"Image": models, code, and papers
Adversarial Robustness Through Local Lipschitzness
Apr 16, 2020
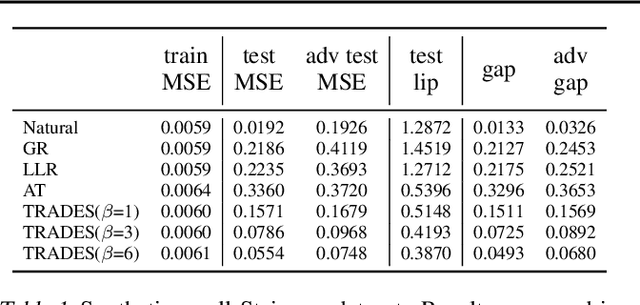
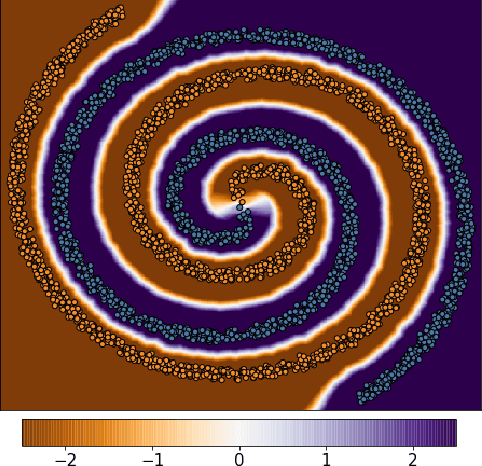
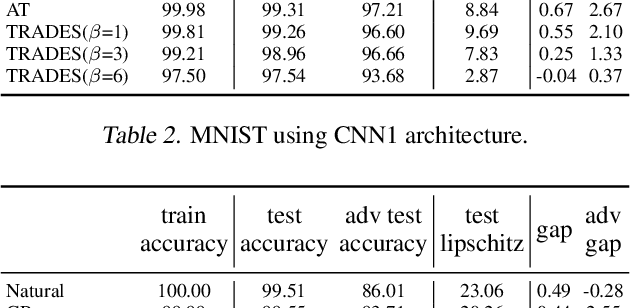
A standard method for improving the robustness of neural networks is adversarial training, where the network is trained on adversarial examples that are close to the training inputs. This produces classifiers that are robust, but it often decreases clean accuracy. Prior work even posits that the tradeoff between robustness and accuracy may be inevitable. We investigate this tradeoff in more depth through the lens of local Lipschitzness. In many image datasets, the classes are separated in the sense that images with different labels are not extremely close in $\ell_\infty$ distance. Using this separation as a starting point, we argue that it is possible to achieve both accuracy and robustness by encouraging the classifier to be locally smooth around the data. More precisely, we consider classifiers that are obtained by rounding locally Lipschitz functions. Theoretically, we show that such classifiers exist for any dataset such that there is a positive distance between the support of different classes. Empirically, we compare the local Lipschitzness of classifiers trained by several methods. Our results show that having a small Lipschitz constant correlates with achieving high clean and robust accuracy, and therefore, the smoothness of the classifier is an important property to consider in the context of adversarial examples. Code available at https://github.com/yangarbiter/robust-local-lipschitz .
EPOS: Estimating 6D Pose of Objects with Symmetries
Apr 01, 2020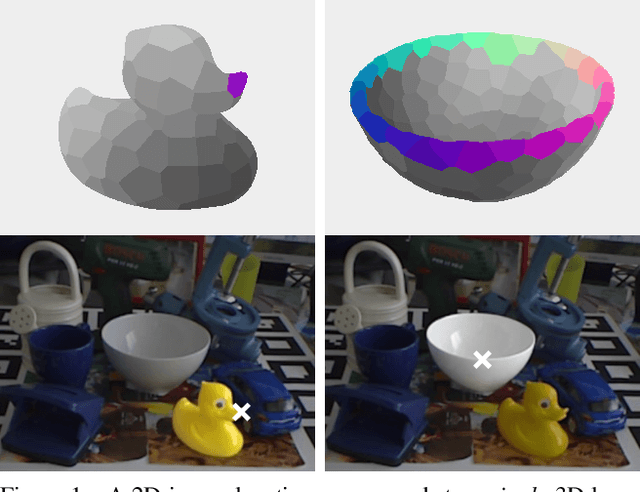
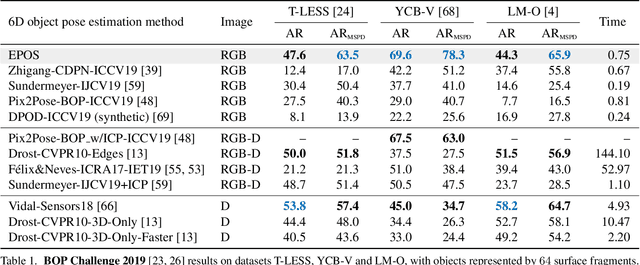
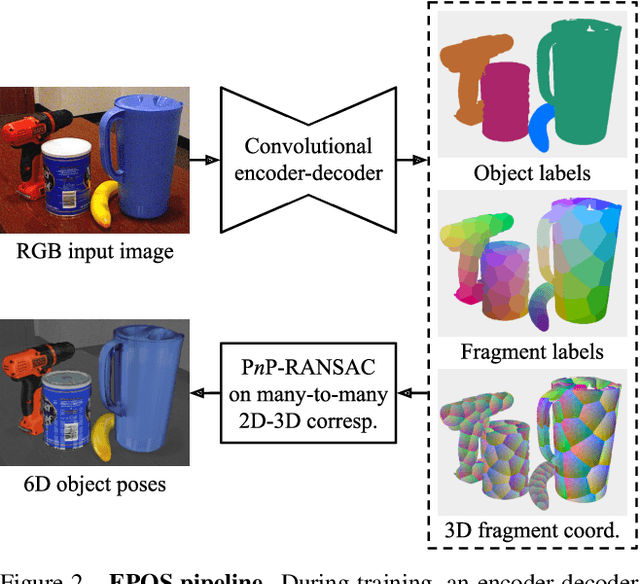
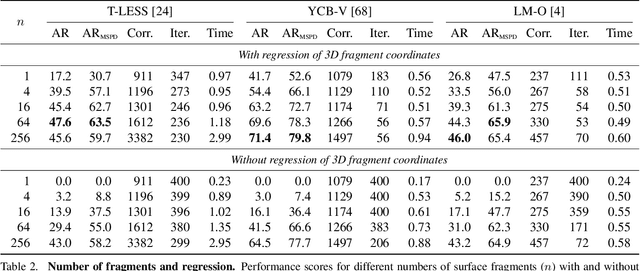
We present a new method for estimating the 6D pose of rigid objects with available 3D models from a single RGB input image. The method is applicable to a broad range of objects, including challenging ones with global or partial symmetries. An object is represented by compact surface fragments which allow handling symmetries in a systematic manner. Correspondences between densely sampled pixels and the fragments are predicted using an encoder-decoder network. At each pixel, the network predicts: (i) the probability of each object's presence, (ii) the probability of the fragments given the object's presence, and (iii) the precise 3D location on each fragment. A data-dependent number of corresponding 3D locations is selected per pixel, and poses of possibly multiple object instances are estimated using a robust and efficient variant of the PnP-RANSAC algorithm. In the BOP Challenge 2019, the method outperforms all RGB and most RGB-D and D methods on the T-LESS and LM-O datasets. On the YCB-V dataset, it is superior to all competitors, with a large margin over the second-best RGB method. Source code is at: cmp.felk.cvut.cz/epos.
Image Super-Resolution via Sparse Bayesian Modeling of Natural Images
Sep 19, 2012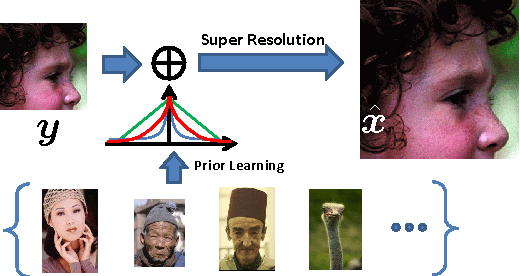
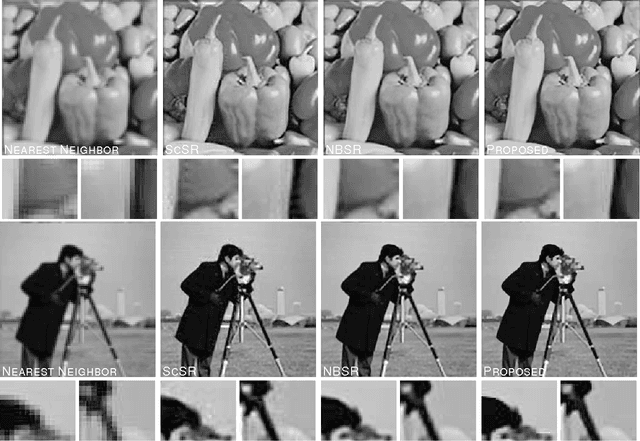


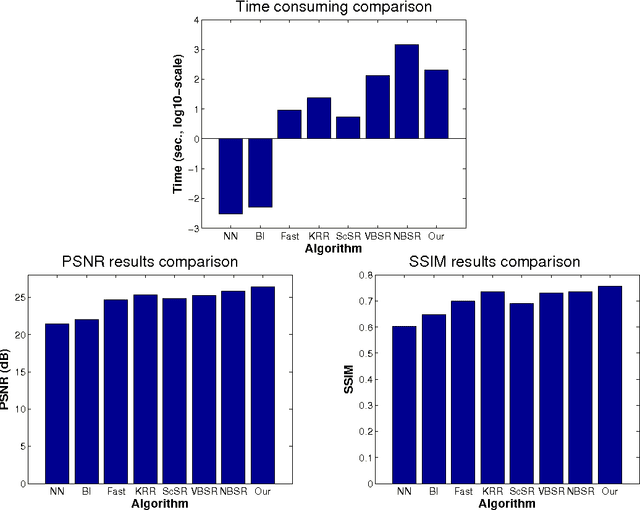
Image super-resolution (SR) is one of the long-standing and active topics in image processing community. A large body of works for image super resolution formulate the problem with Bayesian modeling techniques and then obtain its Maximum-A-Posteriori (MAP) solution, which actually boils down to a regularized regression task over separable regularization term. Although straightforward, this approach cannot exploit the full potential offered by the probabilistic modeling, as only the posterior mode is sought. Also, the separable property of the regularization term can not capture any correlations between the sparse coefficients, which sacrifices much on its modeling accuracy. We propose a Bayesian image SR algorithm via sparse modeling of natural images. The sparsity property of the latent high resolution image is exploited by introducing latent variables into the high-order Markov Random Field (MRF) which capture the content adaptive variance by pixel-wise adaptation. The high-resolution image is estimated via Empirical Bayesian estimation scheme, which is substantially faster than our previous approach based on Markov Chain Monte Carlo sampling [1]. It is shown that the actual cost function for the proposed approach actually incorporates a non-factorial regularization term over the sparse coefficients. Experimental results indicate that the proposed method can generate competitive or better results than \emph{state-of-the-art} SR algorithms.
Deep transformation models: Tackling complex regression problems with neural network based transformation models
Apr 01, 2020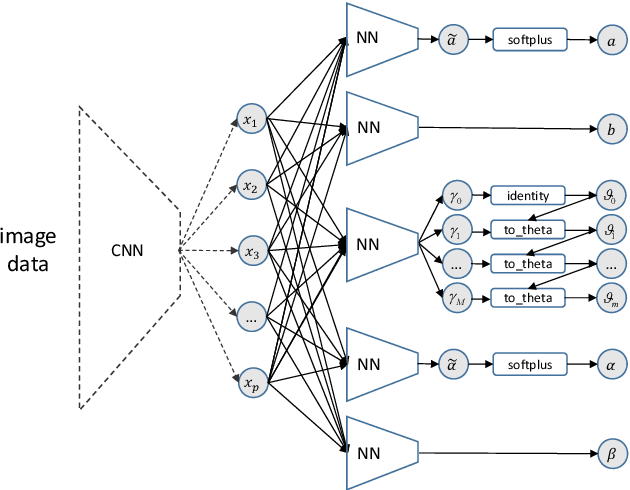
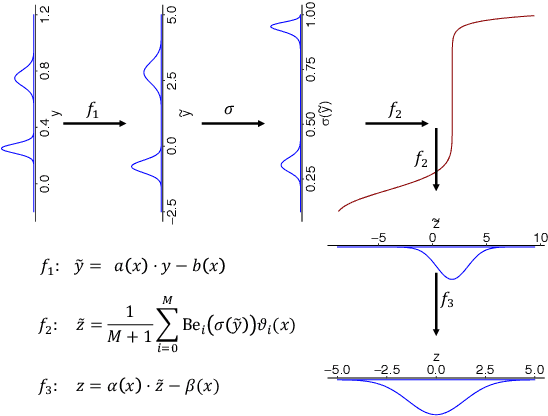
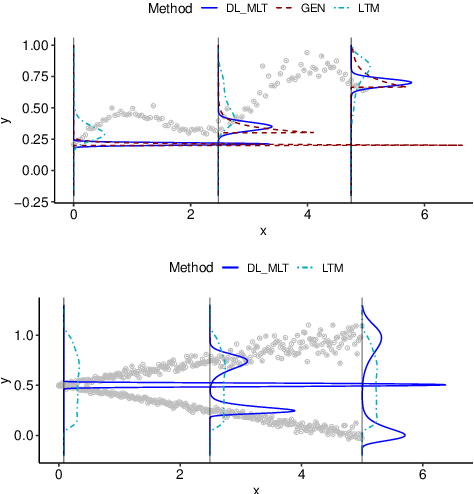
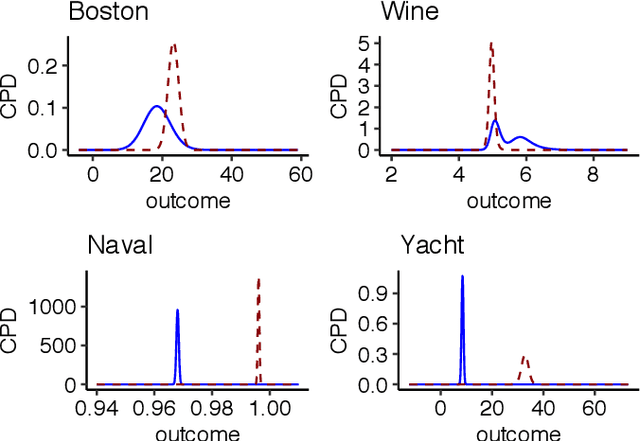
We present a deep transformation model for probabilistic regression. Deep learning is known for outstandingly accurate predictions on complex data but in regression tasks, it is predominantly used to just predict a single number. This ignores the non-deterministic character of most tasks. Especially if crucial decisions are based on the predictions, like in medical applications, it is essential to quantify the prediction uncertainty. The presented deep learning transformation model estimates the whole conditional probability distribution, which is the most thorough way to capture uncertainty about the outcome. We combine ideas from a statistical transformation model (most likely transformation) with recent transformation models from deep learning (normalizing flows) to predict complex outcome distributions. The core of the method is a parameterized transformation function which can be trained with the usual maximum likelihood framework using gradient descent. The method can be combined with existing deep learning architectures. For small machine learning benchmark datasets, we report state of the art performance for most dataset and partly even outperform it. Our method works for complex input data, which we demonstrate by employing a CNN architecture on image data.
A Method for Image Reduction Based on a Generalization of Ordered Weighted Averaging Functions
Jan 15, 2016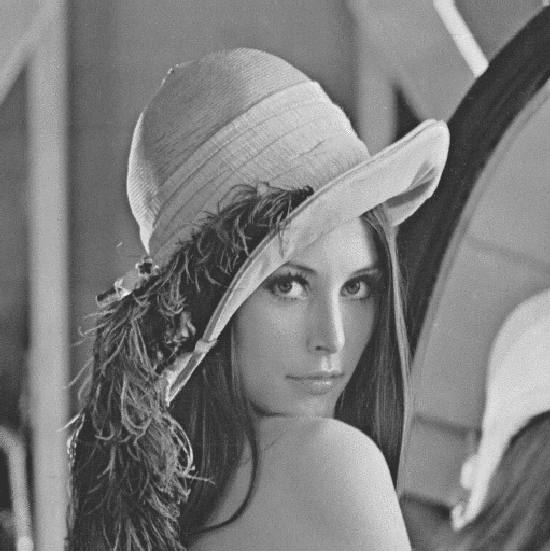
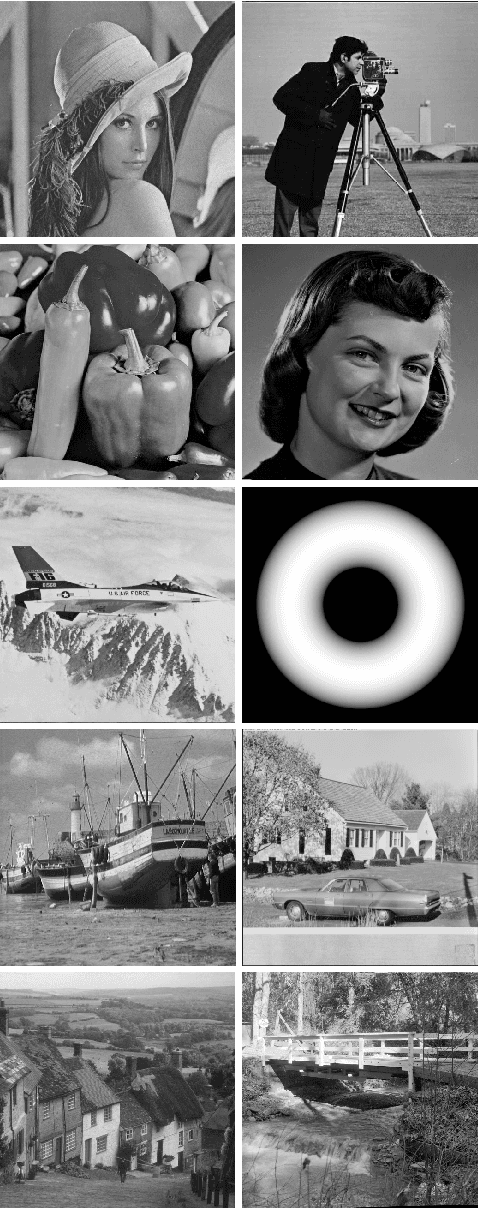
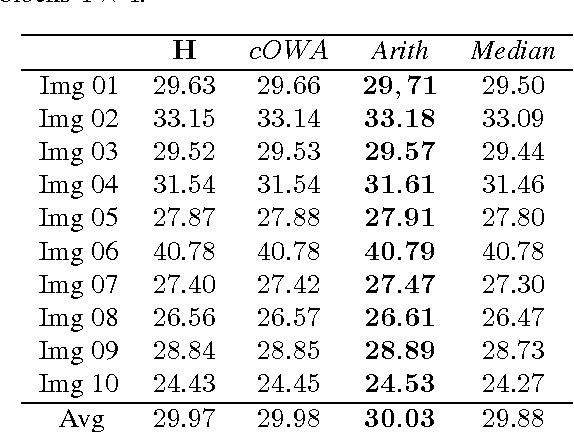

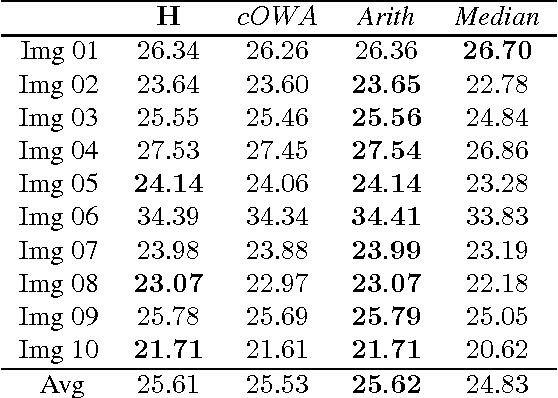
In this paper we propose a special type of aggregation function which generalizes the notion of Ordered Weighted Averaging Function - OWA. The resulting functions are called Dynamic Ordered Weighted Averaging Functions --- DYOWAs. This generalization will be developed in such way that the weight vectors are variables depending on the input vector. Particularly, this operators generalize the aggregation functions: Minimum, Maximum, Arithmetic Mean, Median, etc, which are extensively used in image processing. In this field of research two problems are considered: The determination of methods to reduce images and the construction of techniques which provide noise reduction. The operators described here are able to be used in both cases. In terms of image reduction we apply the methodology provided by Patermain et al. We use the noise reduction operators obtained here to treat the images obtained in the first part of the paper, thus obtaining images with better quality.
Real-Time Face and Landmark Localization for Eyeblink Detection
Jun 01, 2020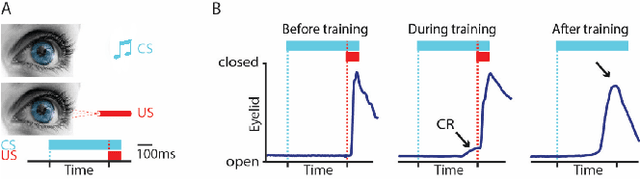
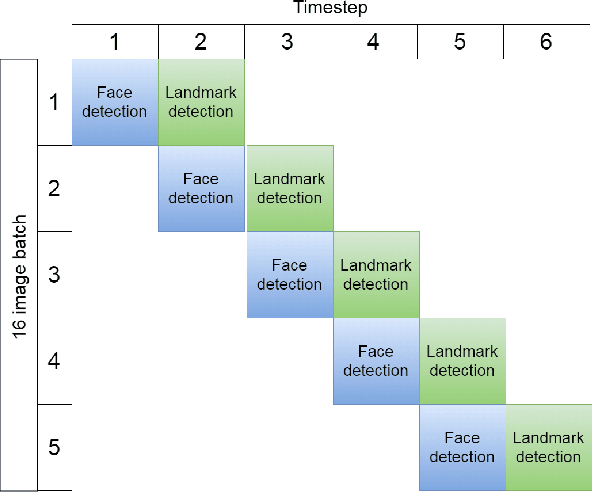
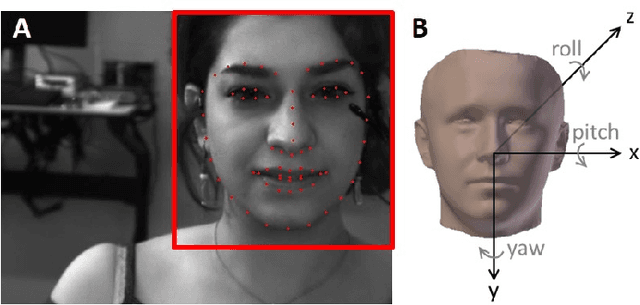
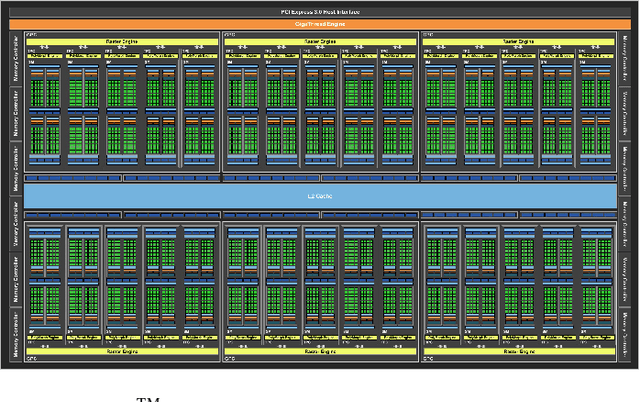
Pavlovian eyeblink conditioning is a powerful experiment used in the field of neuroscience to measure multiple aspects of how we learn in our daily life. To track the movement of the eyelid during an experiment, researchers have traditionally made use of potentiometers or electromyography. More recently, the use of computer vision and image processing alleviated the need for these techniques but currently employed methods require human intervention and are not fast enough to enable real-time processing. In this work, a face- and landmark-detection algorithm have been carefully combined in order to provide fully automated eyelid tracking, and have further been accelerated to make the first crucial step towards online, closed-loop experiments. Such experiments have not been achieved so far and are expected to offer significant insights in the workings of neurological and psychiatric disorders. Based on an extensive literature search, various different algorithms for face detection and landmark detection have been analyzed and evaluated. Two algorithms were identified as most suitable for eyelid detection: the Histogram-of-Oriented-Gradients (HOG) algorithm for face detection and the Ensemble-of-Regression-Trees (ERT) algorithm for landmark detection. These two algorithms have been accelerated on GPU and CPU, achieving speedups of 1,753$\times$ and 11$\times$, respectively. To demonstrate the usefulness of our eyelid-detection algorithm, a research hypothesis was formed and a well-established neuroscientific experiment was employed: eyeblink detection. Our experimental evaluation reveals an overall application runtime of 0.533 ms per frame, which is 1,101$\times$ faster than the sequential implementation and well within the real-time requirements of eyeblink conditioning in humans, i.e. faster than 500 frames per second.
Methods and open-source toolkit for analyzing and visualizing challenge results
Oct 11, 2019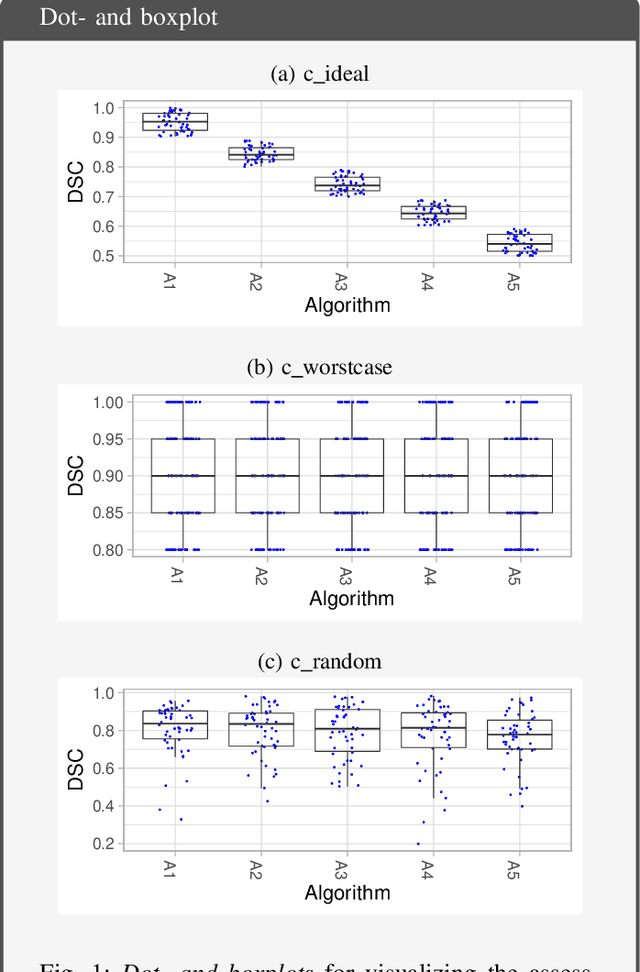
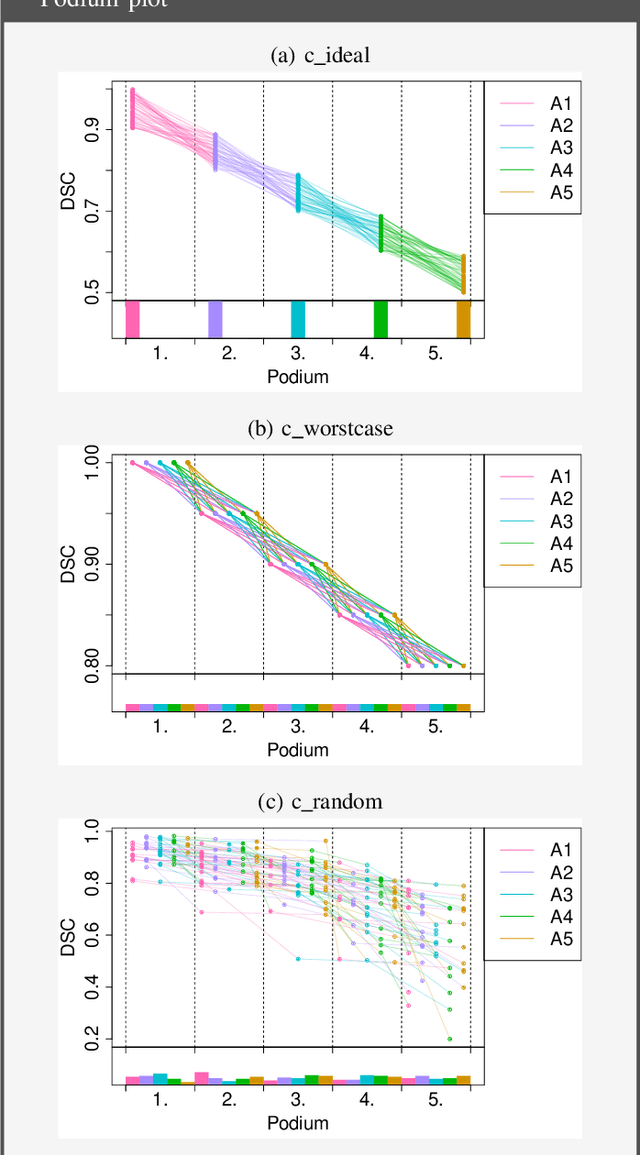
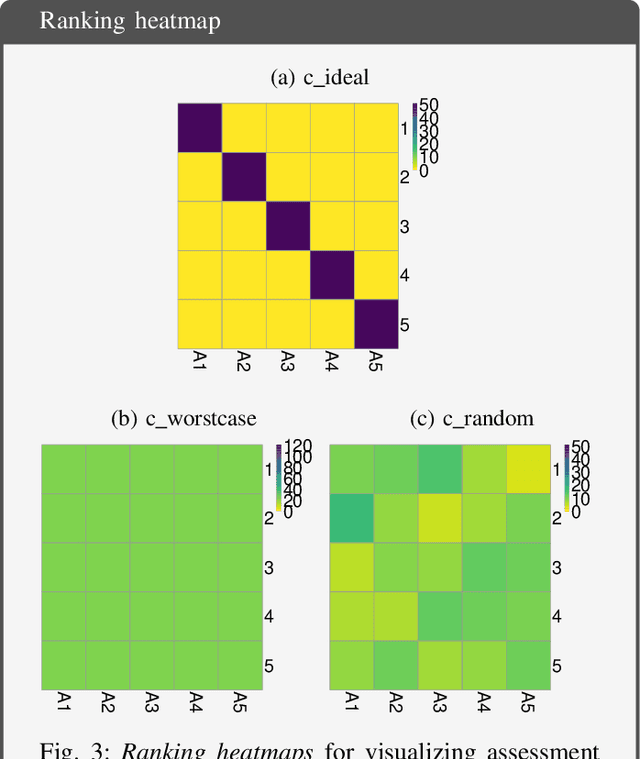
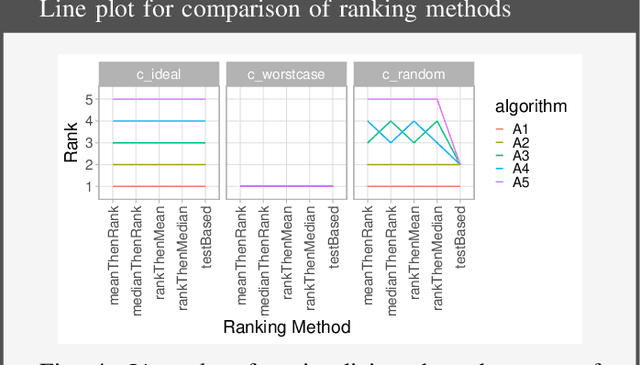
Biomedical challenges have become the de facto standard for benchmarking biomedical image analysis algorithms. While the number of challenges is steadily increasing, surprisingly little effort has been invested in ensuring high quality design, execution and reporting for these international competitions. Specifically, results analysis and visualization in the event of uncertainties have been given almost no attention in the literature. Given these shortcomings, the contribution of this paper is two-fold: (1) We present a set of methods to comprehensively analyze and visualize the results of single-task and multi-task challenges and apply them to a number of simulated and real-life challenges to demonstrate their specific strengths and weaknesses; (2) We release the open-source framework challengeR as part of this work to enable fast and wide adoption of the methodology proposed in this paper. Our approach offers an intuitive way to gain important insights into the relative and absolute performance of algorithms, which cannot be revealed by commonly applied visualization techniques. This is demonstrated by the experiments performed within this work. Our framework could thus become an important tool for analyzing and visualizing challenge results in the field of biomedical image analysis and beyond.
Neural Status Registers
Apr 15, 2020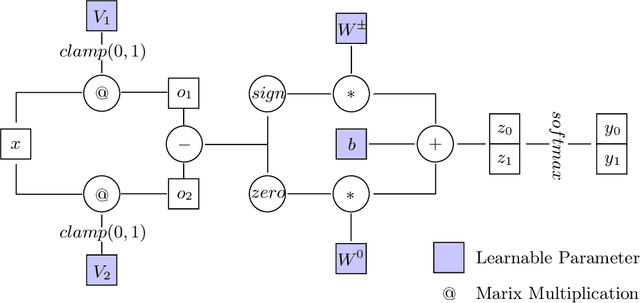
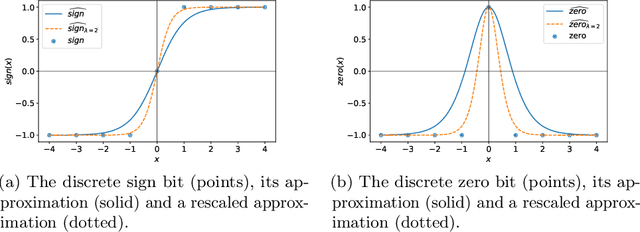
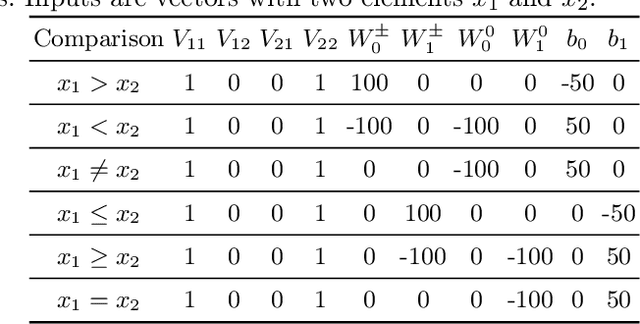
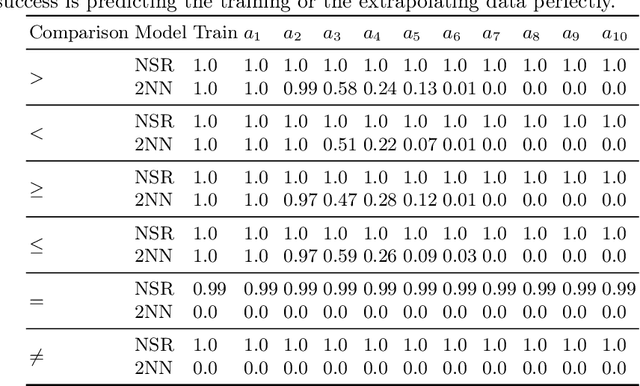
Neural networks excel at approximating functions and finding patterns in complex and challenging domains. Yet, they fail to learn simple but precise computation. Recent work addressed the ability to add, subtract, and multiply numbers but is lacking a component to drive control flow. True computer intelligence should also be able to decide when to perform what operation. In this paper, we introduce the Neural Status Register (NSR), inspired by physical Status Registers. At the heart of the NSR are arithmetic comparisons between inputs. With theoretically principled changes to physical Status Registers, the NSR allows end-to-end differentiation and learns such comparisons reliably. But the NSR also extrapolates: it generalizes to unseen data distributions. For example, the NSR trains on single digits and correctly predicts numbers that are up to 14 orders of magnitude larger. This suggests that the NSR captures the true underlying arithmetic. In follow-up experiments, we use the NSR to control the computation of a downstream arithmetic unit to learn piecewise functions. We can also learn more challenging tasks through redundancy. Finally, we use the NSR to learn an upstream convolutional neural network to compare images of MNIST digits to decide which image contains the larger digit.
PAG-Net: Progressive Attention Guided Depth Super-resolution Network
Nov 22, 2019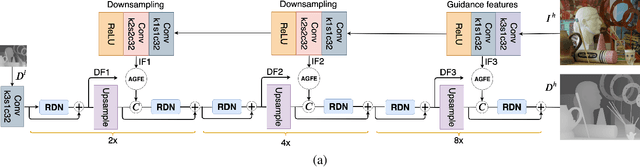
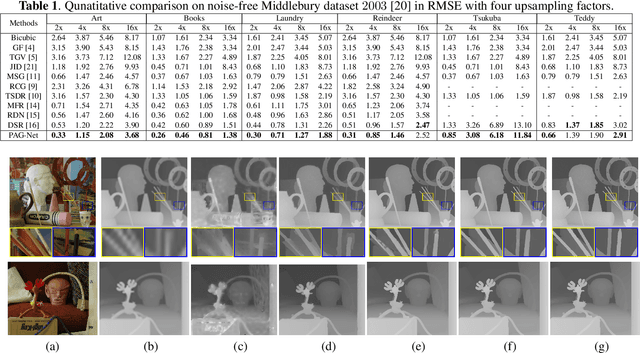
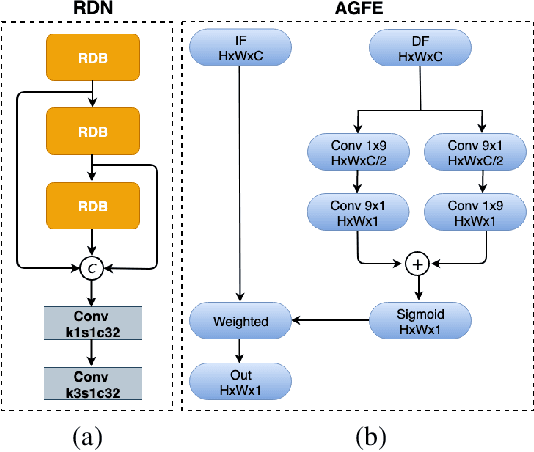

In this paper, we propose a novel method for the challenging problem of guided depth map super-resolution, called PAGNet. It is based on residual dense networks and involves the attention mechanism to suppress the texture copying problem arises due to improper guidance by RGB images. The attention module mainly involves providing the spatial attention to guidance image based on the depth features. We evaluate the proposed trained models on test dataset and provide comparisons with the state-of-the-art depth super-resolution methods.
4DFlowNet: Super-Resolution 4D Flow MRI using Deep Learning and Computational Fluid Dynamics
Apr 15, 2020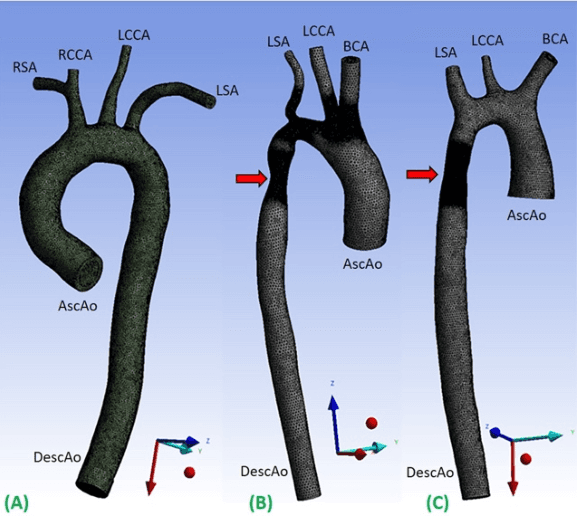
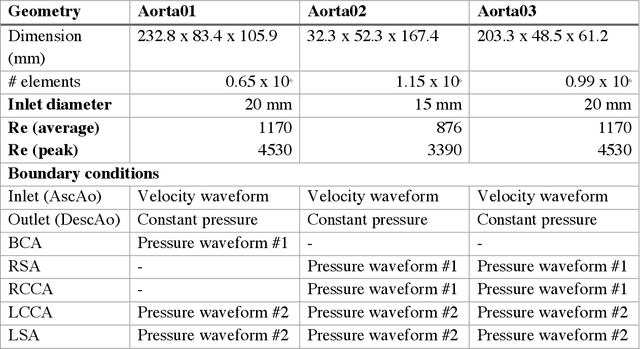
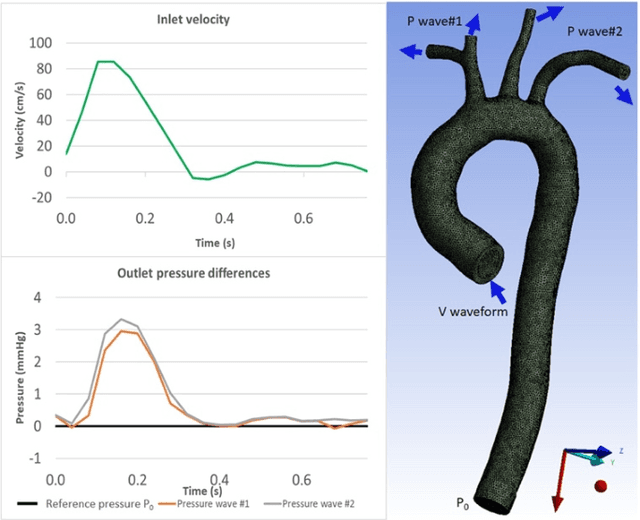
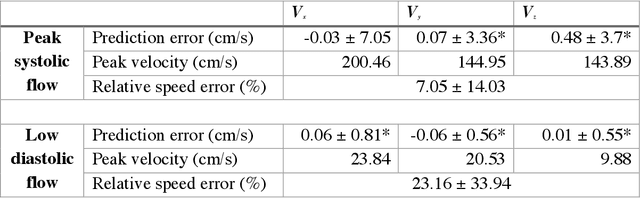
4D-flow magnetic resonance imaging (MRI) is an emerging imaging technique where spatiotemporal 3D blood velocity can be captured with full volumetric coverage in a single non-invasive examination. This enables qualitative and quantitative analysis of hemodynamic flow parameters of the heart and great vessels. An increase in the image resolution would provide more accuracy and allow better assessment of the blood flow, especially for patients with abnormal flows. However, this must be balanced with increasing imaging time. The recent success of deep learning in generating super resolution images shows promise for implementation in medical images. We utilized computational fluid dynamics simulations to generate fluid flow simulations and represent them as synthetic 4D flow MRI data. We built our training dataset to mimic actual 4D flow MRI data with its corresponding noise distribution. Our novel 4DFlowNet network was trained on this synthetic 4D flow data and was capable in producing noise-free super resolution 4D flow phase images with upsample factor of 2. We also tested the 4DFlowNet in actual 4D flow MR images of a phantom and normal volunteer data, and demonstrated comparable results with the actual flow rate measurements giving an absolute relative error of 0.6 to 5.8% and 1.1 to 3.8% in the phantom data and normal volunteer data, respectively.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge