"Information": models, code, and papers
Black-box Coreset Variational Inference
Nov 04, 2022
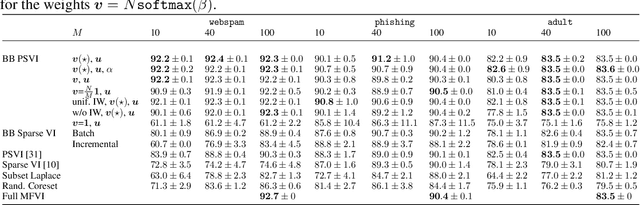
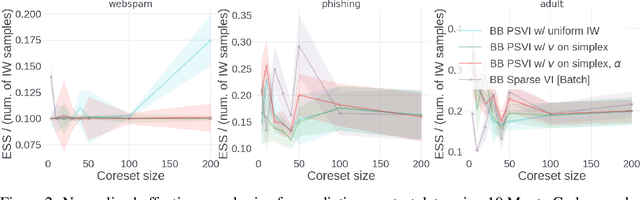
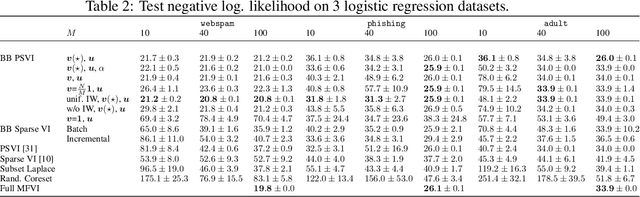
Recent advances in coreset methods have shown that a selection of representative datapoints can replace massive volumes of data for Bayesian inference, preserving the relevant statistical information and significantly accelerating subsequent downstream tasks. Existing variational coreset constructions rely on either selecting subsets of the observed datapoints, or jointly performing approximate inference and optimizing pseudodata in the observed space akin to inducing points methods in Gaussian Processes. So far, both approaches are limited by complexities in evaluating their objectives for general purpose models, and require generating samples from a typically intractable posterior over the coreset throughout inference and testing. In this work, we present a black-box variational inference framework for coresets that overcomes these constraints and enables principled application of variational coresets to intractable models, such as Bayesian neural networks. We apply our techniques to supervised learning problems, and compare them with existing approaches in the literature for data summarization and inference.
Behavior Score-Embedded Brain Encoder Network for Improved Classification of Alzheimer Disease Using Resting State fMRI
Nov 04, 2022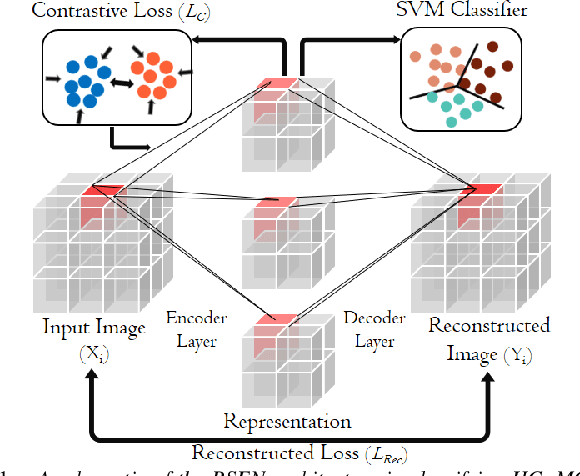
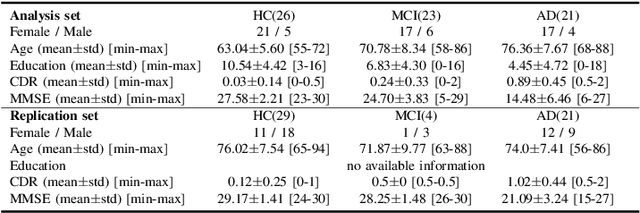
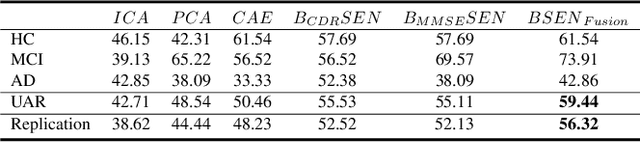
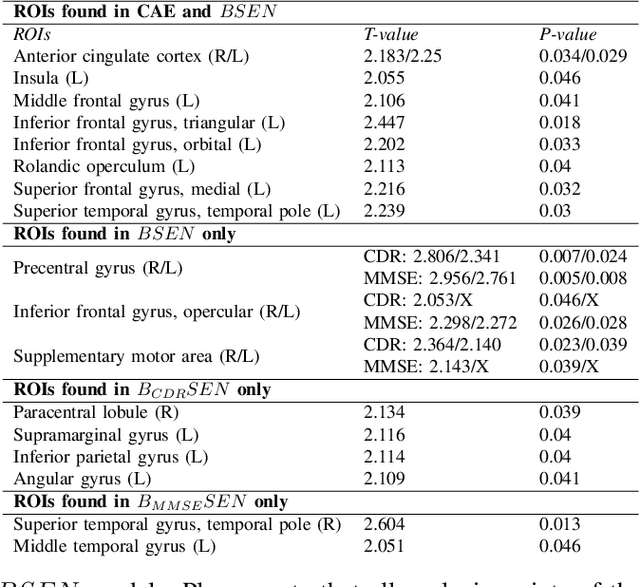
The ability to accurately detect onset of dementia is important in the treatment of the disease. Clinically, the diagnosis of Alzheimer Disease (AD) and Mild Cognitive Impairment (MCI) patients are based on an integrated assessment of psychological tests and brain imaging such as positron emission tomography (PET) and anatomical magnetic resonance imaging (MRI). In this work using two different datasets, we propose a behavior score-embedded encoder network (BSEN) that integrates regularly adminstrated psychological tests information into the encoding procedure of representing subject's restingstate fMRI data for automatic classification tasks. BSEN is based on a 3D convolutional autoencoder structure with contrastive loss jointly optimized using behavior scores from MiniMental State Examination (MMSE) and Clinical Dementia Rating (CDR). Our proposed classification framework of using BSEN achieved an overall recognition accuracy of 59.44% (3-class classification: AD, MCI and Healthy Control), and we further extracted the most discriminative regions between healthy control (HC) and AD patients.
Generation of Anonymous Chest Radiographs Using Latent Diffusion Models for Training Thoracic Abnormality Classification Systems
Nov 04, 2022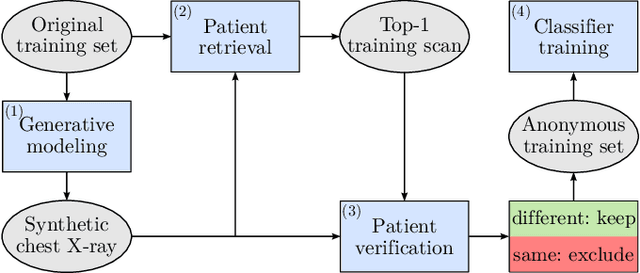
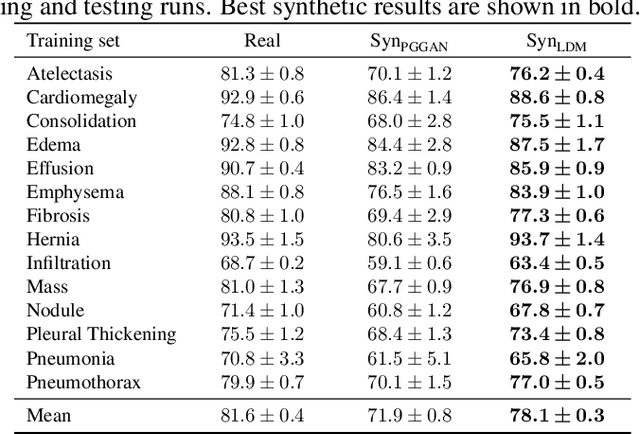
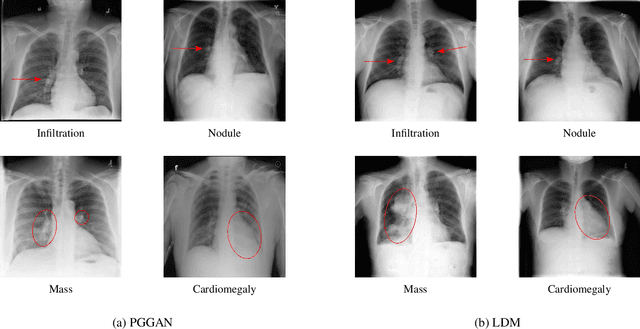
The availability of large-scale chest X-ray datasets is a requirement for developing well-performing deep learning-based algorithms in thoracic abnormality detection and classification. However, biometric identifiers in chest radiographs hinder the public sharing of such data for research purposes due to the risk of patient re-identification. To counteract this issue, synthetic data generation offers a solution for anonymizing medical images. This work employs a latent diffusion model to synthesize an anonymous chest X-ray dataset of high-quality class-conditional images. We propose a privacy-enhancing sampling strategy to ensure the non-transference of biometric information during the image generation process. The quality of the generated images and the feasibility of serving as exclusive training data are evaluated on a thoracic abnormality classification task. Compared to a real classifier, we achieve competitive results with a performance gap of only 3.5% in the area under the receiver operating characteristic curve.
Reducing Two-Way Ranging Variance by Signal-Timing Optimization
Nov 01, 2022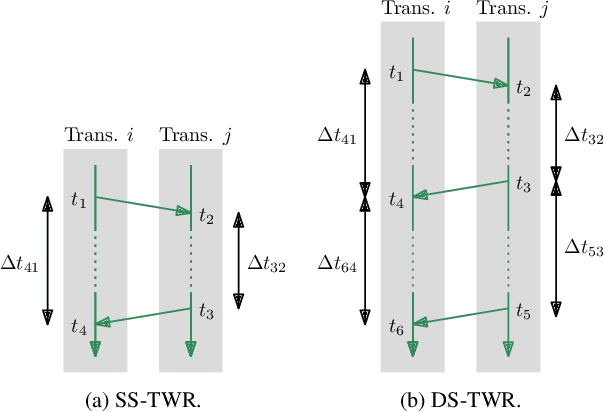
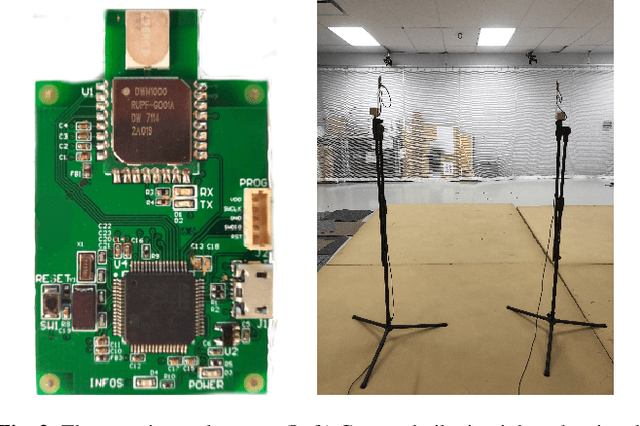
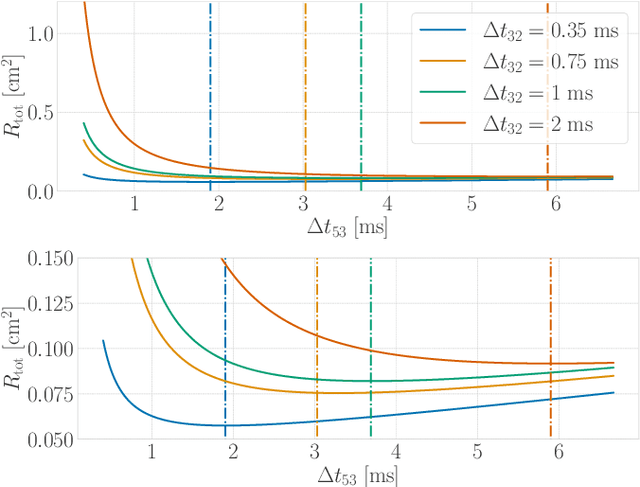

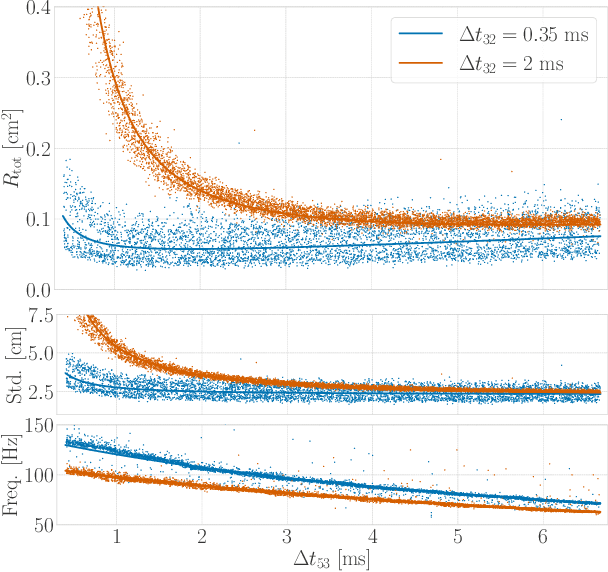
Time-of-flight-based range measurements among transceivers with different clocks requires ranging protocols that accommodate for the varying rates of the clocks. Double-sided two-way ranging (DS-TWR) has recently been widely adopted as a standard protocol due to its accuracy; however, the precision of DS-TWR has not been clearly addressed. In this paper, an analytical model of the variance of DS-TWR is derived as a function of the user-programmed response delays. Consequently, this allows formulating an optimization problem over the response delays in order to maximize the information gained from range measurements by addressing the effect of varying the response delays on the precision and frequency of the measurements. The derived analytical variance model and proposed optimization formulation are validated experimentally with 2 ranging UWB transceivers, where 29 million range measurements are collected.
Expressive-VC: Highly Expressive Voice Conversion with Attention Fusion of Bottleneck and Perturbation Features
Nov 09, 2022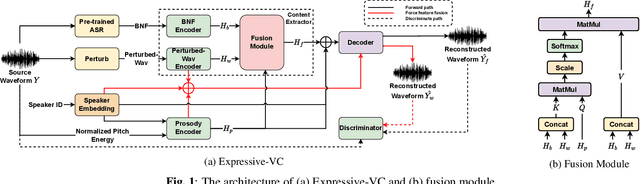
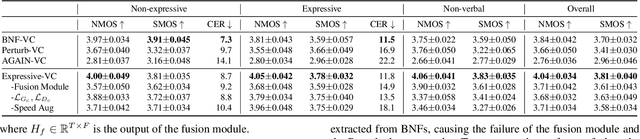
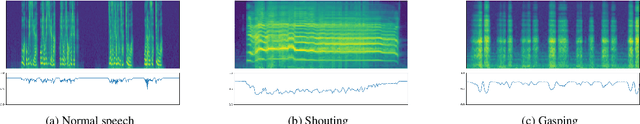

Voice conversion for highly expressive speech is challenging. Current approaches struggle with the balancing between speaker similarity, intelligibility and expressiveness. To address this problem, we propose Expressive-VC, a novel end-to-end voice conversion framework that leverages advantages from both neural bottleneck feature (BNF) approach and information perturbation approach. Specifically, we use a BNF encoder and a Perturbed-Wav encoder to form a content extractor to learn linguistic and para-linguistic features respectively, where BNFs come from a robust pre-trained ASR model and the perturbed wave becomes speaker-irrelevant after signal perturbation. We further fuse the linguistic and para-linguistic features through an attention mechanism, where speaker-dependent prosody features are adopted as the attention query, which result from a prosody encoder with target speaker embedding and normalized pitch and energy of source speech as input. Finally the decoder consumes the integrated features and the speaker-dependent prosody feature to generate the converted speech. Experiments demonstrate that Expressive-VC is superior to several state-of-the-art systems, achieving both high expressiveness captured from the source speech and high speaker similarity with the target speaker; meanwhile intelligibility is well maintained.
A Note on Task-Aware Loss via Reweighing Prediction Loss by Decision-Regret
Nov 09, 2022
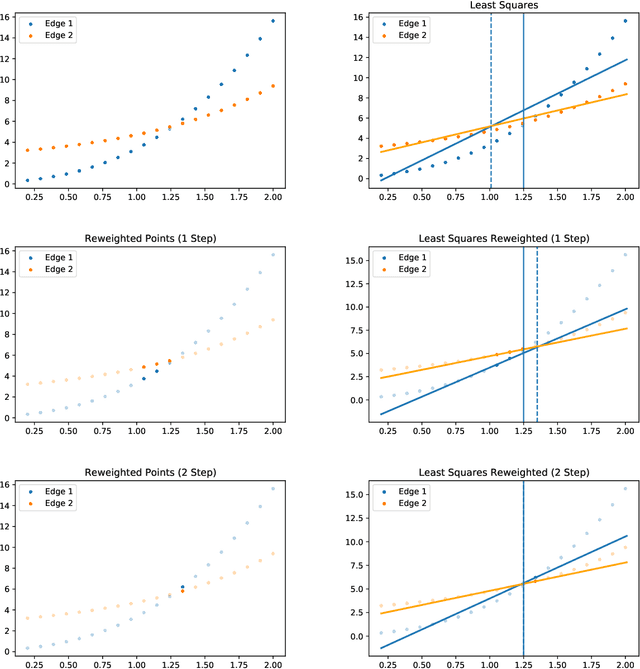
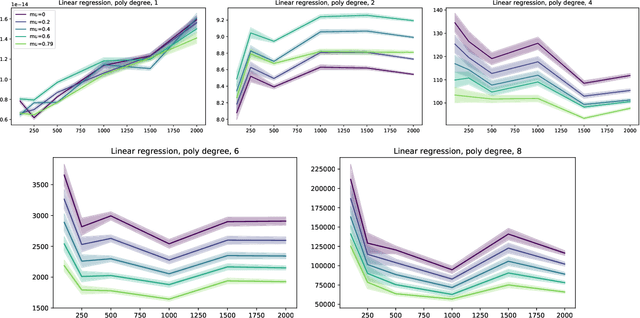
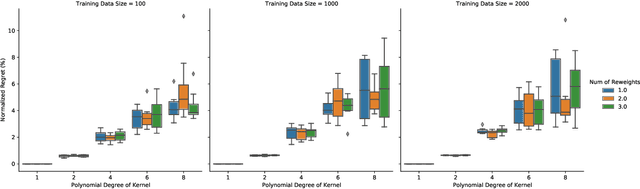
In this short technical note we propose a baseline for decision-aware learning for contextual linear optimization, which solves stochastic linear optimization when cost coefficients can be predicted based on context information. We propose a decision-aware version of predict-then-optimize. We reweigh the prediction error by the decision regret incurred by an (unweighted) pilot estimator of costs to obtain a decision-aware predictor, then optimize with cost predictions from the decision-aware predictor. This method can be motivated as a finite-difference, iterate-independent approximation of the gradients of previously proposed end-to-end learning algorithms; it is also consistent with previously suggested intuition for end-to-end learning. This baseline is computationally easy to implement with readily available reweighted prediction oracles and linear optimization, and can be implemented with convex optimization so long as the prediction error minimization is convex. Empirically, we demonstrate that this approach can lead to improvements over a "predict-then-optimize" framework for settings with misspecified models, and is competitive with other end-to-end approaches. Therefore, due to its simplicity and ease of use, we suggest it as a simple baseline for end-to-end and decision-aware learning.
Nested Named Entity Recognition from Medical Texts: An Adaptive Shared Network Architecture with Attentive CRF
Nov 09, 2022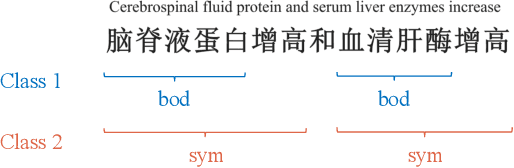

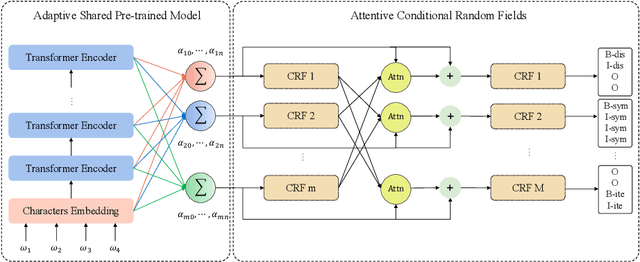
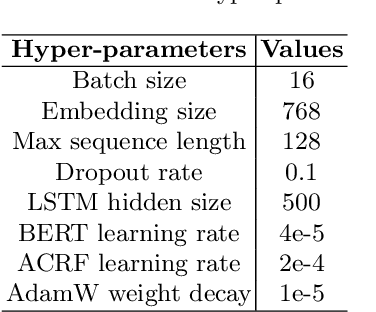
Recognizing useful named entities plays a vital role in medical information processing, which helps drive the development of medical area research. Deep learning methods have achieved good results in medical named entity recognition (NER). However, we find that existing methods face great challenges when dealing with the nested named entities. In this work, we propose a novel method, referred to as ASAC, to solve the dilemma caused by the nested phenomenon, in which the core idea is to model the dependency between different categories of entity recognition. The proposed method contains two key modules: the adaptive shared (AS) part and the attentive conditional random field (ACRF) module. The former part automatically assigns adaptive weights across each task to achieve optimal recognition accuracy in the multi-layer network. The latter module employs the attention operation to model the dependency between different entities. In this way, our model could learn better entity representations by capturing the implicit distinctions and relationships between different categories of entities. Extensive experiments on public datasets verify the effectiveness of our method. Besides, we also perform ablation analyses to deeply understand our methods.
Composite Fixed-Length Ordered Features for Palmprint Template Protection with Diminished Performance Loss
Nov 09, 2022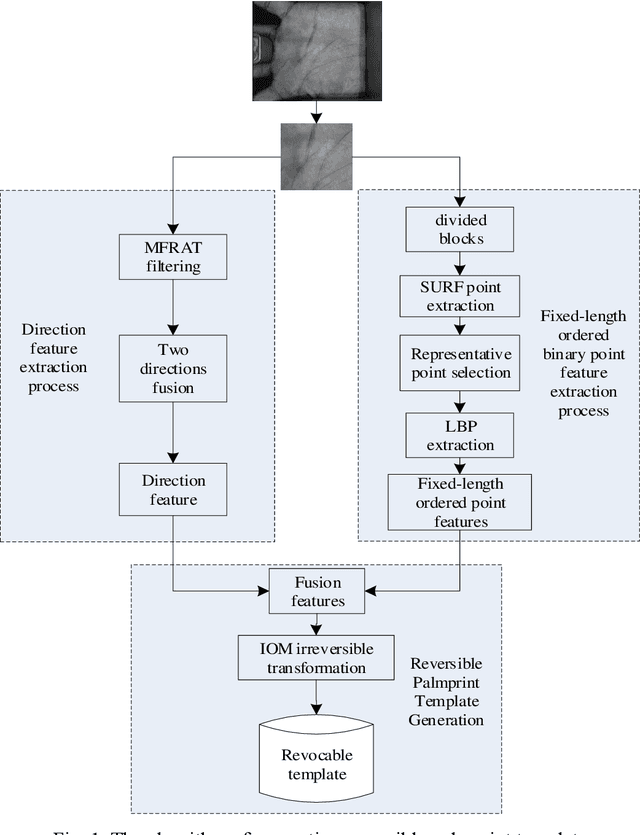
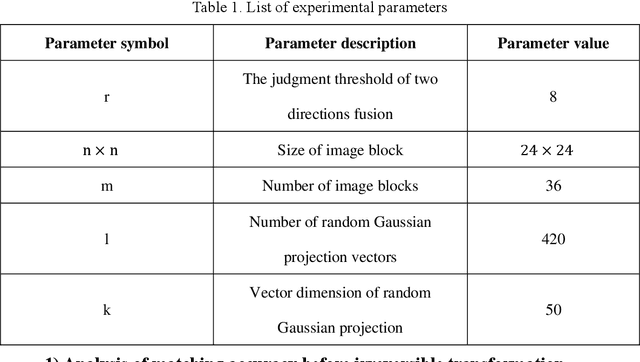
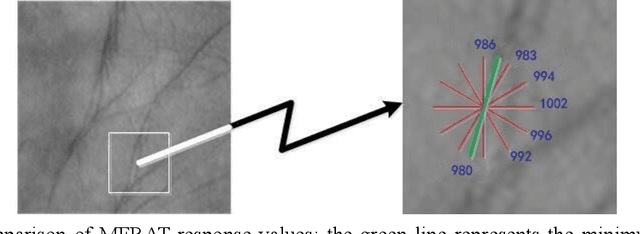
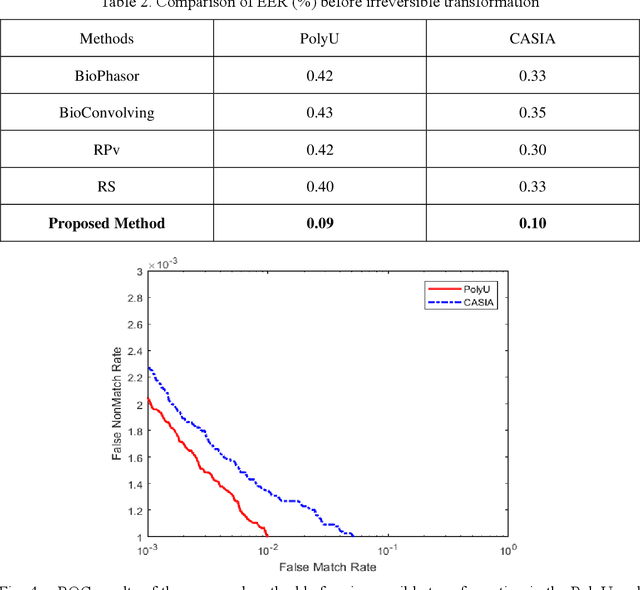
Palmprint recognition has become more and more popular due to its advantages over other biometric modalities such as fingerprint, in that it is larger in area, richer in information and able to work at a distance. However, the issue of palmprint privacy and security (especially palmprint template protection) remains under-studied. Among the very few research works, most of them only use the directional and orientation features of the palmprint with transformation processing, yielding unsatisfactory protection and identification performance. Thus, this paper proposes a palmprint template protection-oriented operator that has a fixed length and is ordered in nature, by fusing point features and orientation features. Firstly, double orientations are extracted with more accuracy based on MFRAT. Then key points of SURF are extracted and converted to be fixed-length and ordered features. Finally, composite features that fuse up the double orientations and SURF points are transformed using the irreversible transformation of IOM to generate the revocable palmprint template. Experiments show that the EER after irreversible transformation on the PolyU and CASIA databases are 0.17% and 0.19% respectively, and the absolute precision loss is 0.08% and 0.07%, respectively, which proves the advantage of our method.
Training self-supervised peptide sequence models on artificially chopped proteins
Nov 09, 2022
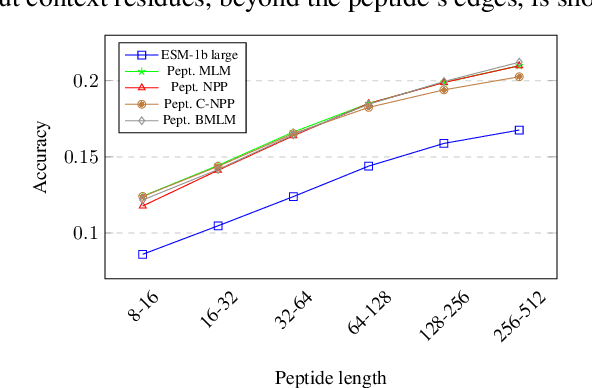
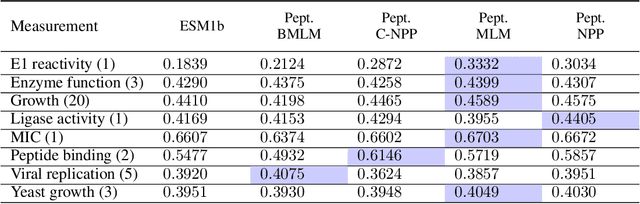
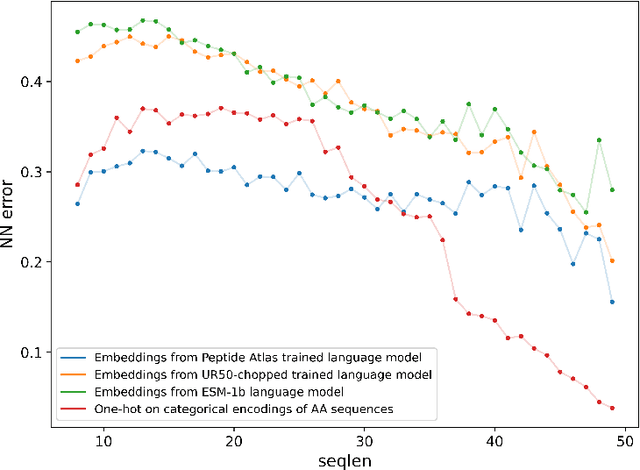
Representation learning for proteins has primarily focused on the global understanding of protein sequences regardless of their length. However, shorter proteins (known as peptides) take on distinct structures and functions compared to their longer counterparts. Unfortunately, there are not as many naturally occurring peptides available to be sequenced and therefore less peptide-specific data to train with. In this paper, we propose a new peptide data augmentation scheme, where we train peptide language models on artificially constructed peptides that are small contiguous subsets of longer, wild-type proteins; we refer to the training peptides as "chopped proteins". We evaluate the representation potential of models trained with chopped proteins versus natural peptides and find that training language models with chopped proteins results in more generalized embeddings for short protein sequences. These peptide-specific models also retain information about the original protein they were derived from better than language models trained on full-length proteins. We compare masked language model training objectives to three novel peptide-specific training objectives: next-peptide prediction, contrastive peptide selection and evolution-weighted MLM. We demonstrate improved zero-shot learning performance for a deep mutational scan peptides benchmark.
GIU-GANs: Global Information Utilization for Generative Adversarial Networks
Jan 25, 2022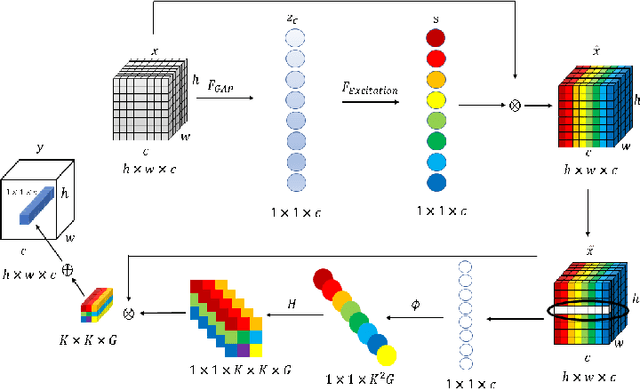
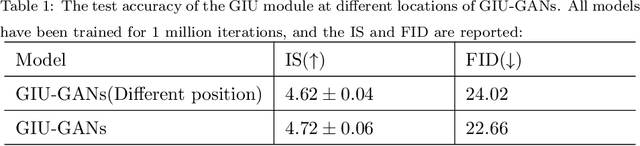

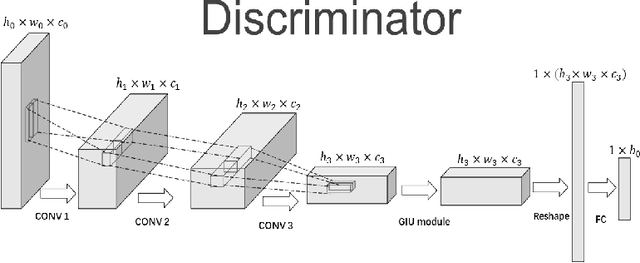

In recent years, with the rapid development of artificial intelligence, image generation based on deep learning has dramatically advanced. Image generation based on Generative Adversarial Networks (GANs) is a promising study. However, since convolutions are limited by spatial-agnostic and channel-specific, features extracted by traditional GANs based on convolution are constrained. Therefore, GANs are unable to capture any more details per image. On the other hand, straightforwardly stacking of convolutions causes too many parameters and layers in GANs, which will lead to a high risk of overfitting. To overcome the aforementioned limitations, in this paper, we propose a new GANs called Involution Generative Adversarial Networks (GIU-GANs). GIU-GANs leverages a brand new module called the Global Information Utilization (GIU) module, which integrates Squeeze-and-Excitation Networks (SENet) and involution to focus on global information by channel attention mechanism, leading to a higher quality of generated images. Meanwhile, Batch Normalization(BN) inevitably ignores the representation differences among noise sampled by the generator, and thus degrade the generated image quality. Thus we introduce Representative Batch Normalization(RBN) to the GANs architecture for this issue. The CIFAR-10 and CelebA datasets are employed to demonstrate the effectiveness of our proposed model. A large number of experiments prove that our model achieves state-of-the-art competitive performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge