Rohan Dharmakumar
Active Sampling for MRI-based Sequential Decision Making
May 07, 2025



Abstract:Despite the superior diagnostic capability of Magnetic Resonance Imaging (MRI), its use as a Point-of-Care (PoC) device remains limited by high cost and complexity. To enable such a future by reducing the magnetic field strength, one key approach will be to improve sampling strategies. Previous work has shown that it is possible to make diagnostic decisions directly from k-space with fewer samples. Such work shows that single diagnostic decisions can be made, but if we aspire to see MRI as a true PoC, multiple and sequential decisions are necessary while minimizing the number of samples acquired. We present a novel multi-objective reinforcement learning framework enabling comprehensive, sequential, diagnostic evaluation from undersampled k-space data. Our approach during inference actively adapts to sequential decisions to optimally sample. To achieve this, we introduce a training methodology that identifies the samples that contribute the best to each diagnostic objective using a step-wise weighting reward function. We evaluate our approach in two sequential knee pathology assessment tasks: ACL sprain detection and cartilage thickness loss assessment. Our framework achieves diagnostic performance competitive with various policy-based benchmarks on disease detection, severity quantification, and overall sequential diagnosis, while substantially saving k-space samples. Our approach paves the way for the future of MRI as a comprehensive and affordable PoC device. Our code is publicly available at https://github.com/vios-s/MRI_Sequential_Active_Sampling
Improved Robustness for Deep Learning-based Segmentation of Multi-Center Myocardial Perfusion MRI Datasets Using Data Adaptive Uncertainty-guided Space-time Analysis
Aug 09, 2024



Abstract:Background. Fully automatic analysis of myocardial perfusion MRI datasets enables rapid and objective reporting of stress/rest studies in patients with suspected ischemic heart disease. Developing deep learning techniques that can analyze multi-center datasets despite limited training data and variations in software and hardware is an ongoing challenge. Methods. Datasets from 3 medical centers acquired at 3T (n = 150 subjects) were included: an internal dataset (inD; n = 95) and two external datasets (exDs; n = 55) used for evaluating the robustness of the trained deep neural network (DNN) models against differences in pulse sequence (exD-1) and scanner vendor (exD-2). A subset of inD (n = 85) was used for training/validation of a pool of DNNs for segmentation, all using the same spatiotemporal U-Net architecture and hyperparameters but with different parameter initializations. We employed a space-time sliding-patch analysis approach that automatically yields a pixel-wise "uncertainty map" as a byproduct of the segmentation process. In our approach, a given test case is segmented by all members of the DNN pool and the resulting uncertainty maps are leveraged to automatically select the "best" one among the pool of solutions. Results. The proposed DAUGS analysis approach performed similarly to the established approach on the internal dataset (p = n.s.) whereas it significantly outperformed on the external datasets (p < 0.005 for exD-1 and exD-2). Moreover, the number of image series with "failed" segmentation was significantly lower for the proposed vs. the established approach (4.3% vs. 17.1%, p < 0.0005). Conclusions. The proposed DAUGS analysis approach has the potential to improve the robustness of deep learning methods for segmentation of multi-center stress perfusion datasets with variations in the choice of pulse sequence, site location or scanner vendor.
The MRI Scanner as a Diagnostic: Image-less Active Sampling
Jun 24, 2024



Abstract:Despite the high diagnostic accuracy of Magnetic Resonance Imaging (MRI), using MRI as a Point-of-Care (POC) disease identification tool poses significant accessibility challenges due to the use of high magnetic field strength and lengthy acquisition times. We ask a simple question: Can we dynamically optimise acquired samples, at the patient level, according to an (automated) downstream decision task, while discounting image reconstruction? We propose an ML-based framework that learns an active sampling strategy, via reinforcement learning, at a patient-level to directly infer disease from undersampled k-space. We validate our approach by inferring Meniscus Tear in undersampled knee MRI data, where we achieve diagnostic performance comparable with ML-based diagnosis, using fully sampled k-space data. We analyse task-specific sampling policies, showcasing the adaptability of our active sampling approach. The introduced frugal sampling strategies have the potential to reduce high field strength requirements that in turn strengthen the viability of MRI-based POC disease identification and associated preliminary screening tools.
Unveiling Fairness Biases in Deep Learning-Based Brain MRI Reconstruction
Sep 25, 2023



Abstract:Deep learning (DL) reconstruction particularly of MRI has led to improvements in image fidelity and reduction of acquisition time. In neuroimaging, DL methods can reconstruct high-quality images from undersampled data. However, it is essential to consider fairness in DL algorithms, particularly in terms of demographic characteristics. This study presents the first fairness analysis in a DL-based brain MRI reconstruction model. The model utilises the U-Net architecture for image reconstruction and explores the presence and sources of unfairness by implementing baseline Empirical Risk Minimisation (ERM) and rebalancing strategies. Model performance is evaluated using image reconstruction metrics. Our findings reveal statistically significant performance biases between the gender and age subgroups. Surprisingly, data imbalance and training discrimination are not the main sources of bias. This analysis provides insights of fairness in DL-based image reconstruction and aims to improve equity in medical AI applications.
Temporal Uncertainty Localization to Enable Human-in-the-loop Analysis of Dynamic Contrast-enhanced Cardiac MRI Datasets
Aug 25, 2023Abstract:Dynamic contrast-enhanced (DCE) cardiac magnetic resonance imaging (CMRI) is a widely used modality for diagnosing myocardial blood flow (perfusion) abnormalities. During a typical free-breathing DCE-CMRI scan, close to 300 time-resolved images of myocardial perfusion are acquired at various contrast "wash in/out" phases. Manual segmentation of myocardial contours in each time-frame of a DCE image series can be tedious and time-consuming, particularly when non-rigid motion correction has failed or is unavailable. While deep neural networks (DNNs) have shown promise for analyzing DCE-CMRI datasets, a "dynamic quality control" (dQC) technique for reliably detecting failed segmentations is lacking. Here we propose a new space-time uncertainty metric as a dQC tool for DNN-based segmentation of free-breathing DCE-CMRI datasets by validating the proposed metric on an external dataset and establishing a human-in-the-loop framework to improve the segmentation results. In the proposed approach, we referred the top 10% most uncertain segmentations as detected by our dQC tool to the human expert for refinement. This approach resulted in a significant increase in the Dice score (p<0.001) and a notable decrease in the number of images with failed segmentation (16.2% to 11.3%) whereas the alternative approach of randomly selecting the same number of segmentations for human referral did not achieve any significant improvement. Our results suggest that the proposed dQC framework has the potential to accurately identify poor-quality segmentations and may enable efficient DNN-based analysis of DCE-CMRI in a human-in-the-loop pipeline for clinical interpretation and reporting of dynamic CMRI datasets.
Semi-supervised Pathology Segmentation with Disentangled Representations
Sep 05, 2020



Abstract:Automated pathology segmentation remains a valuable diagnostic tool in clinical practice. However, collecting training data is challenging. Semi-supervised approaches by combining labelled and unlabelled data can offer a solution to data scarcity. An approach to semi-supervised learning relies on reconstruction objectives (as self-supervision objectives) that learns in a joint fashion suitable representations for the task. Here, we propose Anatomy-Pathology Disentanglement Network (APD-Net), a pathology segmentation model that attempts to learn jointly for the first time: disentanglement of anatomy, modality, and pathology. The model is trained in a semi-supervised fashion with new reconstruction losses directly aiming to improve pathology segmentation with limited annotations. In addition, a joint optimization strategy is proposed to fully take advantage of the available annotations. We evaluate our methods with two private cardiac infarction segmentation datasets with LGE-MRI scans. APD-Net can perform pathology segmentation with few annotations, maintain performance with different amounts of supervision, and outperform related deep learning methods.
* 12 Pages, 4 figures
Disentangle, align and fuse for multimodal and zero-shot image segmentation
Nov 12, 2019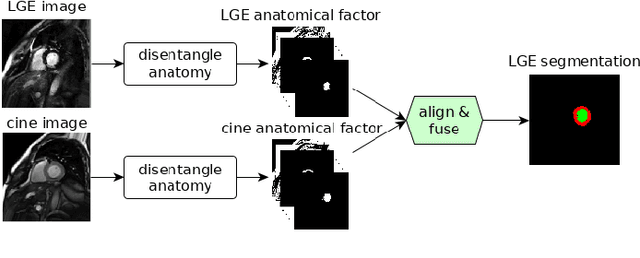
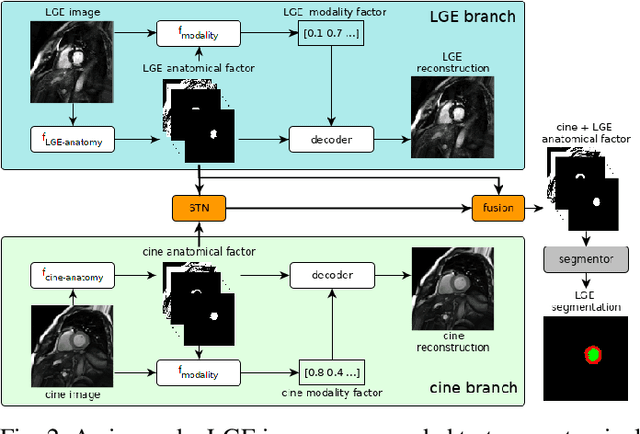
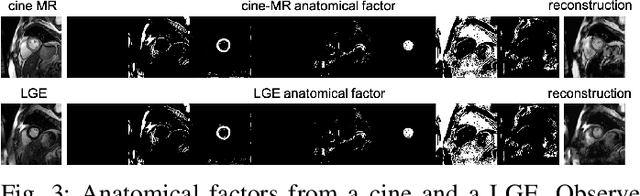

Abstract:Magnetic resonance (MR) protocols rely on several sequences to properly assess pathology and organ status. Yet, despite advances in image analysis we tend to treat each sequence, here termed modality, in isolation. Taking advantage of the information shared between modalities (largely an organ's anatomy) is beneficial for multi-modality multi-input processing and learning. However, we must overcome inherent anatomical misregistrations and disparities in signal intensity across the modalities to claim this benefit. We present a method that offers improved segmentation accuracy of the modality of interest (over a single input model), by learning to leverage information present in other modalities, enabling semi-supervised and zero shot learning. Core to our method is learning a disentangled decomposition into anatomical and imaging factors. Shared anatomical factors from the different inputs are jointly processed and fused to extract more accurate segmentation masks. Image misregistrations are corrected with a Spatial Transformer Network, that non-linearly aligns the anatomical factors. The imaging factor captures signal intensity characteristics across different modality data, and is used for image reconstruction, enabling semi-supervised learning. Temporal and slice pairing between inputs are learned dynamically. We demonstrate applications in Late Gadolinium Enhanced (LGE) and Blood Oxygenation Level Dependent (BOLD) cardiac segmentation, as well as in T2 abdominal segmentation.
Factorised Representation Learning in Cardiac Image Analysis
Mar 22, 2019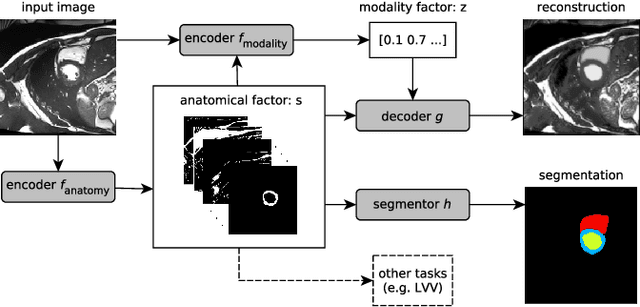
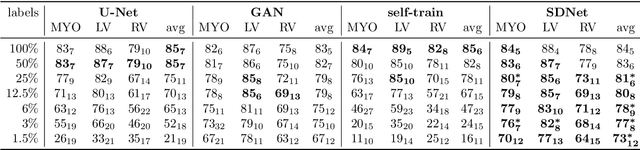
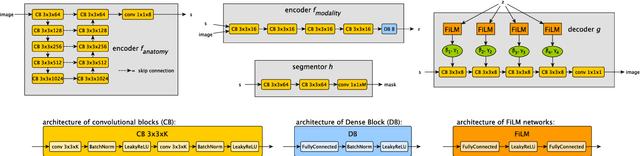
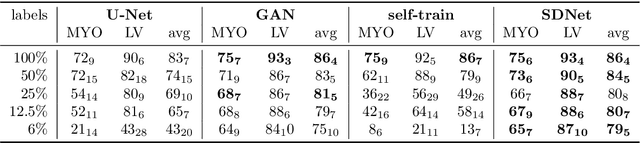
Abstract:Typically, a medical image offers spatial information on the anatomy (and pathology) modulated by imaging specific characteristics. Many imaging modalities including Magnetic Resonance Imaging (MRI) and Computed Tomography (CT) can be interpreted in this way. We can venture further and consider that a medical image naturally factors into some spatial factors depicting anatomy and factors that denote the imaging characteristics. Here, we explicitly learn this decomposed (factorised) representation of imaging data, focusing in particular on cardiac images. We propose Spatial Decomposition Network (SDNet), which factorises 2D medical images into spatial anatomical factors and non-spatial imaging factors. We demonstrate that this high-level representation is ideally suited for several medical image analysis tasks, such as semi-supervised segmentation, multi-task segmentation and regression, and image-to-image synthesis. Specifically, we show that our model can match the performance of fully supervised segmentation models, using only a fraction of the labelled images. Critically, we show that our factorised representation also benefits from supervision obtained either when we use auxiliary tasks to train the model in a multi-task setting (e.g. regressing to known cardiac indices), or when aggregating multimodal data from different sources (e.g. pooling together MRI and CT data). To explore the properties of the learned factorisation, we perform latent-space arithmetic and show that we can synthesise CT from MR and vice versa, by swapping the modality factors. We also demonstrate that the factor holding image specific information can be used to predict the input modality with high accuracy.
Factorised spatial representation learning: application in semi-supervised myocardial segmentation
Nov 02, 2018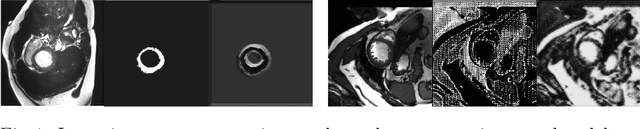
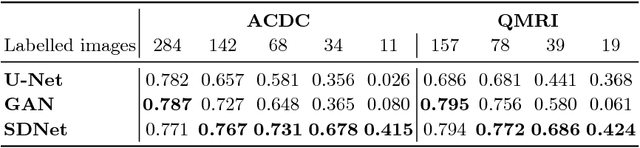
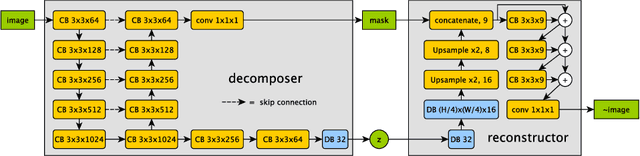
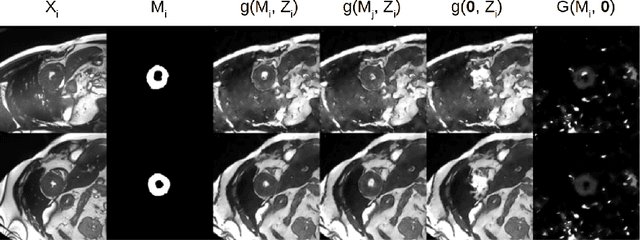
Abstract:The success and generalisation of deep learning algorithms heavily depend on learning good feature representations. In medical imaging this entails representing anatomical information, as well as properties related to the specific imaging setting. Anatomical information is required to perform further analysis, whereas imaging information is key to disentangle scanner variability and potential artefacts. The ability to factorise these would allow for training algorithms only on the relevant information according to the task. To date, such factorisation has not been attempted. In this paper, we propose a methodology of latent space factorisation relying on the cycle-consistency principle. As an example application, we consider cardiac MR segmentation, where we separate information related to the myocardium from other features related to imaging and surrounding substructures. We demonstrate the proposed method's utility in a semi-supervised setting: we use very few labelled images together with many unlabelled images to train a myocardium segmentation neural network. Specifically, we achieve comparable performance to fully supervised networks using a fraction of labelled images in experiments on ACDC and a dataset from Edinburgh Imaging Facility QMRI. Code will be made available at https://github.com/agis85/spatial_factorisation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge