"Image": models, code, and papers
Learning the Regularization in DCE-MR Image Reconstruction for Functional Imaging of Kidneys
Sep 15, 2021
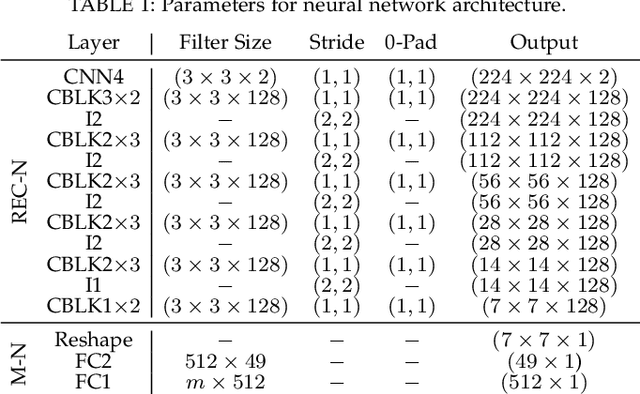
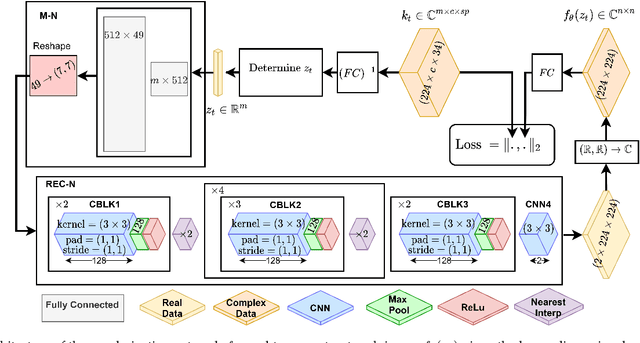
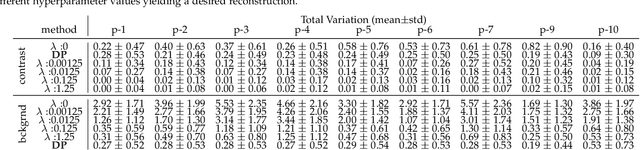
Kidney DCE-MRI aims at both qualitative assessment of kidney anatomy and quantitative assessment of kidney function by estimating the tracer kinetic (TK) model parameters. Accurate estimation of TK model parameters requires an accurate measurement of the arterial input function (AIF) with high temporal resolution. Accelerated imaging is used to achieve high temporal resolution, which yields under-sampling artifacts in the reconstructed images. Compressed sensing (CS) methods offer a variety of reconstruction options. Most commonly, sparsity of temporal differences is encouraged for regularization to reduce artifacts. Increasing regularization in CS methods removes the ambient artifacts but also over-smooths the signal temporally which reduces the parameter estimation accuracy. In this work, we propose a single image trained deep neural network to reduce MRI under-sampling artifacts without reducing the accuracy of functional imaging markers. Instead of regularizing with a penalty term in optimization, we promote regularization by generating images from a lower dimensional representation. In this manuscript we motivate and explain the lower dimensional input design. We compare our approach to CS reconstructions with multiple regularization weights. Proposed approach results in kidney biomarkers that are highly correlated with the ground truth markers estimated using the CS reconstruction which was optimized for functional analysis. At the same time, the proposed approach reduces the artifacts in the reconstructed images.
ECG Heart-beat Classification Using Multimodal Image Fusion
May 28, 2021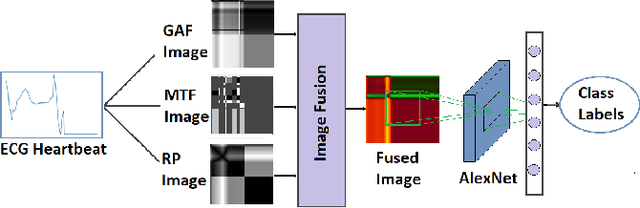
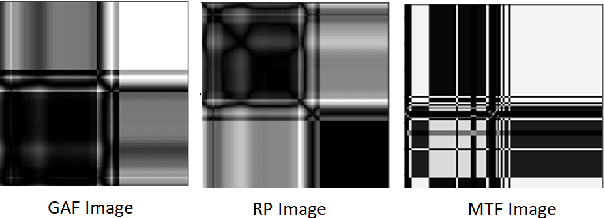
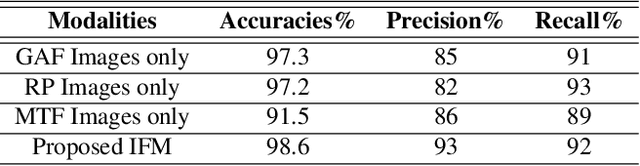

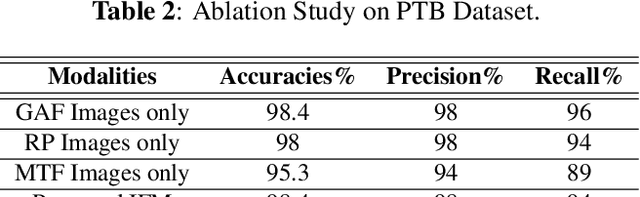
In this paper, we present a novel Image Fusion Model (IFM) for ECG heart-beat classification to overcome the weaknesses of existing machine learning techniques that rely either on manual feature extraction or direct utilization of 1D raw ECG signal. At the input of IFM, we first convert the heart beats of ECG into three different images using Gramian Angular Field (GAF), Recurrence Plot (RP) and Markov Transition Field (MTF) and then fuse these images to create a single imaging modality. We use AlexNet for feature extraction and classification and thus employ end to end deep learning. We perform experiments on PhysioNet MIT-BIH dataset for five different arrhythmias in accordance with the AAMI EC57 standard and on PTB diagnostics dataset for myocardial infarction (MI) classification. We achieved an state of an art results in terms of prediction accuracy, precision and recall.
Personalized visual encoding model construction with small data
Feb 04, 2022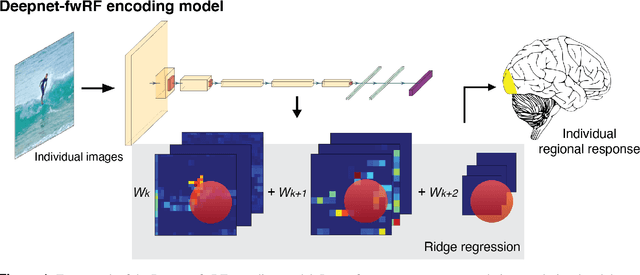
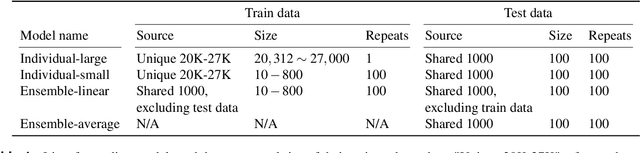
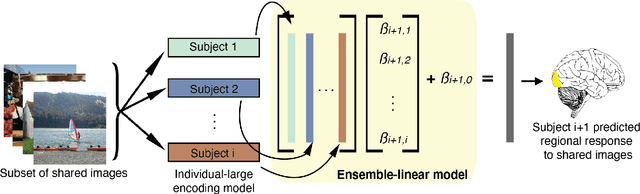
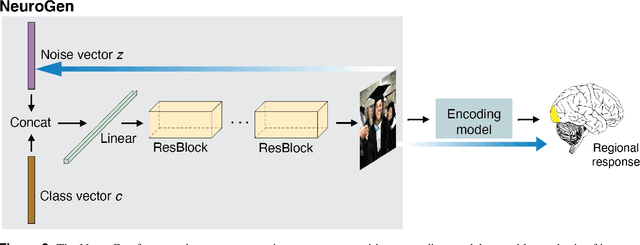
Encoding models that predict brain response patterns to stimuli are one way to capture this relationship between variability in bottom-up neural systems and individual's behavior or pathological state. However, they generally need a large amount of training data to achieve optimal accuracy. Here, we propose and test an alternative personalized ensemble encoding model approach to utilize existing encoding models, to create encoding models for novel individuals with relatively little stimuli-response data. We show that these personalized ensemble encoding models trained with small amounts of data for a specific individual, i.e. ~400 image-response pairs, achieve accuracy not different from models trained on ~24,000 image-response pairs for the same individual. Importantly, the personalized ensemble encoding models preserve patterns of inter-individual variability in the image-response relationship. Additionally, we use our personalized ensemble encoding model within the recently developed NeuroGen framework to generate optimal stimuli designed to maximize specific regions' activations for a specific individual. We show that the inter-individual differences in face area responses to images of dog vs human faces observed previously is replicated using NeuroGen with the ensemble encoding model. Finally, and most importantly, we show the proposed approach is robust against domain shift by validating on a prospectively collected set of image-response data in novel individuals with a different scanner and experimental setup. Our approach shows the potential to use previously collected, deeply sampled data to efficiently create accurate, personalized encoding models and, subsequently, personalized optimal synthetic images for new individuals scanned under different experimental conditions.
Rate Distortion Characteristic Modeling for Neural Image Compression
Jun 24, 2021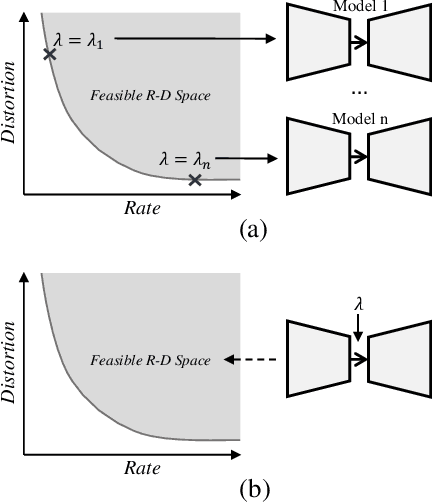

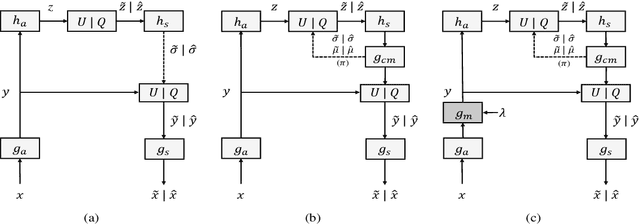
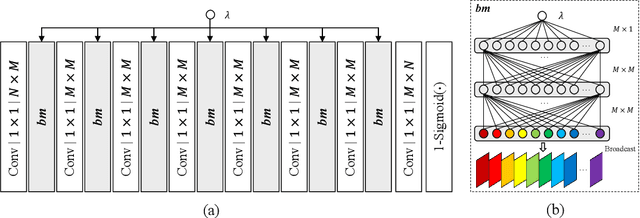
End-to-end optimization capability offers neural image compression (NIC) superior lossy compression performance. However, distinct models are required to be trained to reach different points in the rate-distortion (R-D) space. In this paper, we consider the problem of R-D characteristic analysis and modeling for NIC. We make efforts to formulate the essential mathematical functions to describe the R-D behavior of NIC using deep network and statistical modeling. Thus continuous bit-rate points could be elegantly realized by leveraging such model via a single trained network. In this regard, we propose a plugin-in module to learn the relationship between the target bit-rate and the binary representation for the latent variable of auto-encoder. Furthermore, we model the rate and distortion characteristic of NIC as a function of the coding parameter $\lambda$ respectively. Our experiments show our proposed method is easy to adopt and obtains competitive coding performance with fixed-rate coding approaches, which would benefit the practical deployment of NIC. In addition, the proposed model could be applied to NIC rate control with limited bit-rate error using a single network.
ST-FL: Style Transfer Preprocessing in Federated Learning for COVID-19 Segmentation
Mar 25, 2022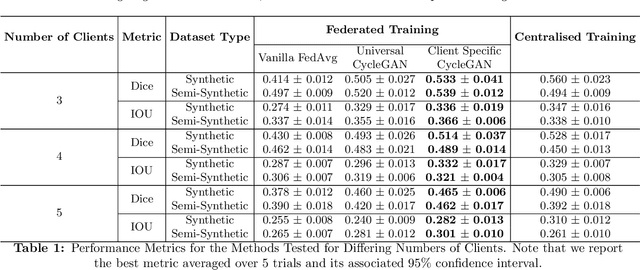
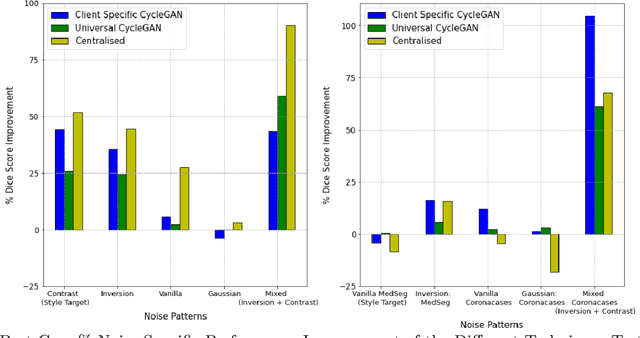
Chest Computational Tomography (CT) scans present low cost, speed and objectivity for COVID-19 diagnosis and deep learning methods have shown great promise in assisting the analysis and interpretation of these images. Most hospitals or countries can train their own models using in-house data, however empirical evidence shows that those models perform poorly when tested on new unseen cases, surfacing the need for coordinated global collaboration. Due to privacy regulations, medical data sharing between hospitals and nations is extremely difficult. We propose a GAN-augmented federated learning model, dubbed ST-FL (Style Transfer Federated Learning), for COVID-19 image segmentation. Federated learning (FL) permits a centralised model to be learned in a secure manner from heterogeneous datasets located in disparate private data silos. We demonstrate that the widely varying data quality on FL client nodes leads to a sub-optimal centralised FL model for COVID-19 chest CT image segmentation. ST-FL is a novel FL framework that is robust in the face of highly variable data quality at client nodes. The robustness is achieved by a denoising CycleGAN model at each client of the federation that maps arbitrary quality images into the same target quality, counteracting the severe data variability evident in real-world FL use-cases. Each client is provided with the target style, which is the same for all clients, and trains their own denoiser. Our qualitative and quantitative results suggest that this FL model performs comparably to, and in some cases better than, a model that has centralised access to all the training data.
StEP: Style-based Encoder Pre-training for Multi-modal Image Synthesis
Apr 14, 2021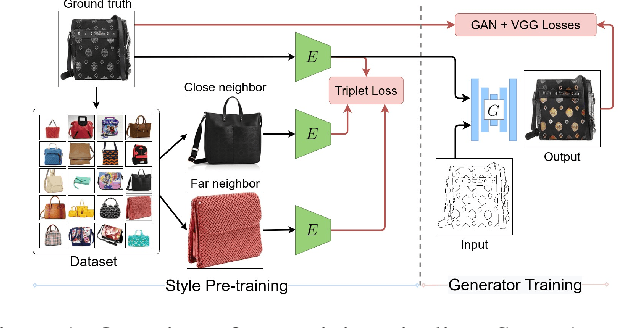
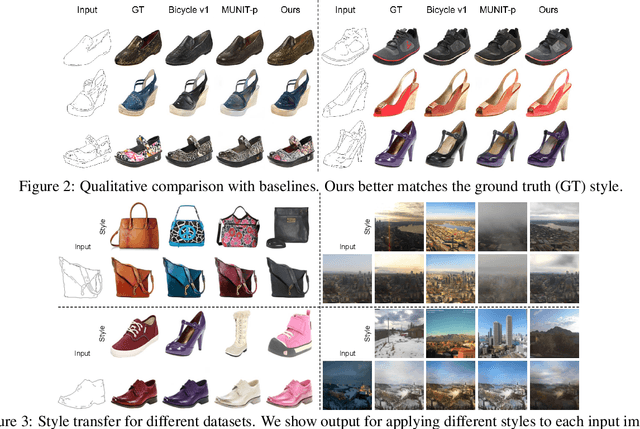
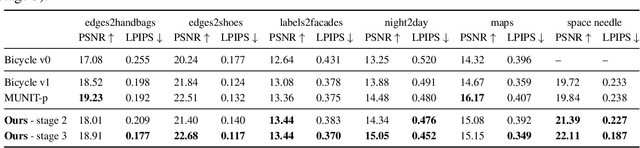

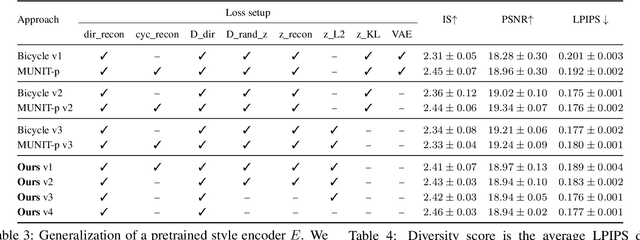
We propose a novel approach for multi-modal Image-to-image (I2I) translation. To tackle the one-to-many relationship between input and output domains, previous works use complex training objectives to learn a latent embedding, jointly with the generator, that models the variability of the output domain. In contrast, we directly model the style variability of images, independent of the image synthesis task. Specifically, we pre-train a generic style encoder using a novel proxy task to learn an embedding of images, from arbitrary domains, into a low-dimensional style latent space. The learned latent space introduces several advantages over previous traditional approaches to multi-modal I2I translation. First, it is not dependent on the target dataset, and generalizes well across multiple domains. Second, it learns a more powerful and expressive latent space, which improves the fidelity of style capture and transfer. The proposed style pre-training also simplifies the training objective and speeds up the training significantly. Furthermore, we provide a detailed study of the contribution of different loss terms to the task of multi-modal I2I translation, and propose a simple alternative to VAEs to enable sampling from unconstrained latent spaces. Finally, we achieve state-of-the-art results on six challenging benchmarks with a simple training objective that includes only a GAN loss and a reconstruction loss.
Accelerated MRI With Deep Linear Convolutional Transform Learning
Apr 17, 2022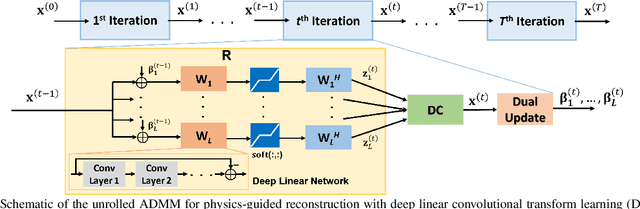
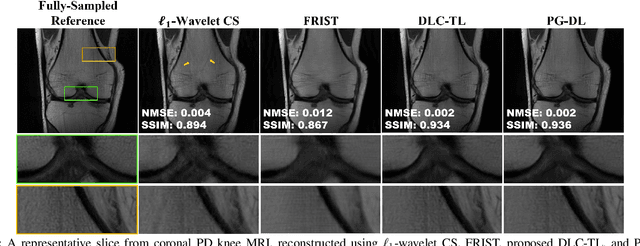
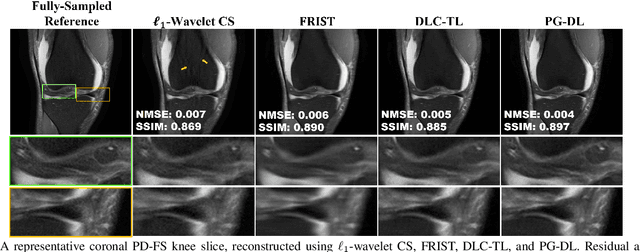
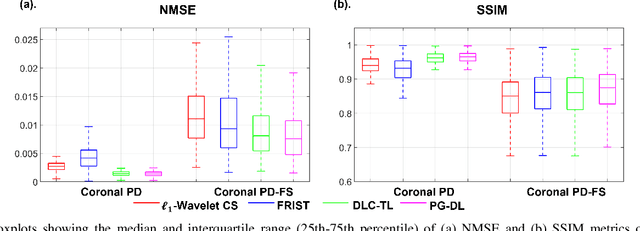
Recent studies show that deep learning (DL) based MRI reconstruction outperforms conventional methods, such as parallel imaging and compressed sensing (CS), in multiple applications. Unlike CS that is typically implemented with pre-determined linear representations for regularization, DL inherently uses a non-linear representation learned from a large database. Another line of work uses transform learning (TL) to bridge the gap between these two approaches by learning linear representations from data. In this work, we combine ideas from CS, TL and DL reconstructions to learn deep linear convolutional transforms as part of an algorithm unrolling approach. Using end-to-end training, our results show that the proposed technique can reconstruct MR images to a level comparable to DL methods, while supporting uniform undersampling patterns unlike conventional CS methods. Our proposed method relies on convex sparse image reconstruction with linear representation at inference time, which may be beneficial for characterizing robustness, stability and generalizability.
Semantic Distillation Guided Salient Object Detection
Mar 08, 2022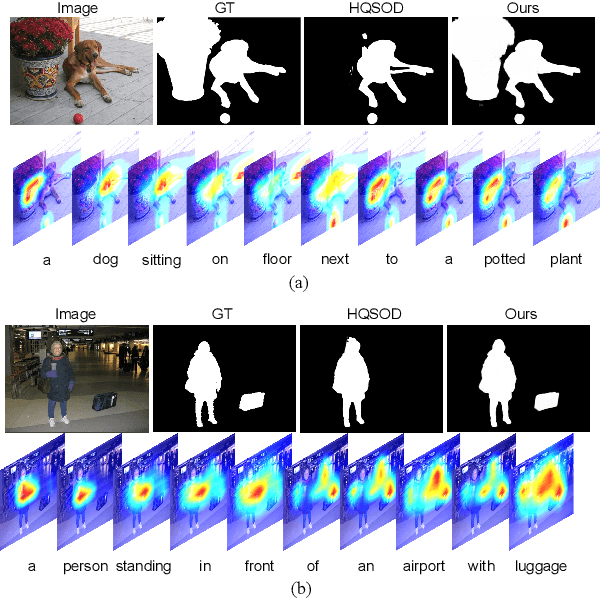
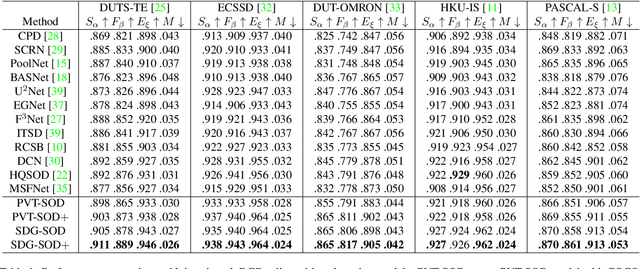
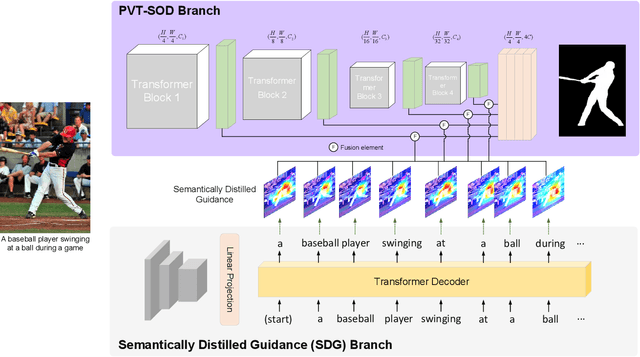
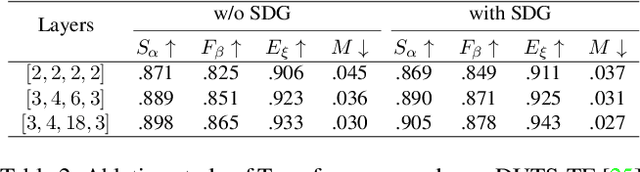
Most existing CNN-based salient object detection methods can identify local segmentation details like hair and animal fur, but often misinterpret the real saliency due to the lack of global contextual information caused by the subjectiveness of the SOD task and the locality of convolution layers. Moreover, due to the unrealistically expensive labeling costs, the current existing SOD datasets are insufficient to cover the real data distribution. The limitation and bias of the training data add additional difficulty to fully exploring the semantic association between object-to-object and object-to-environment in a given image. In this paper, we propose a semantic distillation guided SOD (SDG-SOD) method that produces accurate results by fusing semantically distilled knowledge from generated image captioning into the Vision-Transformer-based SOD framework. SDG-SOD can better uncover inter-objects and object-to-environment saliency and cover the gap between the subjective nature of SOD and its expensive labeling. Comprehensive experiments on five benchmark datasets demonstrate that the SDG-SOD outperforms the state-of-the-art approaches on four evaluation metrics, and largely improves the model performance on DUTS, ECSSD, DUT, HKU-IS, and PASCAL-S datasets.
Low-rank features based double transformation matrices learning for image classification
Jan 28, 2022Linear regression is a supervised method that has been widely used in classification tasks. In order to apply linear regression to classification tasks, a technique for relaxing regression targets was proposed. However, methods based on this technique ignore the pressure on a single transformation matrix due to the complex information contained in the data. A single transformation matrix in this case is too strict to provide a flexible projection, thus it is necessary to adopt relaxation on transformation matrix. This paper proposes a double transformation matrices learning method based on latent low-rank feature extraction. The core idea is to use double transformation matrices for relaxation, and jointly projecting the learned principal and salient features from two directions into the label space, which can share the pressure of a single transformation matrix. Firstly, the low-rank features are learned by the latent low rank representation (LatLRR) method which processes the original data from two directions. In this process, sparse noise is also separated, which alleviates its interference on projection learning to some extent. Then, two transformation matrices are introduced to process the two features separately, and the information useful for the classification is extracted. Finally, the two transformation matrices can be easily obtained by alternate optimization methods. Through such processing, even when a large amount of redundant information is contained in samples, our method can also obtain projection results that are easy to classify. Experiments on multiple data sets demonstrate the effectiveness of our approach for classification, especially for complex scenarios.
Unlabeled Data Guided Semi-supervised Histopathology Image Segmentation
Dec 17, 2020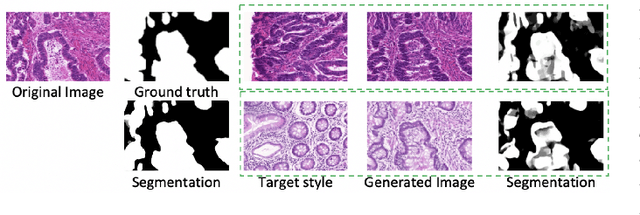
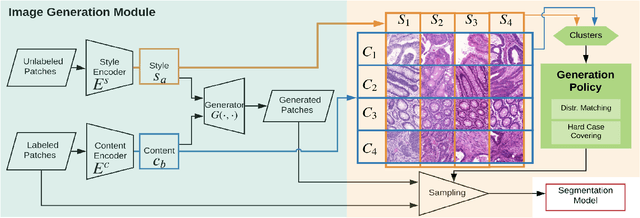
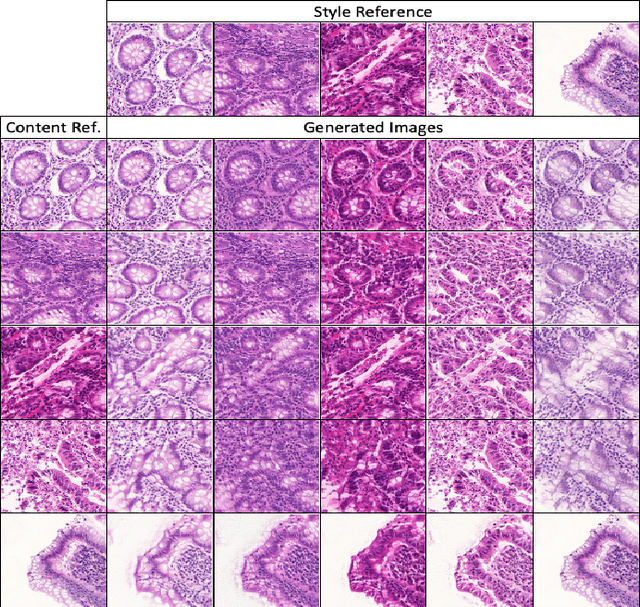
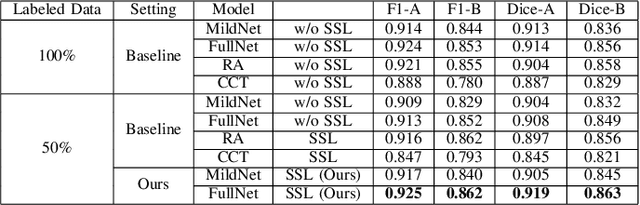
Automatic histopathology image segmentation is crucial to disease analysis. Limited available labeled data hinders the generalizability of trained models under the fully supervised setting. Semi-supervised learning (SSL) based on generative methods has been proven to be effective in utilizing diverse image characteristics. However, it has not been well explored what kinds of generated images would be more useful for model training and how to use such images. In this paper, we propose a new data guided generative method for histopathology image segmentation by leveraging the unlabeled data distributions. First, we design an image generation module. Image content and style are disentangled and embedded in a clustering-friendly space to utilize their distributions. New images are synthesized by sampling and cross-combining contents and styles. Second, we devise an effective data selection policy for judiciously sampling the generated images: (1) to make the generated training set better cover the dataset, the clusters that are underrepresented in the original training set are covered more; (2) to make the training process more effective, we identify and oversample the images of "hard cases" in the data for which annotated training data may be scarce. Our method is evaluated on glands and nuclei datasets. We show that under both the inductive and transductive settings, our SSL method consistently boosts the performance of common segmentation models and attains state-of-the-art results.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge