Cheng Zhuo
Expressivity Enhancement with Efficient Quadratic Neurons for Convolutional Neural Networks
Jun 10, 2023Abstract:Convolutional neural networks (CNNs) have been successfully applied in a range of fields such as image classification and object segmentation. To improve their expressivity, various techniques, such as novel CNN architectures, have been explored. However, the performance gain from such techniques tends to diminish. To address this challenge, many researchers have shifted their focus to increasing the non-linearity of neurons, the fundamental building blocks of neural networks, to enhance the network expressivity. Nevertheless, most of these approaches incur a large number of parameters and thus formidable computation cost inevitably, impairing their efficiency to be deployed in practice. In this work, an efficient quadratic neuron structure is proposed to preserve the non-linearity with only negligible parameter and computation cost overhead. The proposed quadratic neuron can maximize the utilization of second-order computation information to improve the network performance. The experimental results have demonstrated that the proposed quadratic neuron can achieve a higher accuracy and a better computation efficiency in classification tasks compared with both linear neurons and non-linear neurons from previous works.
SFCNeXt: a simple fully convolutional network for effective brain age estimation with small sample size
May 30, 2023Abstract:Deep neural networks (DNN) have been designed to predict the chronological age of a healthy brain from T1-weighted magnetic resonance images (T1 MRIs), and the predicted brain age could serve as a valuable biomarker for the early detection of development-related or aging-related disorders. Recent DNN models for brain age estimations usually rely too much on large sample sizes and complex network structures for multi-stage feature refinement. However, in clinical application scenarios, researchers usually cannot obtain thousands or tens of thousands of MRIs in each data center for thorough training of these complex models. This paper proposes a simple fully convolutional network (SFCNeXt) for brain age estimation in small-sized cohorts with biased age distributions. The SFCNeXt consists of Single Pathway Encoded ConvNeXt (SPEC) and Hybrid Ranking Loss (HRL), aiming to estimate brain ages in a lightweight way with a sufficient exploration of MRI, age, and ranking features of each batch of subjects. Experimental results demonstrate the superiority and efficiency of our approach.
SteppingNet: A Stepping Neural Network with Incremental Accuracy Enhancement
Nov 27, 2022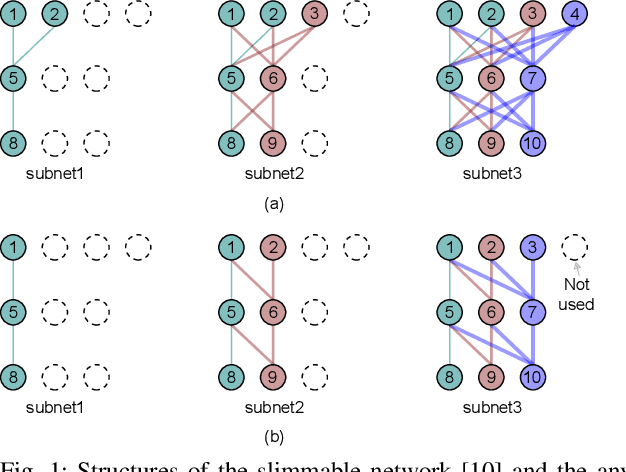

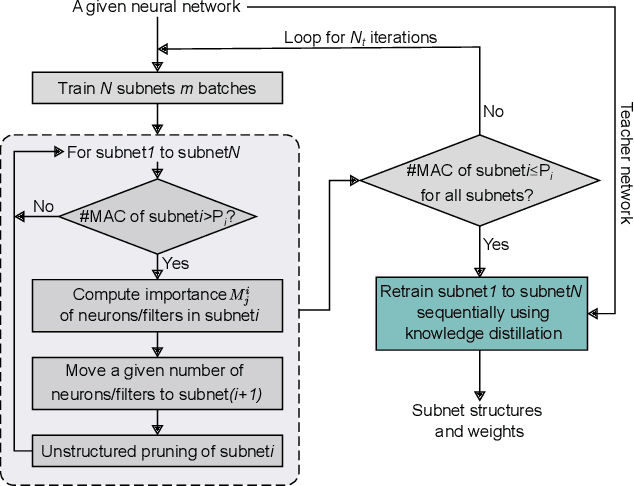
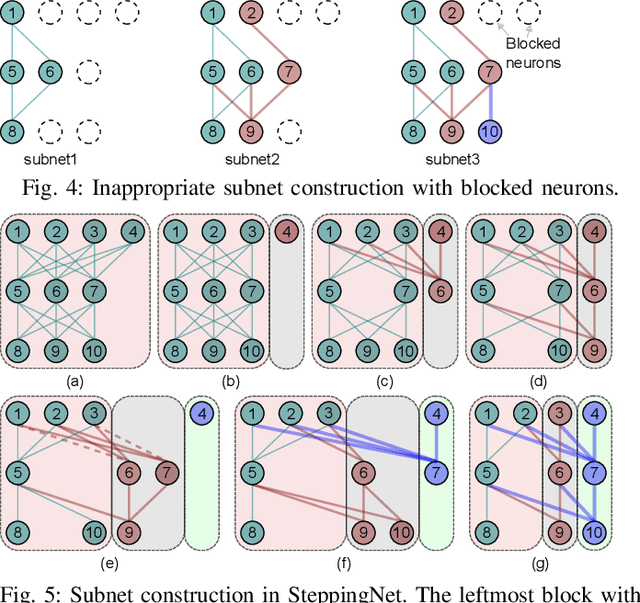
Abstract:Deep neural networks (DNNs) have successfully been applied in many fields in the past decades. However, the increasing number of multiply-and-accumulate (MAC) operations in DNNs prevents their application in resource-constrained and resource-varying platforms, e.g., mobile phones and autonomous vehicles. In such platforms, neural networks need to provide acceptable results quickly and the accuracy of the results should be able to be enhanced dynamically according to the computational resources available in the computing system. To address these challenges, we propose a design framework called SteppingNet. SteppingNet constructs a series of subnets whose accuracy is incrementally enhanced as more MAC operations become available. Therefore, this design allows a trade-off between accuracy and latency. In addition, the larger subnets in SteppingNet are built upon smaller subnets, so that the results of the latter can directly be reused in the former without recomputation. This property allows SteppingNet to decide on-the-fly whether to enhance the inference accuracy by executing further MAC operations. Experimental results demonstrate that SteppingNet provides an effective incremental accuracy improvement and its inference accuracy consistently outperforms the state-of-the-art work under the same limit of computational resources.
HDNet: Hierarchical Dynamic Network for Gait Recognition using Millimeter-Wave Radar
Nov 01, 2022



Abstract:Gait recognition is widely used in diversified practical applications. Currently, the most prevalent approach is to recognize human gait from RGB images, owing to the progress of computer vision technologies. Nevertheless, the perception capability of RGB cameras deteriorates in rough circumstances, and visual surveillance may cause privacy invasion. Due to the robustness and non-invasive feature of millimeter wave (mmWave) radar, radar-based gait recognition has attracted increasing attention in recent years. In this research, we propose a Hierarchical Dynamic Network (HDNet) for gait recognition using mmWave radar. In order to explore more dynamic information, we propose point flow as a novel point clouds descriptor. We also devise a dynamic frame sampling module to promote the efficiency of computation without deteriorating performance noticeably. To prove the superiority of our methods, we perform extensive experiments on two public mmWave radar-based gait recognition datasets, and the results demonstrate that our model is superior to existing state-of-the-art methods.
A Homogeneous Processing Fabric for Matrix-Vector Multiplication and Associative Search Using Ferroelectric Time-Domain Compute-in-Memory
Sep 24, 2022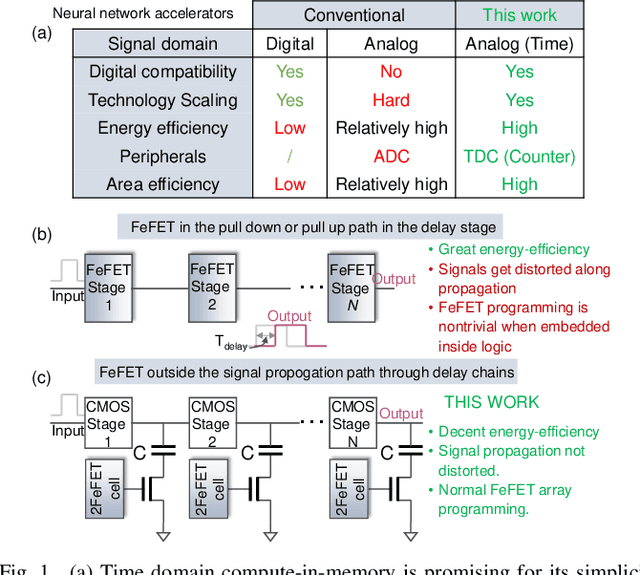
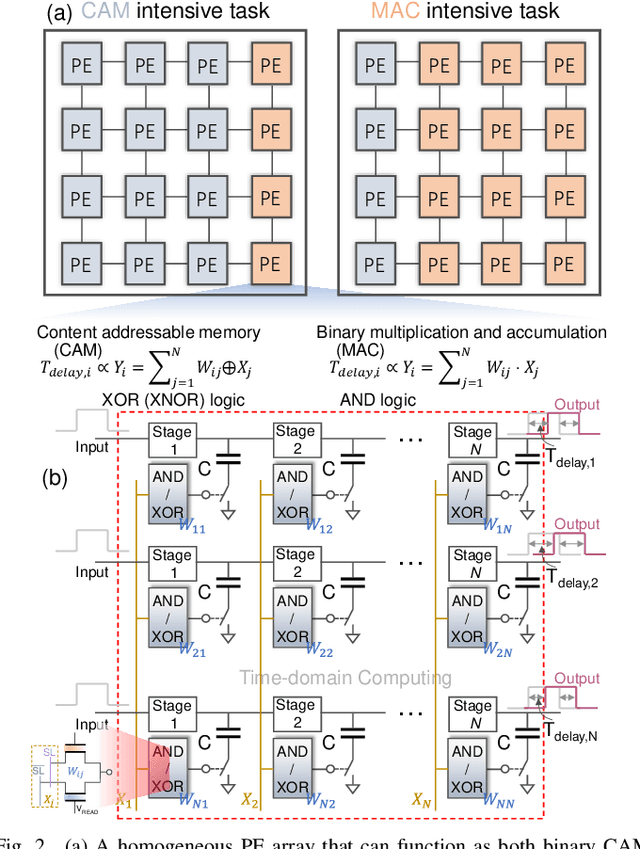
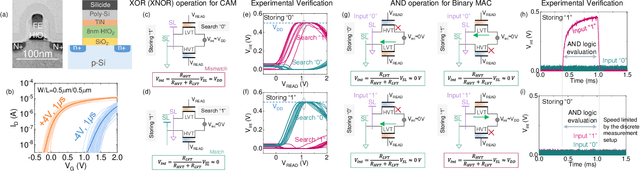
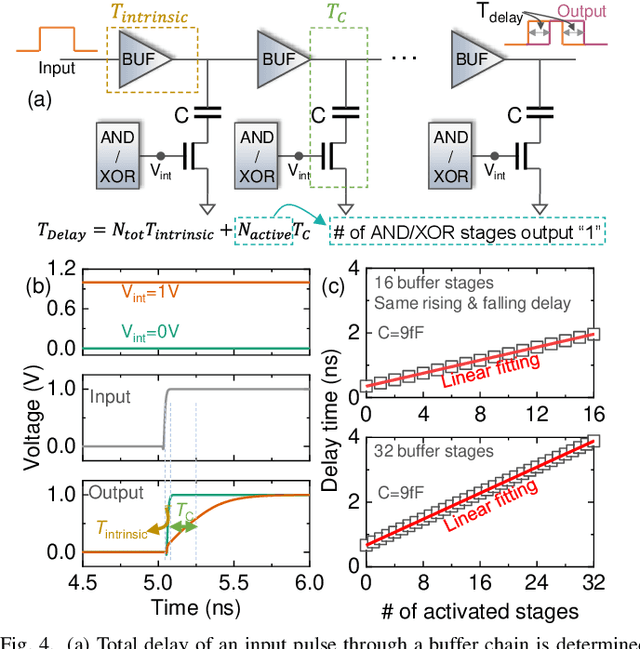
Abstract:In this work, we propose a ferroelectric FET(FeFET) time-domain compute-in-memory (TD-CiM) array as a homogeneous processing fabric for binary multiplication-accumulation (MAC) and content addressable memory (CAM). We demonstrate that: i) the XOR(XNOR)/AND logic function can be realized using a single cell composed of 2FeFETs connected in series; ii) a two-phase computation in an inverter chain with each stage featuring the XOR/AND cell to control the associated capacitor loading and the computation results of binary MAC and CAM are reflected in the chain output signal delay, illustrating full digital compatibility; iii) comprehensive theoretical and experimental validation of the proposed 2FeFET cell and inverter delay chains and their robustness against FeFET variation; iv) the homogeneous processing fabric is applied in hyperdimensional computing to show dynamic and fine-grain resource allocation to accommodate different tasks requiring varying demands over the binary MAC and CAM resources.
GANDSE: Generative Adversarial Network based Design Space Exploration for Neural Network Accelerator Design
Aug 01, 2022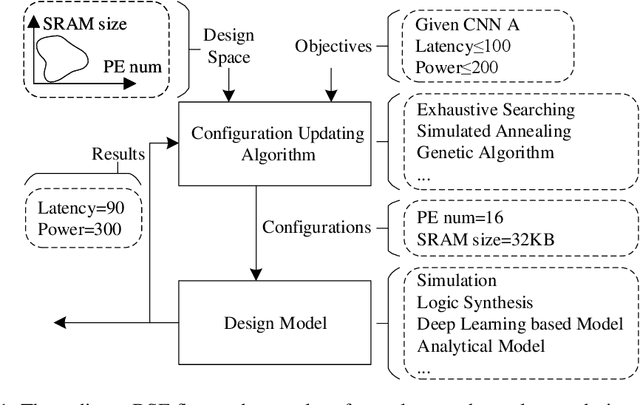
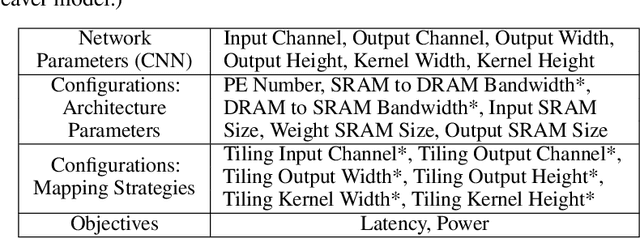

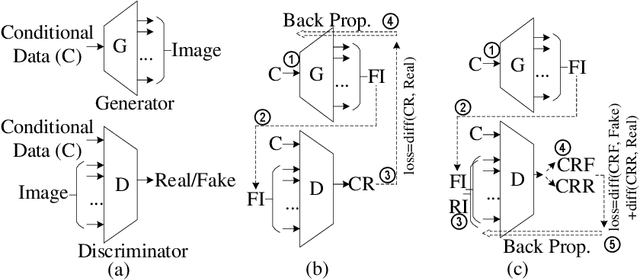
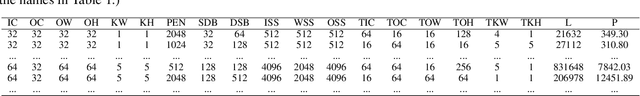
Abstract:With the popularity of deep learning, the hardware implementation platform of deep learning has received increasing interest. Unlike the general purpose devices, e.g., CPU, or GPU, where the deep learning algorithms are executed at the software level, neural network hardware accelerators directly execute the algorithms to achieve higher both energy efficiency and performance improvements. However, as the deep learning algorithms evolve frequently, the engineering effort and cost of designing the hardware accelerators are greatly increased. To improve the design quality while saving the cost, design automation for neural network accelerators was proposed, where design space exploration algorithms are used to automatically search the optimized accelerator design within a design space. Nevertheless, the increasing complexity of the neural network accelerators brings the increasing dimensions to the design space. As a result, the previous design space exploration algorithms are no longer effective enough to find an optimized design. In this work, we propose a neural network accelerator design automation framework named GANDSE, where we rethink the problem of design space exploration, and propose a novel approach based on the generative adversarial network (GAN) to support an optimized exploration for high dimension large design space. The experiments show that GANDSE is able to find the more optimized designs in negligible time compared with approaches including multilayer perceptron and deep reinforcement learning.
RT-DNAS: Real-time Constrained Differentiable Neural Architecture Search for 3D Cardiac Cine MRI Segmentation
Jun 13, 2022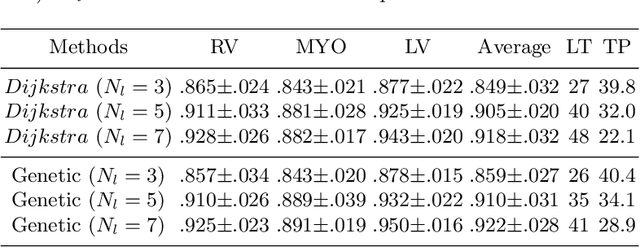

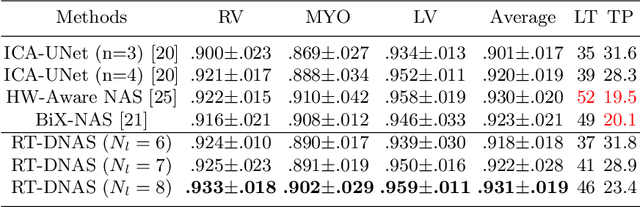
Abstract:Accurately segmenting temporal frames of cine magnetic resonance imaging (MRI) is a crucial step in various real-time MRI guided cardiac interventions. To achieve fast and accurate visual assistance, there are strict requirements on the maximum latency and minimum throughput of the segmentation framework. State-of-the-art neural networks on this task are mostly hand-crafted to satisfy these constraints while achieving high accuracy. On the other hand, while existing literature have demonstrated the power of neural architecture search (NAS) in automatically identifying the best neural architectures for various medical applications, they are mostly guided by accuracy, sometimes with computation complexity, and the importance of real-time constraints are overlooked. A major challenge is that such constraints are non-differentiable and are thus not compatible with the widely used differentiable NAS frameworks. In this paper, we present a strategy that directly handles real-time constraints in a differentiable NAS framework named RT-DNAS. Experiments on extended 2017 MICCAI ACDC dataset show that compared with state-of-the-art manually and automatically designed architectures, RT-DNAS is able to identify ones with better accuracy while satisfying the real-time constraints.
OTFPF: Optimal Transport-Based Feature Pyramid Fusion Network for Brain Age Estimation with 3D Overlapped ConvNeXt
May 11, 2022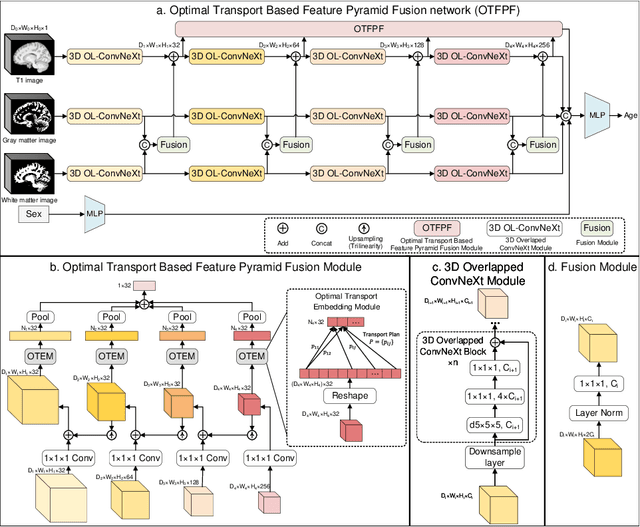
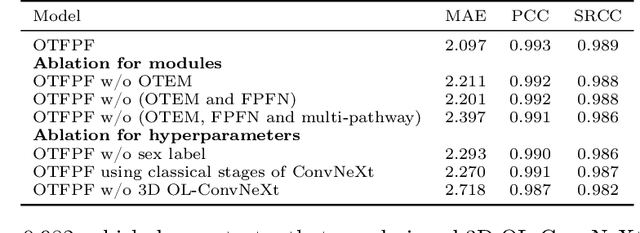
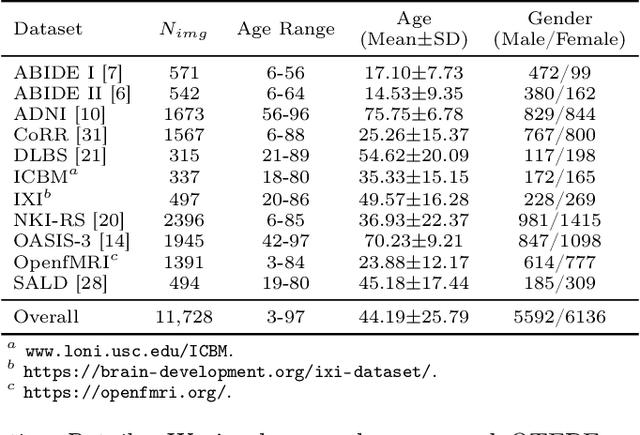
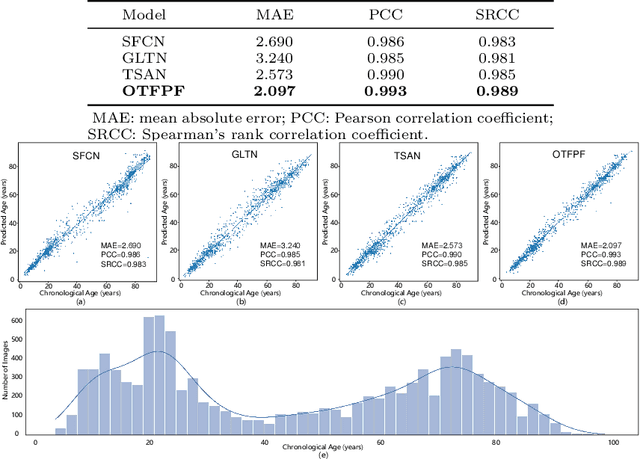

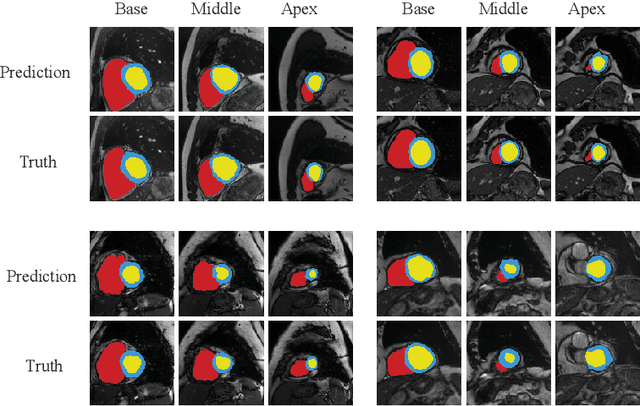
Abstract:Chronological age of healthy brain is able to be predicted using deep neural networks from T1-weighted magnetic resonance images (T1 MRIs), and the predicted brain age could serve as an effective biomarker for detecting aging-related diseases or disorders. In this paper, we propose an end-to-end neural network architecture, referred to as optimal transport based feature pyramid fusion (OTFPF) network, for the brain age estimation with T1 MRIs. The OTFPF consists of three types of modules: Optimal Transport based Feature Pyramid Fusion (OTFPF) module, 3D overlapped ConvNeXt (3D OL-ConvNeXt) module and fusion module. These modules strengthen the OTFPF network's understanding of each brain's semi-multimodal and multi-level feature pyramid information, and significantly improve its estimation performances. Comparing with recent state-of-the-art models, the proposed OTFPF converges faster and performs better. The experiments with 11,728 MRIs aged 3-97 years show that OTFPF network could provide accurate brain age estimation, yielding mean absolute error (MAE) of 2.097, Pearson's correlation coefficient (PCC) of 0.993 and Spearman's rank correlation coefficient (SRCC) of 0.989, between the estimated and chronological ages. Widespread quantitative experiments and ablation experiments demonstrate the superiority and rationality of OTFPF network. The codes and implement details will be released on GitHub: https://github.com/ZJU-Brain/OTFPF after final decision.
Worst-Case Dynamic Power Distribution Network Noise Prediction Using Convolutional Neural Network
Apr 27, 2022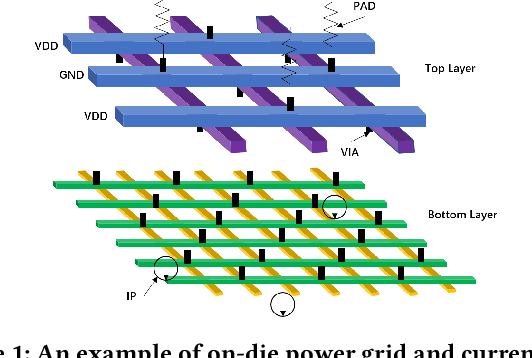
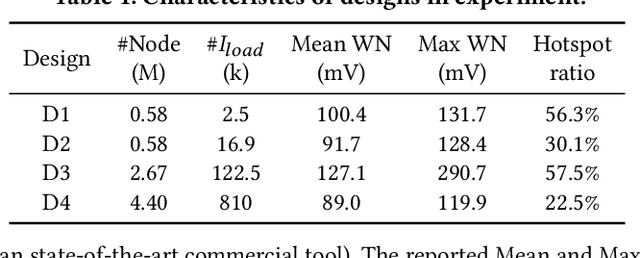
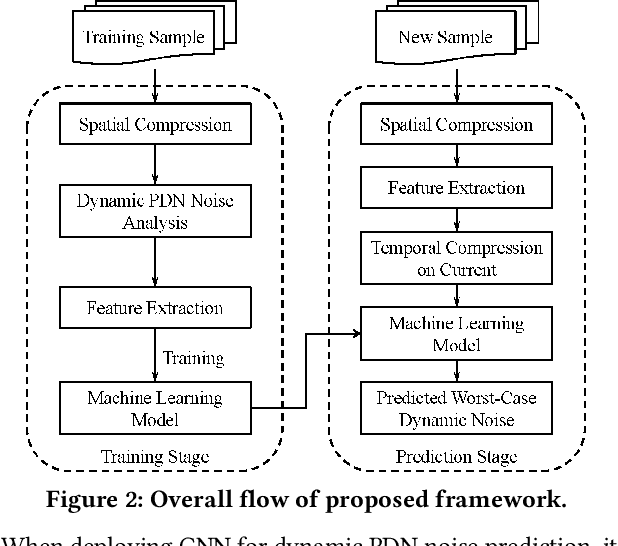
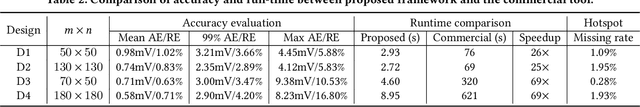
Abstract:Worst-case dynamic PDN noise analysis is an essential step in PDN sign-off to ensure the performance and reliability of chips. However, with the growing PDN size and increasing scenarios to be validated, it becomes very time- and resource-consuming to conduct full-stack PDN simulation to check the worst-case noise for different test vectors. Recently, various works have proposed machine learning based methods for supply noise prediction, many of which still suffer from large training overhead, inefficiency, or non-scalability. Thus, this paper proposed an efficient and scalable framework for the worst-case dynamic PDN noise prediction. The framework first reduces the spatial and temporal redundancy in the PDN and input current vector, and then employs efficient feature extraction as well as a novel convolutional neural network architecture to predict the worst-case dynamic PDN noise. Experimental results show that the proposed framework consistently outperforms the commercial tool and the state-of-the-art machine learning method with only 0.63-1.02% mean relative error and 25-69$\times$ speedup.
An Ultra-Compact Single FeFET Binary and Multi-Bit Associative Search Engine
Mar 15, 2022Abstract:Content addressable memory (CAM) is widely used in associative search tasks for its highly parallel pattern matching capability. To accommodate the increasingly complex and data-intensive pattern matching tasks, it is critical to keep improving the CAM density to enhance the performance and area efficiency. In this work, we demonstrate: i) a novel ultra-compact 1FeFET CAM design that enables parallel associative search and in-memory hamming distance calculation; ii) a multi-bit CAM for exact search using the same CAM cell; iii) compact device designs that integrate the series resistor current limiter into the intrinsic FeFET structure to turn the 1FeFET1R into an effective 1FeFET cell; iv) a successful 2-step search operation and a sufficient sensing margin of the proposed binary and multi-bit 1FeFET1R CAM array with sizes of practical interests in both experiments and simulations, given the existing unoptimized FeFET device variation; v) 89.9x speedup and 66.5x energy efficiency improvement over the state-of-the art alignment tools on GPU in accelerating genome pattern matching applications through the hyperdimensional computing paradigm.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge