Chengjia Wang
Max-Fusion U-Net for Multi-Modal Pathology Segmentation with Attention and Dynamic Resampling
Sep 05, 2020



Abstract:Automatic segmentation of multi-sequence (multi-modal) cardiac MR (CMR) images plays a significant role in diagnosis and management for a variety of cardiac diseases. However, the performance of relevant algorithms is significantly affected by the proper fusion of the multi-modal information. Furthermore, particular diseases, such as myocardial infarction, display irregular shapes on images and occupy small regions at random locations. These facts make pathology segmentation of multi-modal CMR images a challenging task. In this paper, we present the Max-Fusion U-Net that achieves improved pathology segmentation performance given aligned multi-modal images of LGE, T2-weighted, and bSSFP modalities. Specifically, modality-specific features are extracted by dedicated encoders. Then they are fused with the pixel-wise maximum operator. Together with the corresponding encoding features, these representations are propagated to decoding layers with U-Net skip-connections. Furthermore, a spatial-attention module is applied in the last decoding layer to encourage the network to focus on those small semantically meaningful pathological regions that trigger relatively high responses by the network neurons. We also use a simple image patch extraction strategy to dynamically resample training examples with varying spacial and batch sizes. With limited GPU memory, this strategy reduces the imbalance of classes and forces the model to focus on regions around the interested pathology. It further improves segmentation accuracy and reduces the mis-classification of pathology. We evaluate our methods using the Myocardial pathology segmentation (MyoPS) combining the multi-sequence CMR dataset which involves three modalities. Extensive experiments demonstrate the effectiveness of the proposed model which outperforms the related baselines.
* 13 pages, 7 figures, conference paper
Annealing Genetic GAN for Minority Oversampling
Aug 05, 2020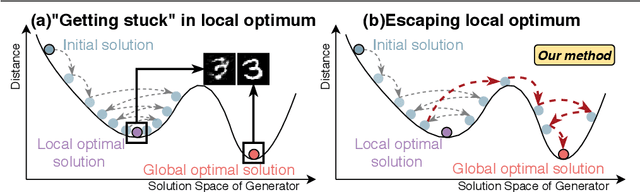
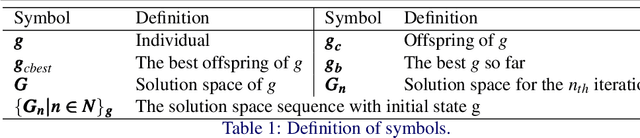

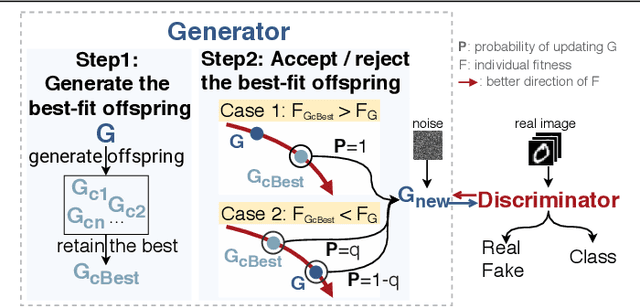
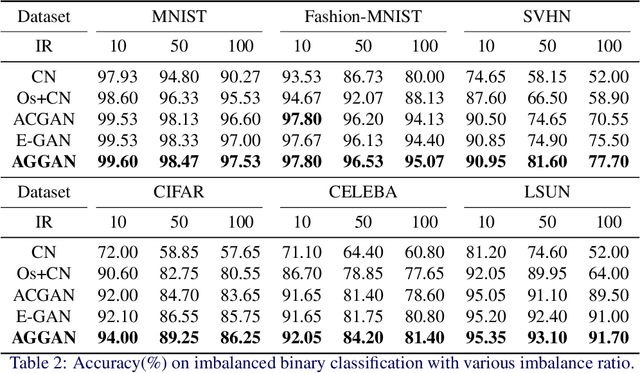
Abstract:The key to overcome class imbalance problems is to capture the distribution of minority class accurately. Generative Adversarial Networks (GANs) have shown some potentials to tackle class imbalance problems due to their capability of reproducing data distributions given ample training data samples. However, the scarce samples of one or more classes still pose a great challenge for GANs to learn accurate distributions for the minority classes. In this work, we propose an Annealing Genetic GAN (AGGAN) method, which aims to reproduce the distributions closest to the ones of the minority classes using only limited data samples. Our AGGAN renovates the training of GANs as an evolutionary process that incorporates the mechanism of simulated annealing. In particular, the generator uses different training strategies to generate multiple offspring and retain the best. Then, we use the Metropolis criterion in the simulated annealing to decide whether we should update the best offspring for the generator. As the Metropolis criterion allows a certain chance to accept the worse solutions, it enables our AGGAN steering away from the local optimum. According to both theoretical analysis and experimental studies on multiple imbalanced image datasets, we prove that the proposed training strategy can enable our AGGAN to reproduce the distributions of minority classes from scarce samples and provide an effective and robust solution for the class imbalance problem.
Deep Attentive Wasserstein Generative Adversarial Networks for MRI Reconstruction with Recurrent Context-Awareness
Jun 23, 2020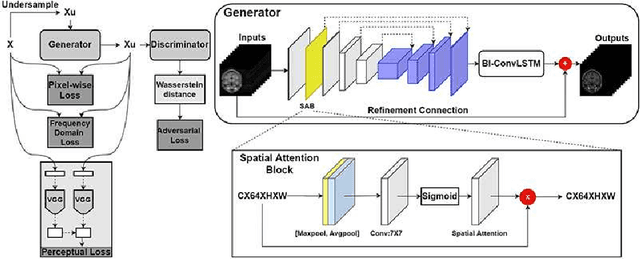
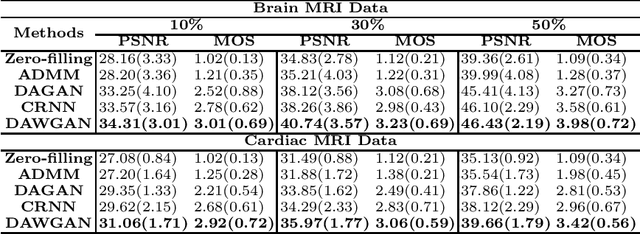
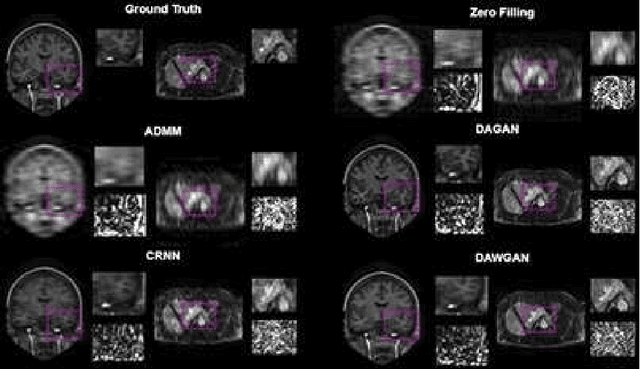
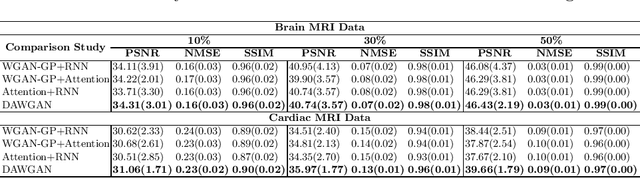
Abstract:The performance of traditional compressive sensing-based MRI (CS-MRI) reconstruction is affected by its slow iterative procedure and noise-induced artefacts. Although many deep learning-based CS-MRI methods have been proposed to mitigate the problems of traditional methods, they have not been able to achieve more robust results at higher acceleration factors. Most of the deep learning-based CS-MRI methods still can not fully mine the information from the k-space, which leads to unsatisfactory results in the MRI reconstruction. In this study, we propose a new deep learning-based CS-MRI reconstruction method to fully utilise the relationship among sequential MRI slices by coupling Wasserstein Generative Adversarial Networks (WGAN) with Recurrent Neural Networks. Further development of an attentive unit enables our model to reconstruct more accurate anatomical structures for the MRI data. By experimenting on different MRI datasets, we have demonstrated that our method can not only achieve better results compared to the state-of-the-arts but can also effectively reduce residual noise generated during the reconstruction process.
Learning to synthesise the ageing brain without longitudinal data
Dec 12, 2019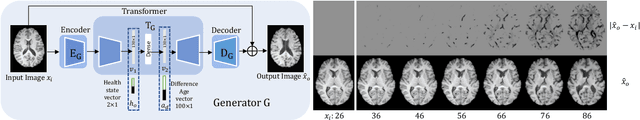
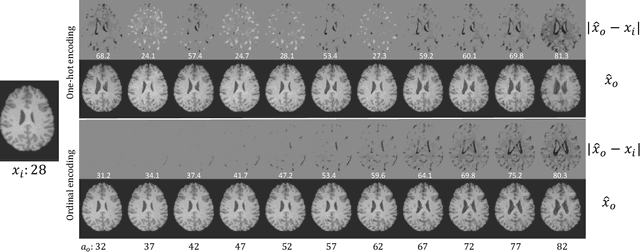
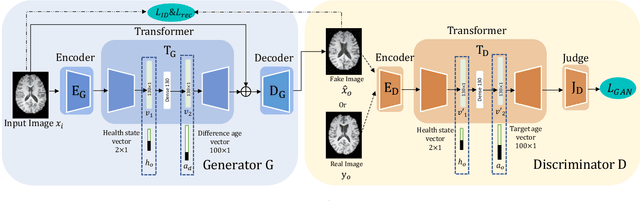
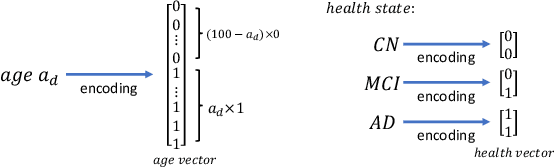
Abstract:Brain ageing is a continuous process that is affected by many factors including neurodegenerative diseases. Understanding this process is of great value for both neuroscience research and clinical applications. However, revealing underlying mechanisms is challenging due to the lack of longitudinal data. In this paper, we propose a deep learning-based method that learns to simulate subject-specific brain ageing trajectories without relying on longitudinal data. Our method synthesises aged images using a network conditioned on two clinical variables: age as a continuous variable, and health state, i.e. status of Alzheimer's Disease (AD) for this work, as an ordinal variable. We adopt an adversarial loss to learn the joint distribution of brain appearance and clinical variables and define reconstruction losses that help preserve subject identity. To demonstrate our model, we compare with several approaches using two widely used datasets: Cam-CAN and ADNI. We use ground-truth longitudinal data from ADNI to evaluate the quality of synthesised images. A pre-trained age predictor, which estimates the apparent age of a brain image, is used to assess age accuracy. In addition, we show that we can train the model on Cam-CAN data and evaluate on the longitudinal data from ADNI, indicating the generalisation power of our approach. Both qualitative and quantitative results show that our method can progressively simulate the ageing process by synthesising realistic brain images. The code will be made publicly available at: https://github.com/xiat0616/BrainAgeing.
Disentangle, align and fuse for multimodal and zero-shot image segmentation
Nov 12, 2019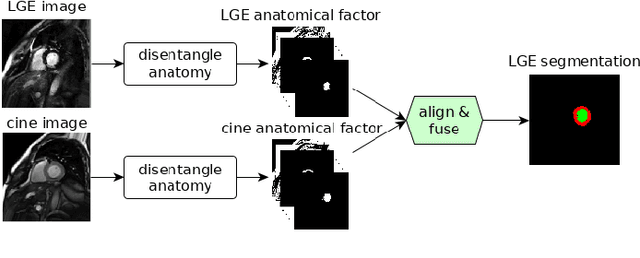
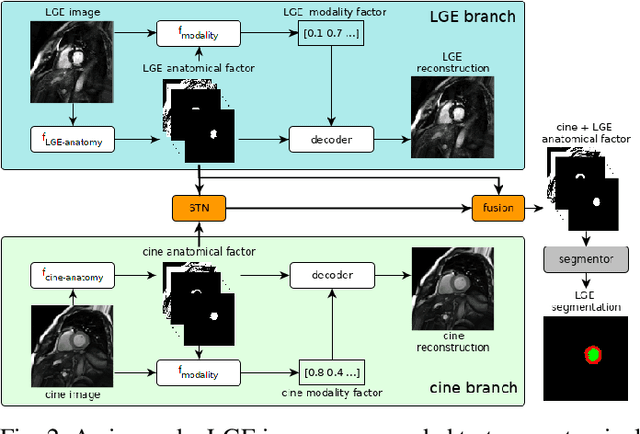
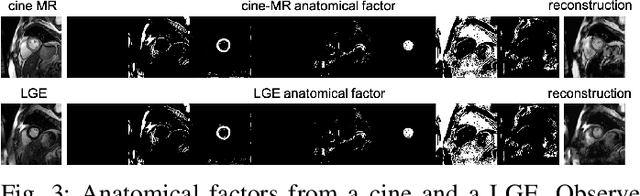

Abstract:Magnetic resonance (MR) protocols rely on several sequences to properly assess pathology and organ status. Yet, despite advances in image analysis we tend to treat each sequence, here termed modality, in isolation. Taking advantage of the information shared between modalities (largely an organ's anatomy) is beneficial for multi-modality multi-input processing and learning. However, we must overcome inherent anatomical misregistrations and disparities in signal intensity across the modalities to claim this benefit. We present a method that offers improved segmentation accuracy of the modality of interest (over a single input model), by learning to leverage information present in other modalities, enabling semi-supervised and zero shot learning. Core to our method is learning a disentangled decomposition into anatomical and imaging factors. Shared anatomical factors from the different inputs are jointly processed and fused to extract more accurate segmentation masks. Image misregistrations are corrected with a Spatial Transformer Network, that non-linearly aligns the anatomical factors. The imaging factor captures signal intensity characteristics across different modality data, and is used for image reconstruction, enabling semi-supervised learning. Temporal and slice pairing between inputs are learned dynamically. We demonstrate applications in Late Gadolinium Enhanced (LGE) and Blood Oxygenation Level Dependent (BOLD) cardiac segmentation, as well as in T2 abdominal segmentation.
Recurrent Aggregation Learning for Multi-View Echocardiographic Sequences Segmentation
Jul 24, 2019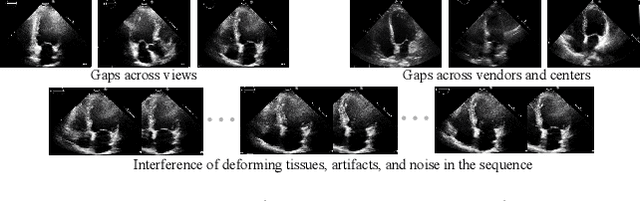
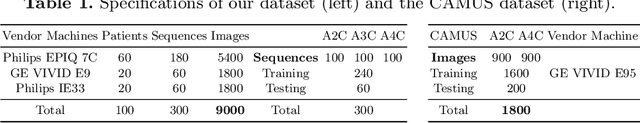
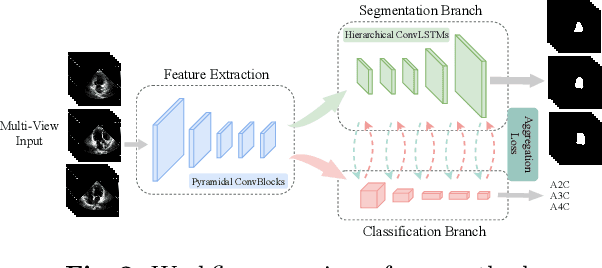
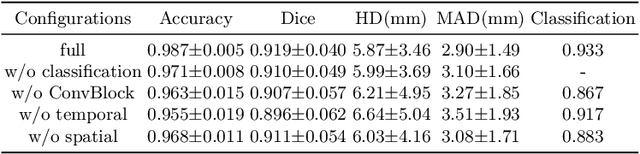
Abstract:Multi-view echocardiographic sequences segmentation is crucial for clinical diagnosis. However, this task is challenging due to limited labeled data, huge noise, and large gaps across views. Here we propose a recurrent aggregation learning method to tackle this challenging task. By pyramid ConvBlocks, multi-level and multi-scale features are extracted efficiently. Hierarchical ConvLSTMs next fuse these features and capture spatial-temporal information in multi-level and multi-scale space. We further introduce a double-branch aggregation mechanism for segmentation and classification which are mutually promoted by deep aggregation of multi-level and multi-scale features. The segmentation branch provides information to guide the classification while the classification branch affords multi-view regularization to refine segmentations and further lessen gaps across views. Our method is built as an end-to-end framework for segmentation and classification. Adequate experiments on our multi-view dataset (9000 labeled images) and the CAMUS dataset (1800 labeled images) corroborate that our method achieves not only superior segmentation and classification accuracy but also prominent temporal stability.
FIRE: Unsupervised bi-directional inter-modality registration using deep networks
Jul 11, 2019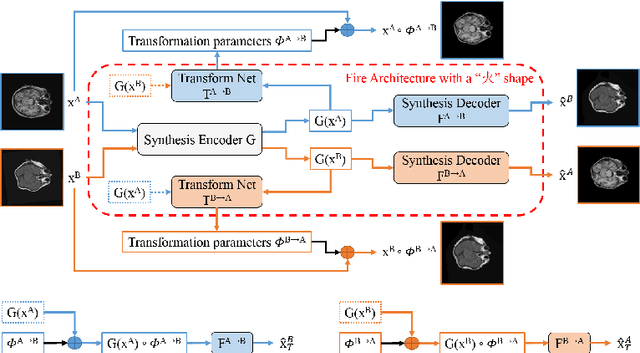
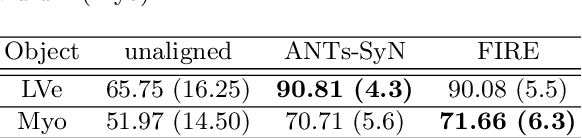
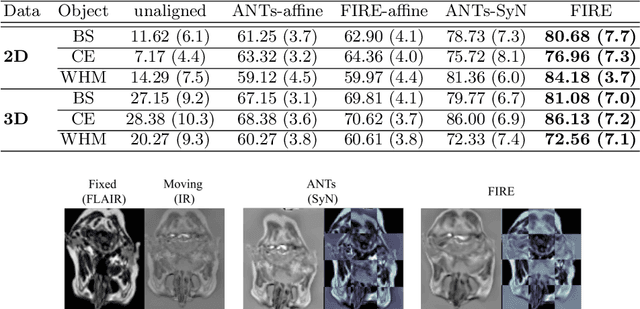
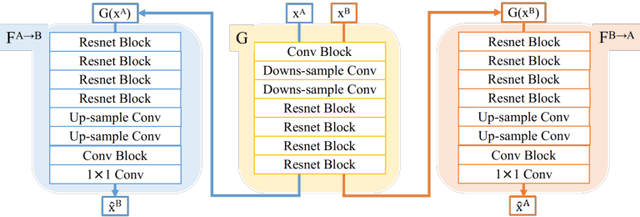

Abstract:Inter-modality image registration is an critical preprocessing step for many applications within the routine clinical pathway. This paper presents an unsupervised deep inter-modality registration network that can learn the optimal affine and non-rigid transformations simultaneously. Inverse-consistency is an important property commonly ignored in recent deep learning based inter-modality registration algorithms. We address this issue through the proposed multi-task architecture and the new comprehensive transformation network. Specifically, the proposed model learns a modality-independent latent representation to perform cycle-consistent cross-modality synthesis, and use an inverse-consistent loss to learn a pair of transformations to align the synthesized image with the target. We name this proposed framework as FIRE due to the shape of its structure. Our method shows comparable and better performances with the popular baseline method in experiments on multi-sequence brain MR data and intra-modality 4D cardiac Cine-MR data.
Evaluation of Algorithms for Multi-Modality Whole Heart Segmentation: An Open-Access Grand Challenge
Feb 21, 2019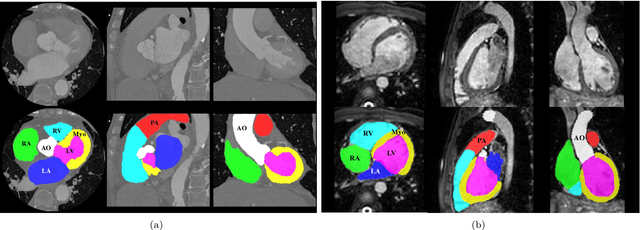
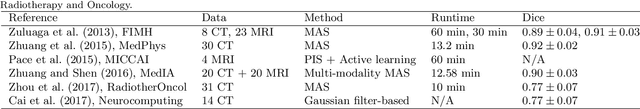

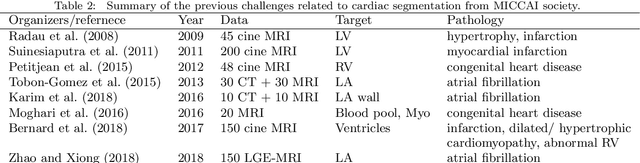
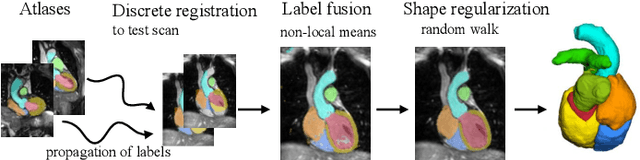
Abstract:Knowledge of whole heart anatomy is a prerequisite for many clinical applications. Whole heart segmentation (WHS), which delineates substructures of the heart, can be very valuable for modeling and analysis of the anatomy and functions of the heart. However, automating this segmentation can be arduous due to the large variation of the heart shape, and different image qualities of the clinical data. To achieve this goal, a set of training data is generally needed for constructing priors or for training. In addition, it is difficult to perform comparisons between different methods, largely due to differences in the datasets and evaluation metrics used. This manuscript presents the methodologies and evaluation results for the WHS algorithms selected from the submissions to the Multi-Modality Whole Heart Segmentation (MM-WHS) challenge, in conjunction with MICCAI 2017. The challenge provides 120 three-dimensional cardiac images covering the whole heart, including 60 CT and 60 MRI volumes, all acquired in clinical environments with manual delineation. Ten algorithms for CT data and eleven algorithms for MRI data, submitted from twelve groups, have been evaluated. The results show that many of the deep learning (DL) based methods achieved high accuracy, even though the number of training datasets was limited. A number of them also reported poor results in the blinded evaluation, probably due to overfitting in their training. The conventional algorithms, mainly based on multi-atlas segmentation, demonstrated robust and stable performance, even though the accuracy is not as good as the best DL method in CT segmentation. The challenge, including the provision of the annotated training data and the blinded evaluation for submitted algorithms on the test data, continues as an ongoing benchmarking resource via its homepage (\url{www.sdspeople.fudan.edu.cn/zhuangxiahai/0/mmwhs/}).
Unsupervised learning for cross-domain medical image synthesis using deformation invariant cycle consistency networks
Aug 12, 2018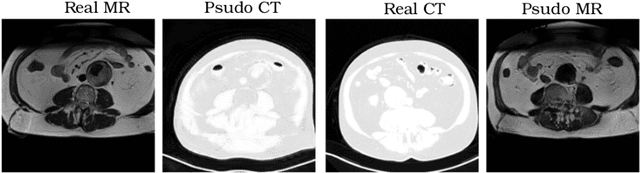
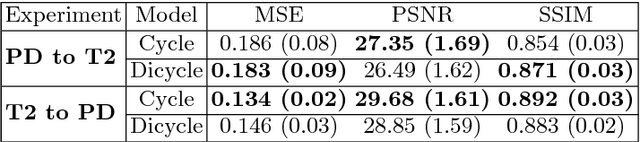
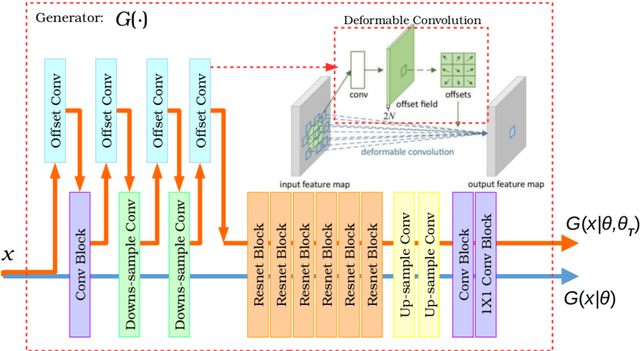
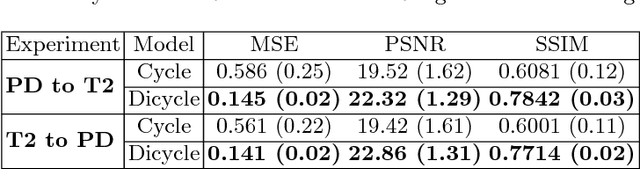
Abstract:Recently, the cycle-consistent generative adversarial networks (CycleGAN) has been widely used for synthesis of multi-domain medical images. The domain-specific nonlinear deformations captured by CycleGAN make the synthesized images difficult to be used for some applications, for example, generating pseudo-CT for PET-MR attenuation correction. This paper presents a deformation-invariant CycleGAN (DicycleGAN) method using deformable convolutional layers and new cycle-consistency losses. Its robustness dealing with data that suffer from domain-specific nonlinear deformations has been evaluated through comparison experiments performed on a multi-sequence brain MR dataset and a multi-modality abdominal dataset. Our method has displayed its ability to generate synthesized data that is aligned with the source while maintaining a proper quality of signal compared to CycleGAN-generated data. The proposed model also obtained comparable performance with CycleGAN when data from the source and target domains are alignable through simple affine transformations.
A two-stage 3D Unet framework for multi-class segmentation on full resolution image
Apr 12, 2018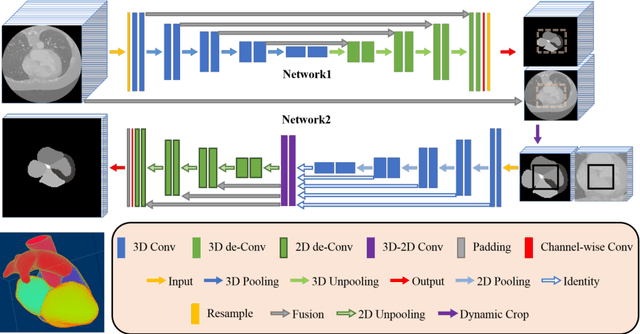
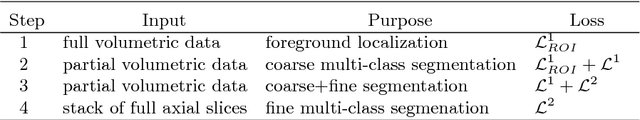

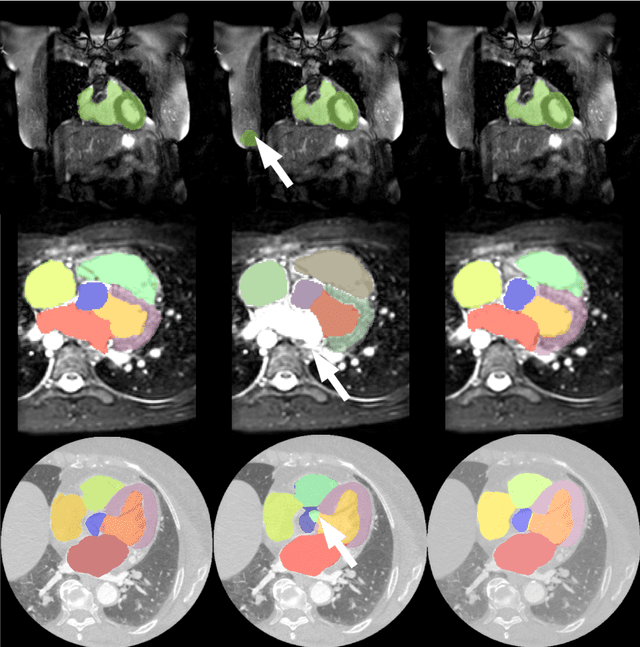
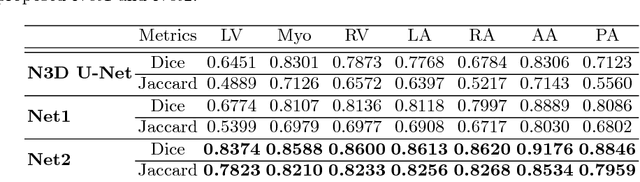
Abstract:Deep convolutional neural networks (CNNs) have been intensively used for multi-class segmentation of data from different modalities and achieved state-of-the-art performances. However, a common problem when dealing with large, high resolution 3D data is that the volumes input into the deep CNNs has to be either cropped or downsampled due to limited memory capacity of computing devices. These operations lead to loss of resolution and increment of class imbalance in the input data batches, which can downgrade the performances of segmentation algorithms. Inspired by the architecture of image super-resolution CNN (SRCNN) and self-normalization network (SNN), we developed a two-stage modified Unet framework that simultaneously learns to detect a ROI within the full volume and to classify voxels without losing the original resolution. Experiments on a variety of multi-modal volumes demonstrated that, when trained with a simply weighted dice coefficients and our customized learning procedure, this framework shows better segmentation performances than state-of-the-art Deep CNNs with advanced similarity metrics.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge