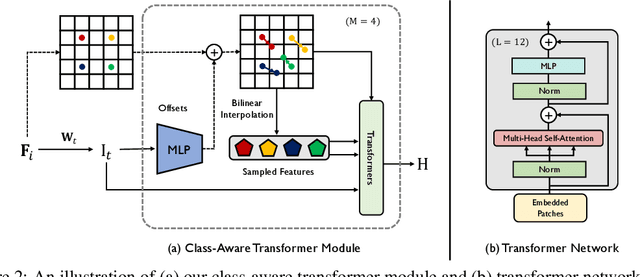
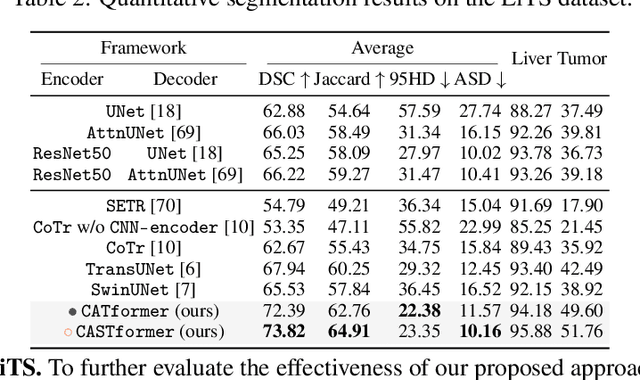
Abstract:Transformers have made remarkable progress towards modeling long-range dependencies within the medical image analysis domain. However, current transformer-based models suffer from several disadvantages: (1) existing methods fail to capture the important features of the images due to the naive tokenization scheme; (2) the models suffer from information loss because they only consider single-scale feature representations; and (3) the segmentation label maps generated by the models are not accurate enough without considering rich semantic contexts and anatomical textures. In this work, we present CA-GANformer, a novel type of generative adversarial transformers, for medical image segmentation. First, we take advantage of the pyramid structure to construct multi-scale representations and handle multi-scale variations. We then design a novel class-aware transformer module to better learn the discriminative regions of objects with semantic structures. Lastly, we utilize an adversarial training strategy that boosts segmentation accuracy and correspondingly allows a transformer-based discriminator to capture high-level semantically correlated contents and low-level anatomical features. Our experiments demonstrate that CA-GANformer dramatically outperforms previous state-of-the-art transformer-based approaches on three benchmarks, obtaining 2.54%-5.88% absolute improvements in Dice over previous models. Further qualitative experiments provide a more detailed picture of the model's inner workings, shed light on the challenges in improved transparency, and demonstrate that transfer learning can greatly improve performance and reduce the size of medical image datasets in training, making CA-GANformer a strong starting point for downstream medical image analysis tasks. Codes and models will be available to the public.
Auto-Encoding Knowledge Graph for Unsupervised Medical Report Generation
Nov 15, 2021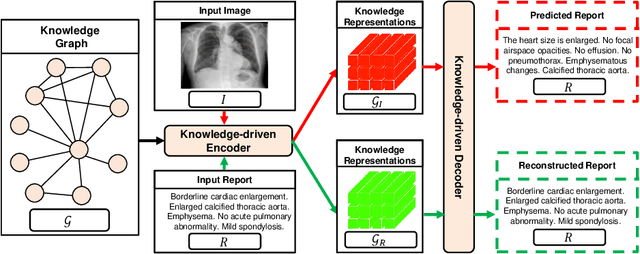
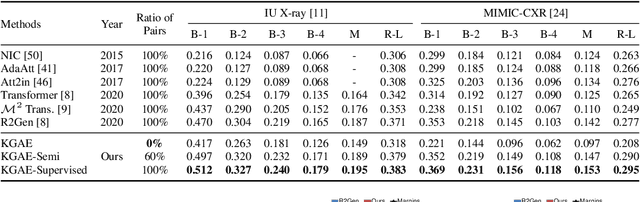
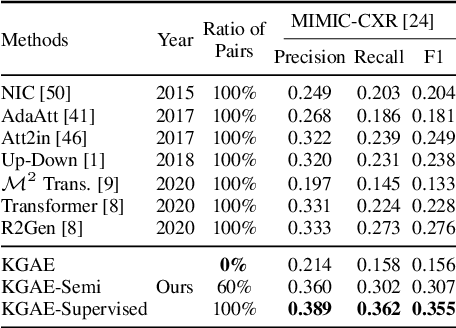
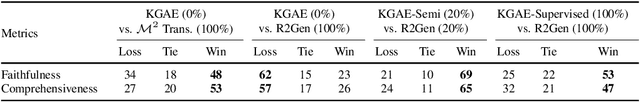
Abstract:Medical report generation, which aims to automatically generate a long and coherent report of a given medical image, has been receiving growing research interests. Existing approaches mainly adopt a supervised manner and heavily rely on coupled image-report pairs. However, in the medical domain, building a large-scale image-report paired dataset is both time-consuming and expensive. To relax the dependency on paired data, we propose an unsupervised model Knowledge Graph Auto-Encoder (KGAE) which accepts independent sets of images and reports in training. KGAE consists of a pre-constructed knowledge graph, a knowledge-driven encoder and a knowledge-driven decoder. The knowledge graph works as the shared latent space to bridge the visual and textual domains; The knowledge-driven encoder projects medical images and reports to the corresponding coordinates in this latent space and the knowledge-driven decoder generates a medical report given a coordinate in this space. Since the knowledge-driven encoder and decoder can be trained with independent sets of images and reports, KGAE is unsupervised. The experiments show that the unsupervised KGAE generates desirable medical reports without using any image-report training pairs. Moreover, KGAE can also work in both semi-supervised and supervised settings, and accept paired images and reports in training. By further fine-tuning with image-report pairs, KGAE consistently outperforms the current state-of-the-art models on two datasets.
O2NA: An Object-Oriented Non-Autoregressive Approach for Controllable Video Captioning
Aug 05, 2021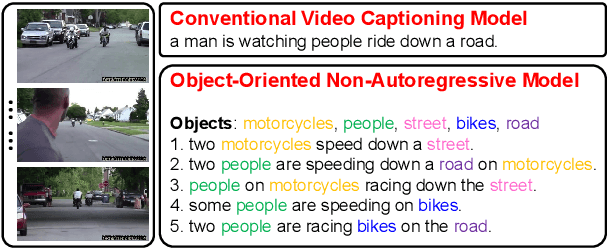
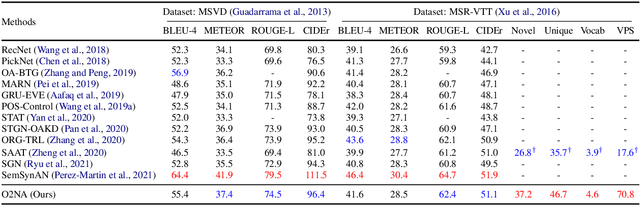
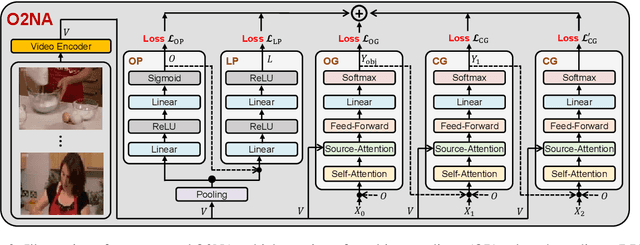
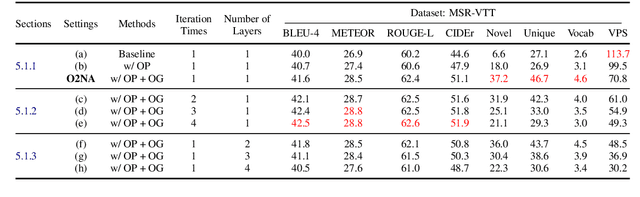
Abstract:Video captioning combines video understanding and language generation. Different from image captioning that describes a static image with details of almost every object, video captioning usually considers a sequence of frames and biases towards focused objects, e.g., the objects that stay in focus regardless of the changing background. Therefore, detecting and properly accommodating focused objects is critical in video captioning. To enforce the description of focused objects and achieve controllable video captioning, we propose an Object-Oriented Non-Autoregressive approach (O2NA), which performs caption generation in three steps: 1) identify the focused objects and predict their locations in the target caption; 2) generate the related attribute words and relation words of these focused objects to form a draft caption; and 3) combine video information to refine the draft caption to a fluent final caption. Since the focused objects are generated and located ahead of other words, it is difficult to apply the word-by-word autoregressive generation process; instead, we adopt a non-autoregressive approach. The experiments on two benchmark datasets, i.e., MSR-VTT and MSVD, demonstrate the effectiveness of O2NA, which achieves results competitive with the state-of-the-arts but with both higher diversity and higher inference speed.
Audio-Oriented Multimodal Machine Comprehension: Task, Dataset and Model
Jul 04, 2021
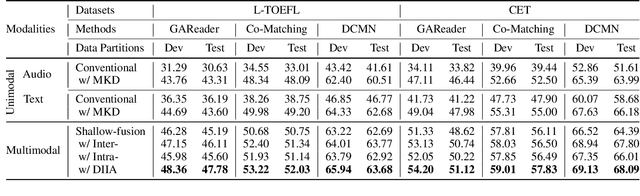
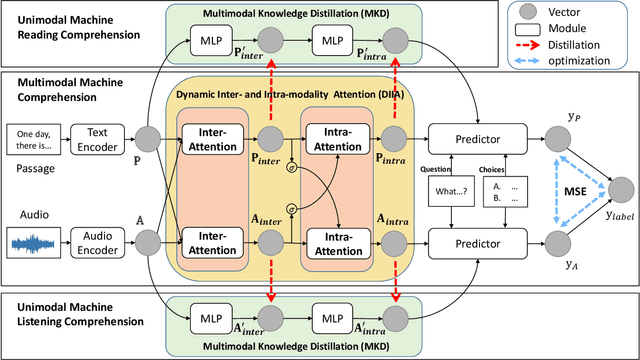
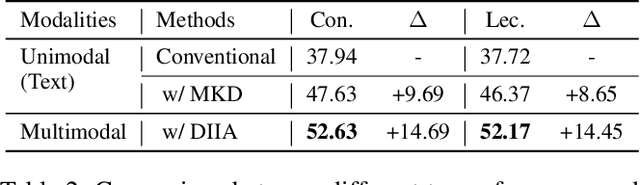
Abstract:While Machine Comprehension (MC) has attracted extensive research interests in recent years, existing approaches mainly belong to the category of Machine Reading Comprehension task which mines textual inputs (paragraphs and questions) to predict the answers (choices or text spans). However, there are a lot of MC tasks that accept audio input in addition to the textual input, e.g. English listening comprehension test. In this paper, we target the problem of Audio-Oriented Multimodal Machine Comprehension, and its goal is to answer questions based on the given audio and textual information. To solve this problem, we propose a Dynamic Inter- and Intra-modality Attention (DIIA) model to effectively fuse the two modalities (audio and textual). DIIA can work as an independent component and thus be easily integrated into existing MC models. Moreover, we further develop a Multimodal Knowledge Distillation (MKD) module to enable our multimodal MC model to accurately predict the answers based only on either the text or the audio. As a result, the proposed approach can handle various tasks including: Audio-Oriented Multimodal Machine Comprehension, Machine Reading Comprehension and Machine Listening Comprehension, in a single model, making fair comparisons possible between our model and the existing unimodal MC models. Experimental results and analysis prove the effectiveness of the proposed approaches. First, the proposed DIIA boosts the baseline models by up to 21.08% in terms of accuracy; Second, under the unimodal scenarios, the MKD module allows our multimodal MC model to significantly outperform the unimodal models by up to 18.87%, which are trained and tested with only audio or textual data.
Exploring and Distilling Posterior and Prior Knowledge for Radiology Report Generation
Jun 26, 2021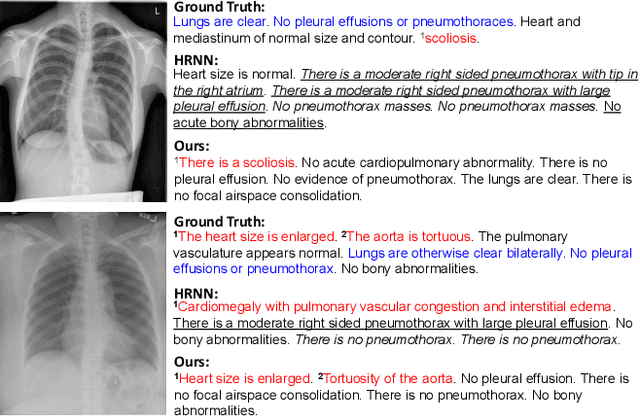
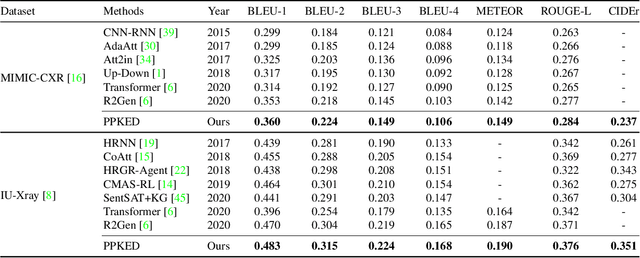
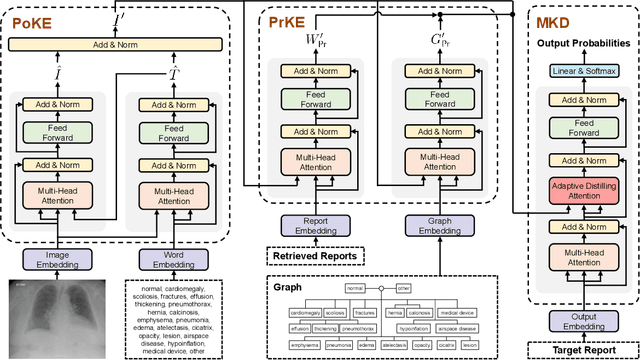
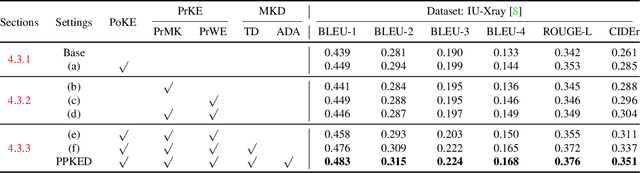
Abstract:Automatically generating radiology reports can improve current clinical practice in diagnostic radiology. On one hand, it can relieve radiologists from the heavy burden of report writing; On the other hand, it can remind radiologists of abnormalities and avoid the misdiagnosis and missed diagnosis. Yet, this task remains a challenging job for data-driven neural networks, due to the serious visual and textual data biases. To this end, we propose a Posterior-and-Prior Knowledge Exploring-and-Distilling approach (PPKED) to imitate the working patterns of radiologists, who will first examine the abnormal regions and assign the disease topic tags to the abnormal regions, and then rely on the years of prior medical knowledge and prior working experience accumulations to write reports. Thus, the PPKED includes three modules: Posterior Knowledge Explorer (PoKE), Prior Knowledge Explorer (PrKE) and Multi-domain Knowledge Distiller (MKD). In detail, PoKE explores the posterior knowledge, which provides explicit abnormal visual regions to alleviate visual data bias; PrKE explores the prior knowledge from the prior medical knowledge graph (medical knowledge) and prior radiology reports (working experience) to alleviate textual data bias. The explored knowledge is distilled by the MKD to generate the final reports. Evaluated on MIMIC-CXR and IU-Xray datasets, our method is able to outperform previous state-of-the-art models on these two datasets.
Exploring Semantic Relationships for Unpaired Image Captioning
Jun 20, 2021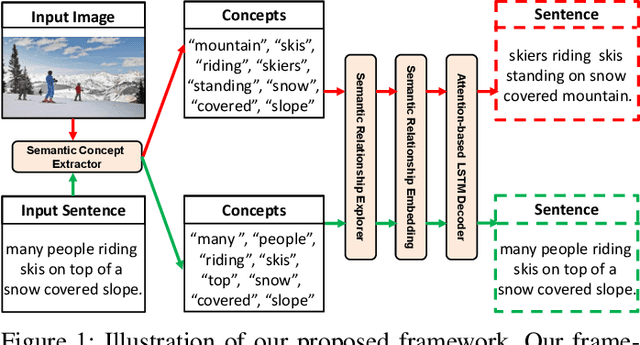
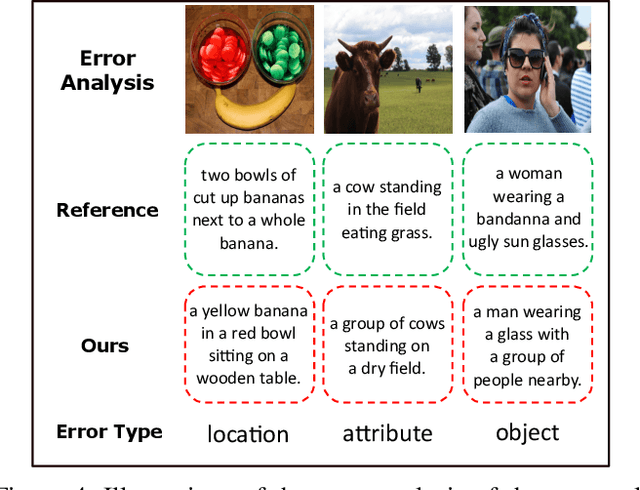
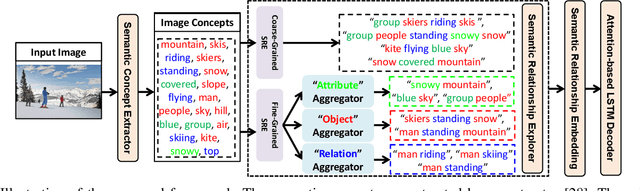
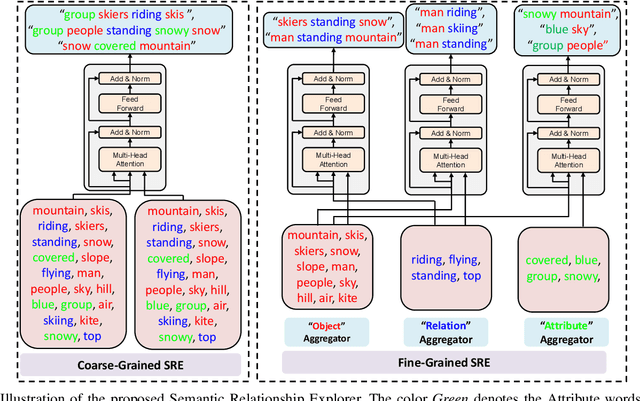

Abstract:Recently, image captioning has aroused great interest in both academic and industrial worlds. Most existing systems are built upon large-scale datasets consisting of image-sentence pairs, which, however, are time-consuming to construct. In addition, even for the most advanced image captioning systems, it is still difficult to realize deep image understanding. In this work, we achieve unpaired image captioning by bridging the vision and the language domains with high-level semantic information. The motivation stems from the fact that the semantic concepts with the same modality can be extracted from both images and descriptions. To further improve the quality of captions generated by the model, we propose the Semantic Relationship Explorer, which explores the relationships between semantic concepts for better understanding of the image. Extensive experiments on MSCOCO dataset show that we can generate desirable captions without paired datasets. Furthermore, the proposed approach boosts five strong baselines under the paired setting, where the most significant improvement in CIDEr score reaches 8%, demonstrating that it is effective and generalizes well to a wide range of models.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge