Steen Moeller
Accelerated MRI With Deep Linear Convolutional Transform Learning
Apr 17, 2022
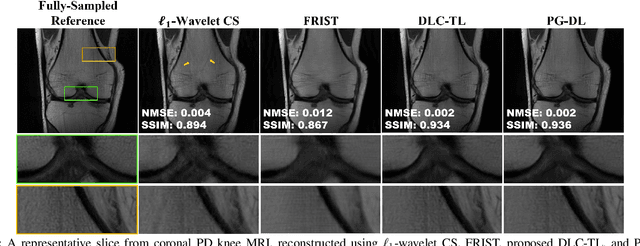
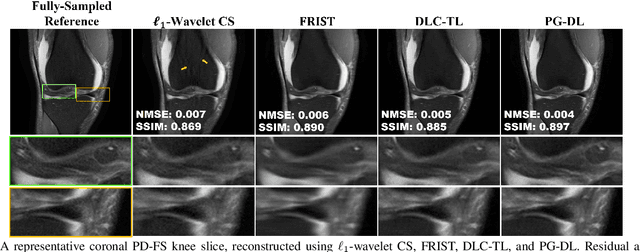
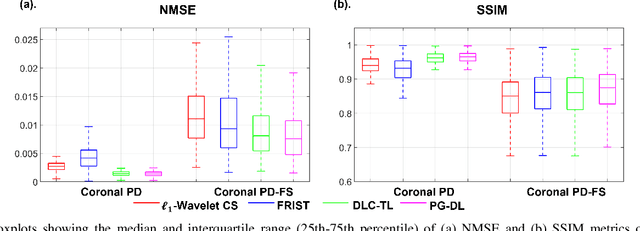
Abstract:Recent studies show that deep learning (DL) based MRI reconstruction outperforms conventional methods, such as parallel imaging and compressed sensing (CS), in multiple applications. Unlike CS that is typically implemented with pre-determined linear representations for regularization, DL inherently uses a non-linear representation learned from a large database. Another line of work uses transform learning (TL) to bridge the gap between these two approaches by learning linear representations from data. In this work, we combine ideas from CS, TL and DL reconstructions to learn deep linear convolutional transforms as part of an algorithm unrolling approach. Using end-to-end training, our results show that the proposed technique can reconstruct MR images to a level comparable to DL methods, while supporting uniform undersampling patterns unlike conventional CS methods. Our proposed method relies on convex sparse image reconstruction with linear representation at inference time, which may be beneficial for characterizing robustness, stability and generalizability.
20-fold Accelerated 7T fMRI Using Referenceless Self-Supervised Deep Learning Reconstruction
May 12, 2021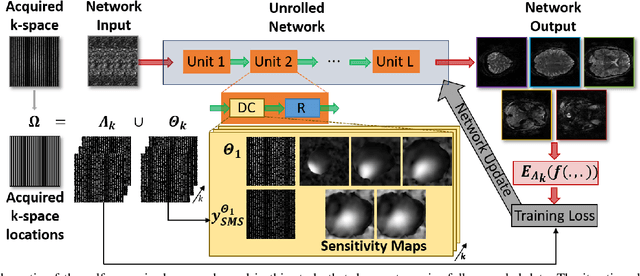
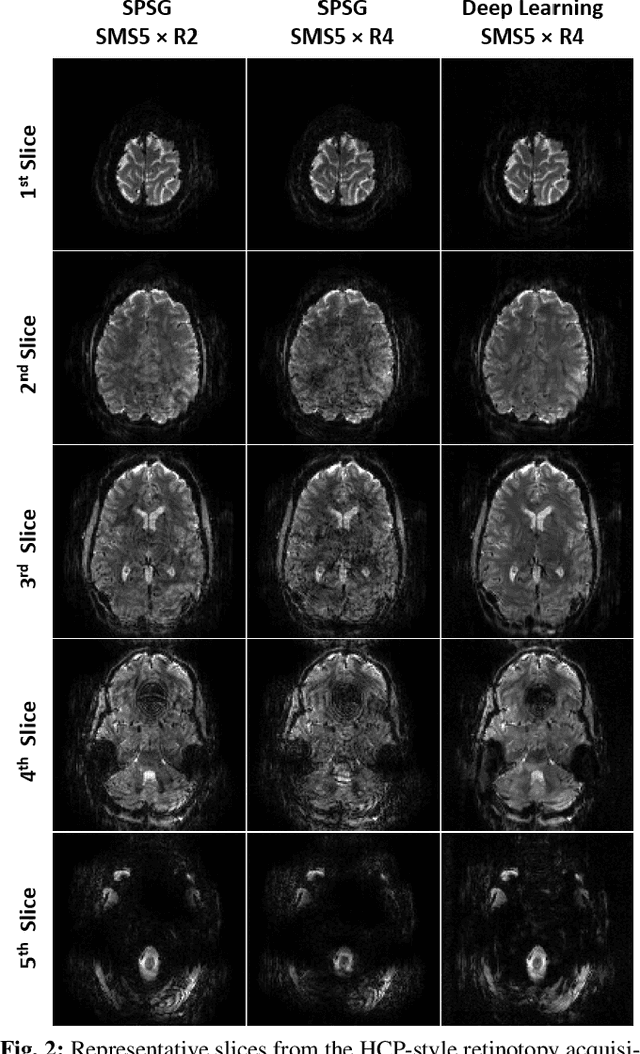
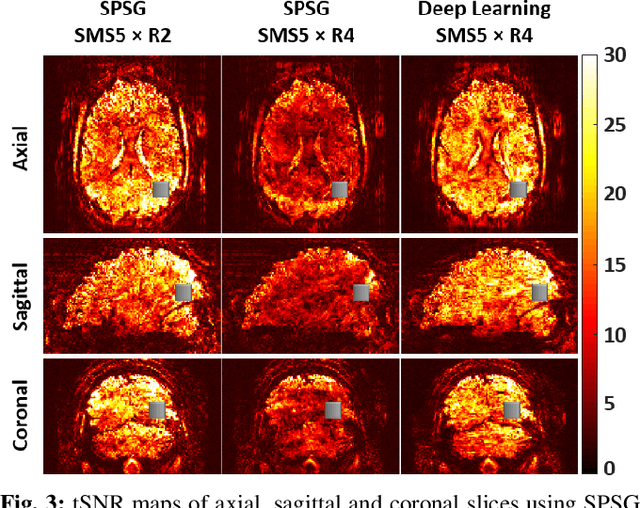
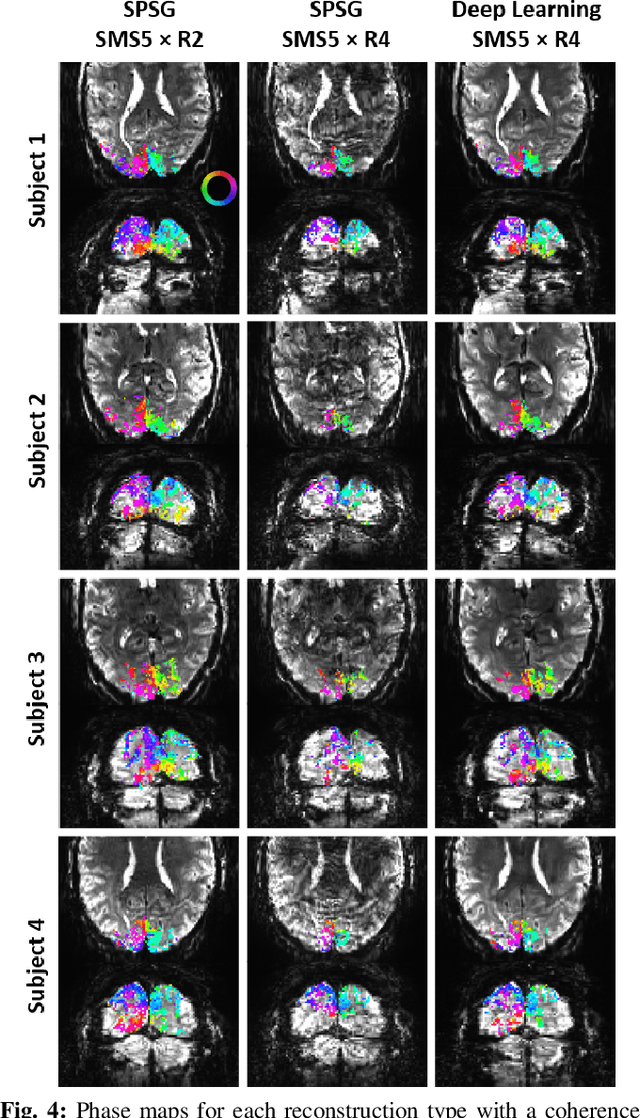
Abstract:High spatial and temporal resolution across the whole brain is essential to accurately resolve neural activities in fMRI. Therefore, accelerated imaging techniques target improved coverage with high spatio-temporal resolution. Simultaneous multi-slice (SMS) imaging combined with in-plane acceleration are used in large studies that involve ultrahigh field fMRI, such as the Human Connectome Project. However, for even higher acceleration rates, these methods cannot be reliably utilized due to aliasing and noise artifacts. Deep learning (DL) reconstruction techniques have recently gained substantial interest for improving highly-accelerated MRI. Supervised learning of DL reconstructions generally requires fully-sampled training datasets, which is not available for high-resolution fMRI studies. To tackle this challenge, self-supervised learning has been proposed for training of DL reconstruction with only undersampled datasets, showing similar performance to supervised learning. In this study, we utilize a self-supervised physics-guided DL reconstruction on a 5-fold SMS and 4-fold in-plane accelerated 7T fMRI data. Our results show that our self-supervised DL reconstruction produce high-quality images at this 20-fold acceleration, substantially improving on existing methods, while showing similar functional precision and temporal effects in the subsequent analysis compared to a standard 10-fold accelerated acquisition.
Improved Simultaneous Multi-Slice Functional MRI Using Self-supervised Deep Learning
May 10, 2021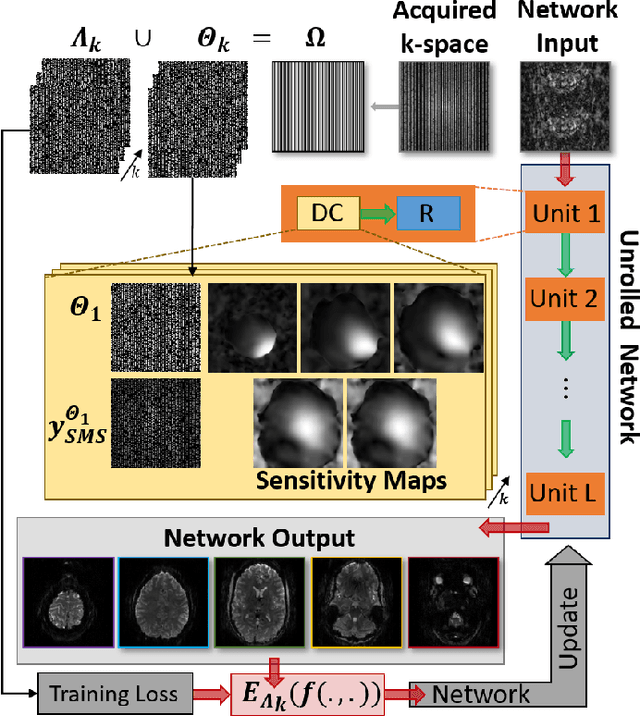
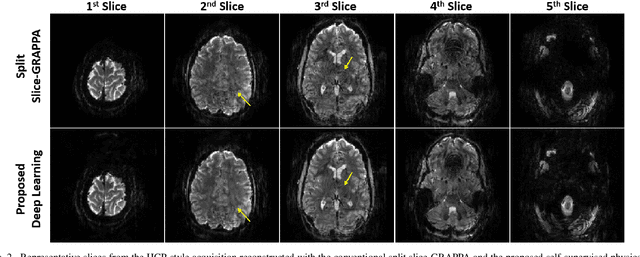
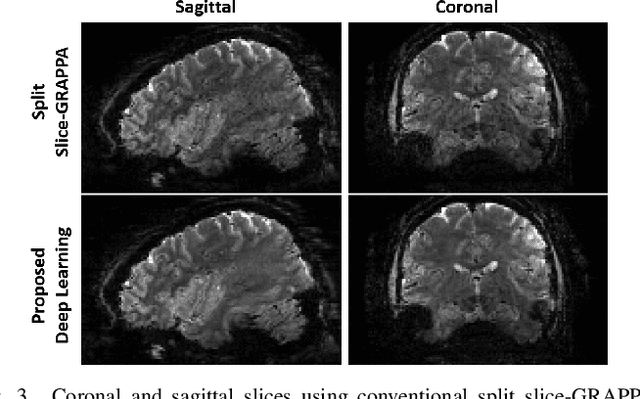
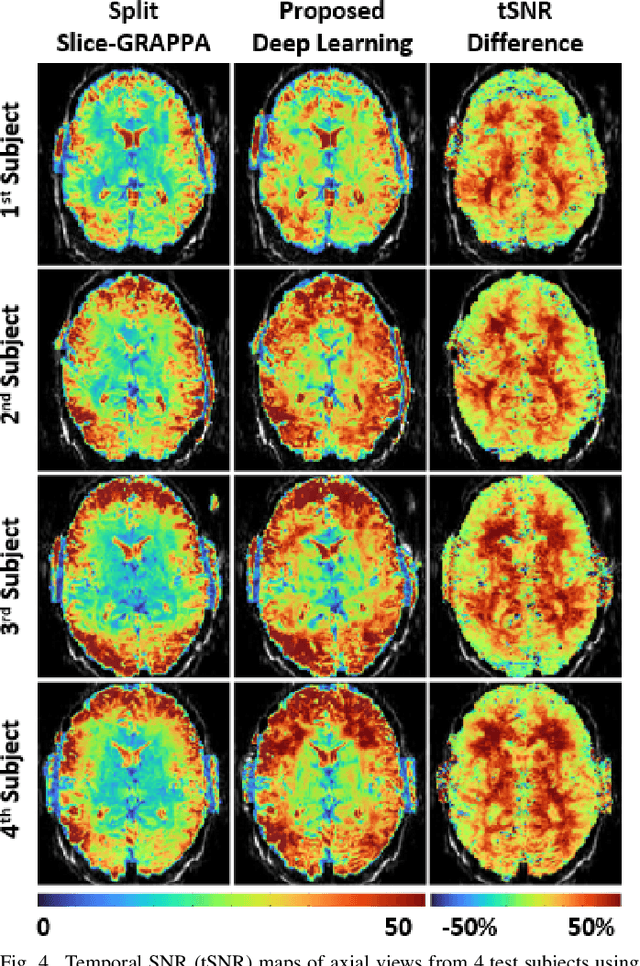
Abstract:Functional MRI (fMRI) is commonly used for interpreting neural activities across the brain. Numerous accelerated fMRI techniques aim to provide improved spatiotemporal resolutions. Among these, simultaneous multi-slice (SMS) imaging has emerged as a powerful strategy, becoming a part of large-scale studies, such as the Human Connectome Project. However, when SMS imaging is combined with in-plane acceleration for higher acceleration rates, conventional SMS reconstruction methods may suffer from noise amplification and other artifacts. Recently, deep learning (DL) techniques have gained interest for improving MRI reconstruction. However, these methods are typically trained in a supervised manner that necessitates fully-sampled reference data, which is not feasible in highly-accelerated fMRI acquisitions. Self-supervised learning that does not require fully-sampled data has recently been proposed and has shown similar performance to supervised learning. However, it has only been applied for in-plane acceleration. Furthermore the effect of DL reconstruction on subsequent fMRI analysis remains unclear. In this work, we extend self-supervised DL reconstruction to SMS imaging. Our results on prospectively 10-fold accelerated 7T fMRI data show that self-supervised DL reduces reconstruction noise and suppresses residual artifacts. Subsequent fMRI analysis remains unaltered by DL processing, while the improved temporal signal-to-noise ratio produces higher coherence estimates between task runs.
On Instabilities of Conventional Multi-Coil MRI Reconstruction to Small Adverserial Perturbations
Feb 25, 2021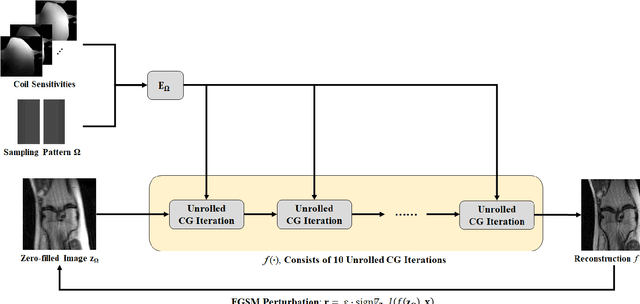
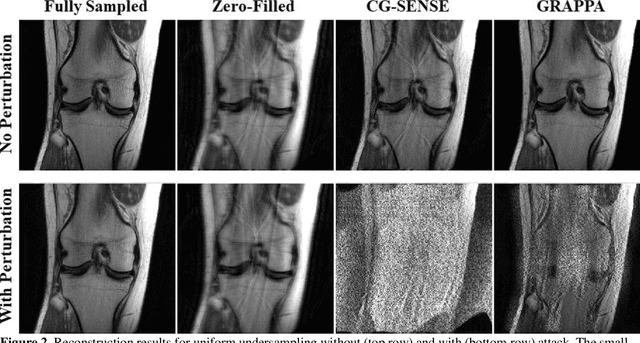
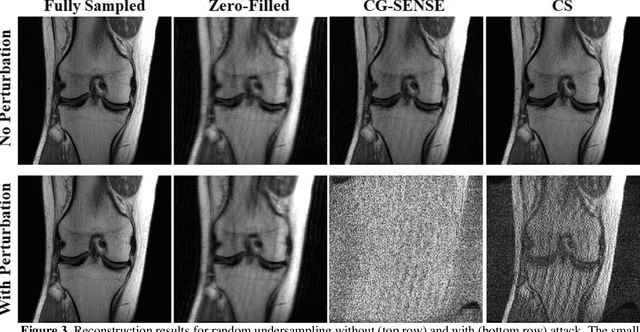
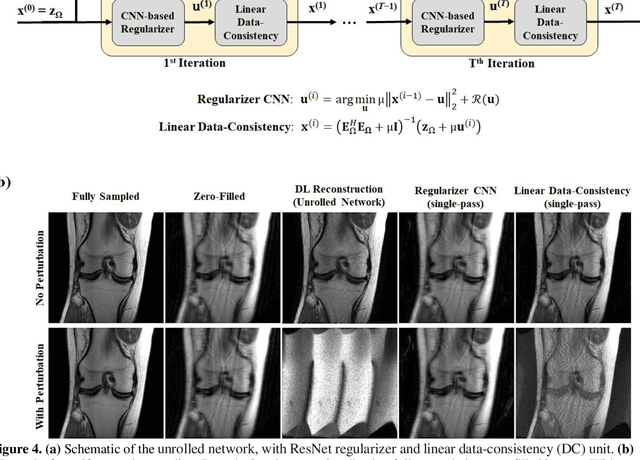
Abstract:Although deep learning (DL) has received much attention in accelerated MRI, recent studies suggest small perturbations may lead to instabilities in DL-based reconstructions, leading to concern for their clinical application. However, these works focus on single-coil acquisitions, which is not practical. We investigate instabilities caused by small adversarial attacks for multi-coil acquisitions. Our results suggest that, parallel imaging and multi-coil CS exhibit considerable instabilities against small adversarial perturbations.
Self-Supervised Physics-Guided Deep Learning Reconstruction For High-Resolution 3D LGE CMR
Nov 18, 2020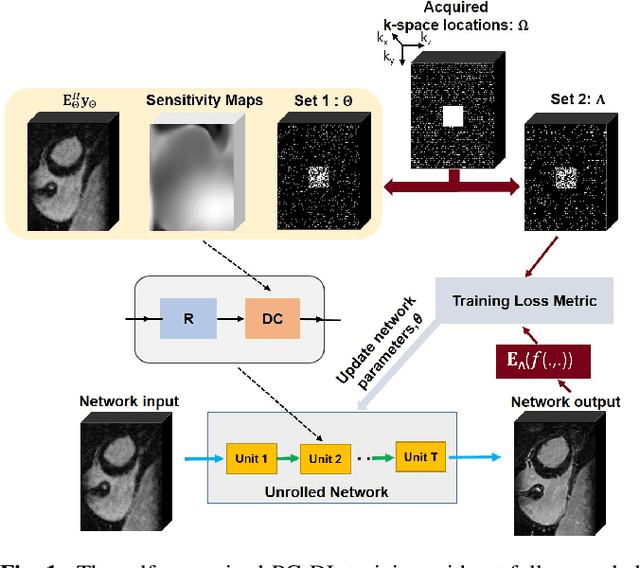
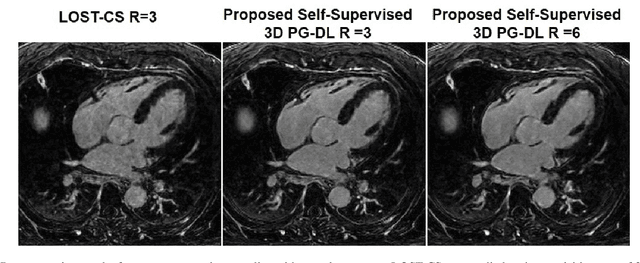
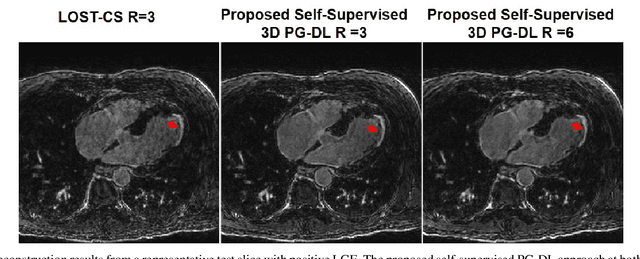
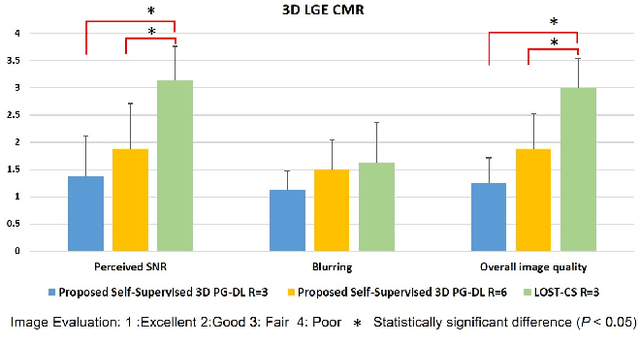
Abstract:Late gadolinium enhancement (LGE) cardiac MRI (CMR) is the clinical standard for diagnosis of myocardial scar. 3D isotropic LGE CMR provides improved coverage and resolution compared to 2D imaging. However, image acceleration is required due to long scan times and contrast washout. Physics-guided deep learning (PG-DL) approaches have recently emerged as an improved accelerated MRI strategy. Training of PG-DL methods is typically performed in supervised manner requiring fully-sampled data as reference, which is challenging in 3D LGE CMR. Recently, a self-supervised learning approach was proposed to enable training PG-DL techniques without fully-sampled data. In this work, we extend this self-supervised learning approach to 3D imaging, while tackling challenges related to small training database sizes of 3D volumes. Results and a reader study on prospectively accelerated 3D LGE show that the proposed approach at 6-fold acceleration outperforms the clinically utilized compressed sensing approach at 3-fold acceleration.
Improved Supervised Training of Physics-Guided Deep Learning Image Reconstruction with Multi-Masking
Oct 26, 2020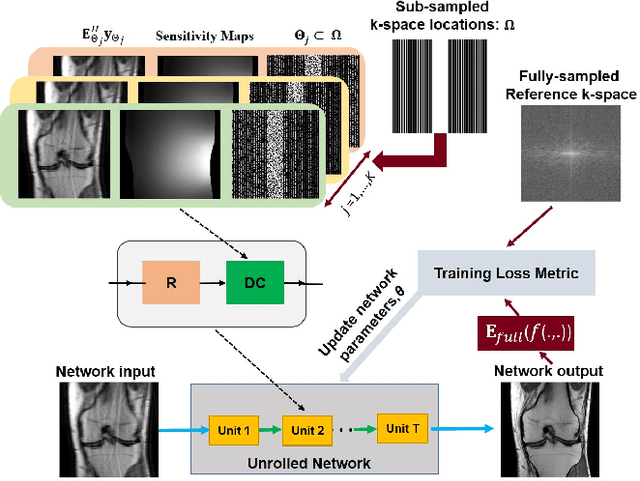
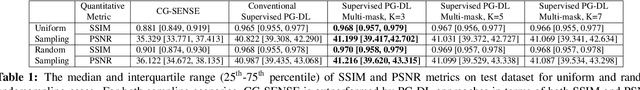
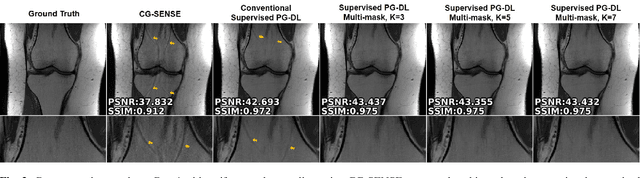
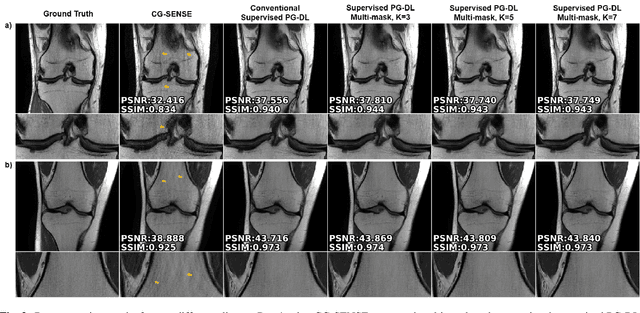
Abstract:Physics-guided deep learning (PG-DL) via algorithm unrolling has received significant interest for improved image reconstruction, including MRI applications. These methods unroll an iterative optimization algorithm into a series of regularizer and data consistency units. The unrolled networks are typically trained end-to-end using a supervised approach. Current supervised PG-DL approaches use all of the available sub-sampled measurements in their data consistency units. Thus, the network learns to fit the rest of the measurements. In this study, we propose to improve the performance and robustness of supervised training by utilizing randomness by retrospectively selecting only a subset of all the available measurements for data consistency units. The process is repeated multiple times using different random masks during training for further enhancement. Results on knee MRI show that the proposed multi-mask supervised PG-DL enhances reconstruction performance compared to conventional supervised PG-DL approaches.
Multi-Mask Self-Supervised Learning for Physics-Guided Neural Networks in Highly Accelerated MRI
Aug 13, 2020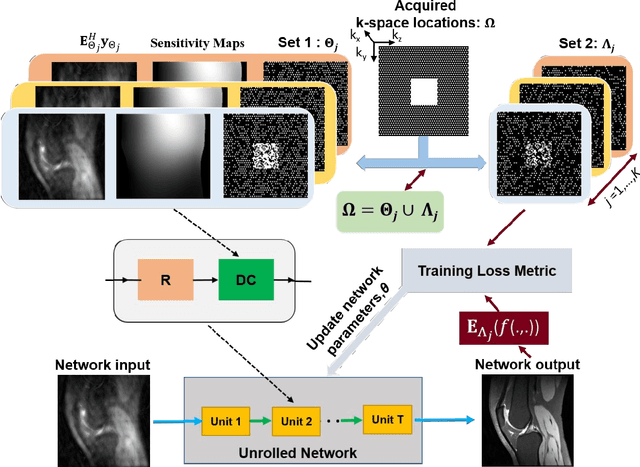
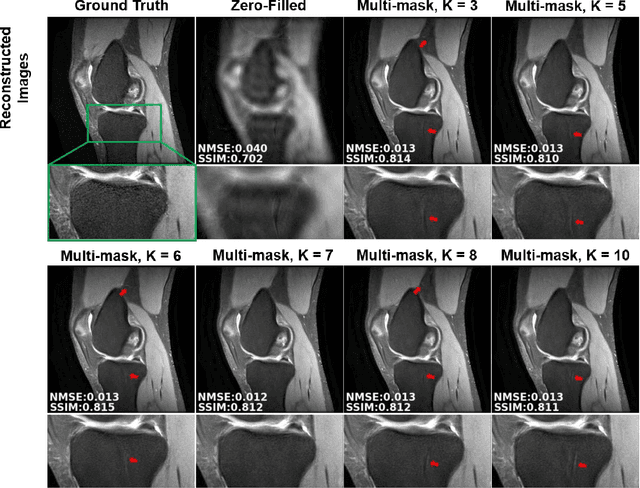
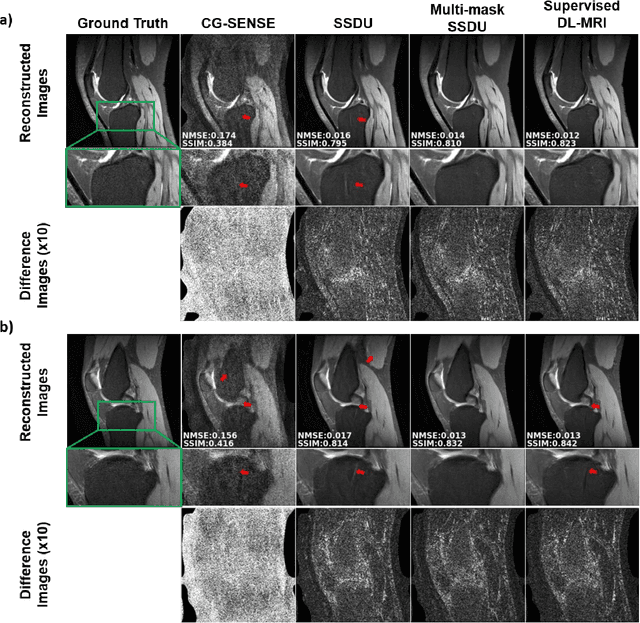
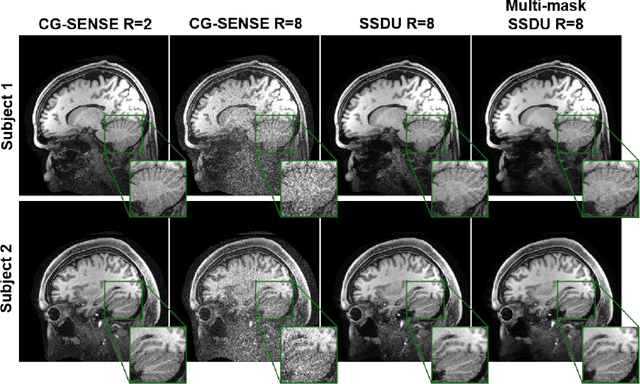
Abstract:Purpose: To develop an improved self-supervised learning strategy that efficiently uses the acquired data for training a physics-guided reconstruction network without a database of fully-sampled data. Methods: Currently self-supervised learning for physics-guided reconstruction networks splits acquired undersampled data into two disjoint sets, where one is used for data consistency (DC) in the unrolled network and the other to define the training loss. The proposed multi-mask self-supervised learning via data undersampling (SSDU) splits acquired measurements into multiple pairs of disjoint sets for each training sample, while using one of these sets for DC units and the other for defining loss, thereby more efficiently using the undersampled data. Multi-mask SSDU is applied on fully-sampled 3D knee and prospectively undersampled 3D brain MRI datasets, which are retrospectively subsampled to acceleration rate (R)=8, and compared to CG-SENSE and single-mask SSDU DL-MRI, as well as supervised DL-MRI when fully-sampled data is available. Results: Results on knee MRI show that the proposed multi-mask SSDU outperforms SSDU and performs closely with supervised DL-MRI, while significantly outperforming CG-SENSE. A clinical reader study further ranks the multi-mask SSDU higher than supervised DL-MRI in terms of SNR and aliasing artifacts. Results on brain MRI show that multi-mask SSDU achieves better reconstruction quality compared to SSDU and CG-SENSE. Reader study demonstrates that multi-mask SSDU at R=8 significantly improves reconstruction compared to single-mask SSDU at R=8, as well as CG-SENSE at R=2. Conclusion: The proposed multi-mask SSDU approach enables improved training of physics-guided neural networks without fully-sampled data, by enabling efficient use of the undersampled data with multiple masks.
High-Fidelity Accelerated MRI Reconstruction by Scan-Specific Fine-Tuning of Physics-Based Neural Networks
May 12, 2020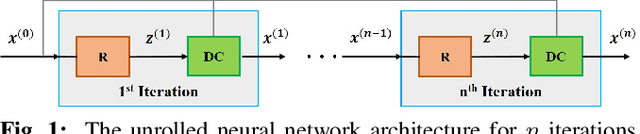
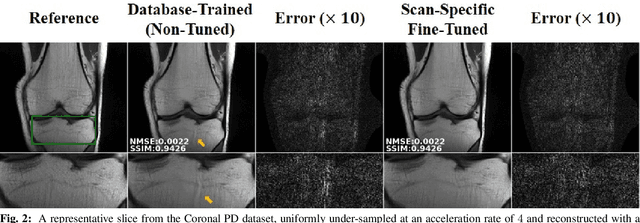
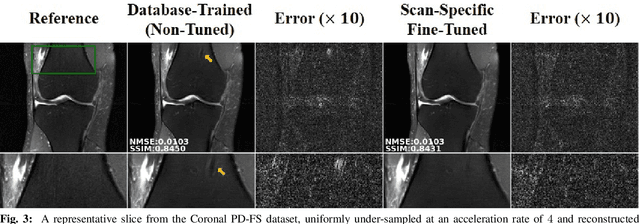
Abstract:Long scan duration remains a challenge for high-resolution MRI. Deep learning has emerged as a powerful means for accelerated MRI reconstruction by providing data-driven regularizers that are directly learned from data. These data-driven priors typically remain unchanged for future data in the testing phase once they are learned during training. In this study, we propose to use a transfer learning approach to fine-tune these regularizers for new subjects using a self-supervision approach. While the proposed approach can compromise the extremely fast reconstruction time of deep learning MRI methods, our results on knee MRI indicate that such adaptation can substantially reduce the remaining artifacts in reconstructed images. In addition, the proposed approach has the potential to reduce the risks of generalization to rare pathological conditions, which may be unavailable in the training data.
Self-Supervised Learning of Physics-Based Reconstruction Neural Networks without Fully-Sampled Reference Data
Dec 16, 2019
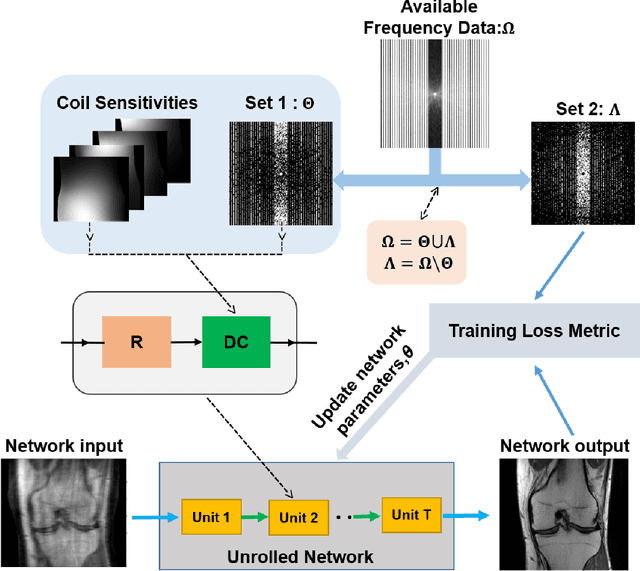
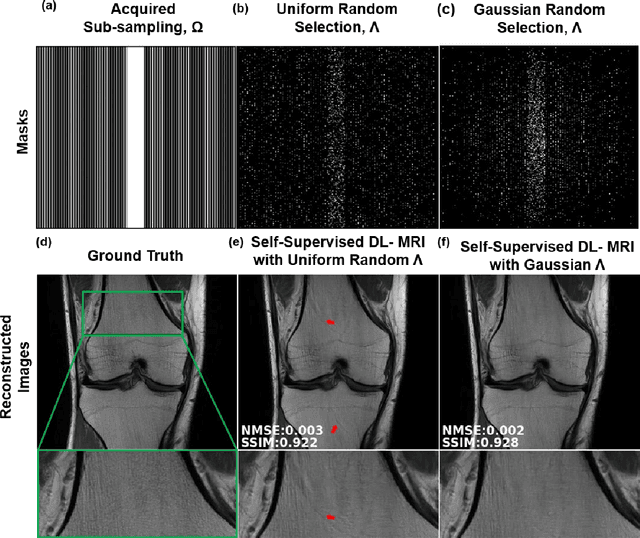
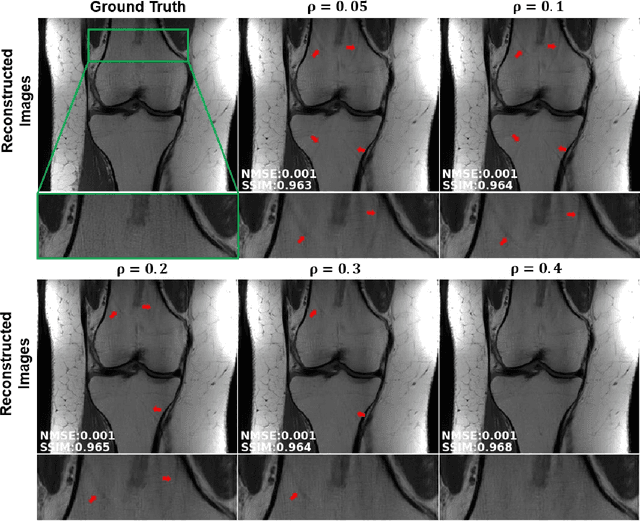
Abstract:Purpose: To develop a strategy for training a physics-driven MRI reconstruction neural network without a database of fully-sampled datasets. Theory and Methods: Self-supervised learning via data under-sampling (SSDU) for physics-based deep learning (DL) reconstruction partitions available measurements into two sets, one of which is used in the data consistency units in the unrolled network and the other is used to define the loss for training. The proposed training without fully-sampled data is compared to fully-supervised training with ground-truth data, as well as conventional compressed sensing and parallel imaging methods using the publicly available fastMRI knee database. The same physics-based neural network is used for both proposed SSDU and supervised training. The SSDU training is also applied to prospectively 2-fold accelerated high-resolution brain datasets at different acceleration rates, and compared to parallel imaging. Results: Results on five different knee sequences at acceleration rate of 4 shows that proposed self-supervised approach performs closely with supervised learning, while significantly outperforming conventional compressed sensing and parallel imaging, as characterized by quantitative metrics and a clinical reader study. The results on prospectively sub-sampled brain datasets, where supervised learning cannot be employed due to lack of ground-truth reference, show that the proposed self-supervised approach successfully perform reconstruction at high acceleration rates (4, 6 and 8). Image readings indicate improved visual reconstruction quality with the proposed approach compared to parallel imaging at acquisition acceleration. Conclusion: The proposed SSDU approach allows training of physics-based DL-MRI reconstruction without fully-sampled data, while achieving comparable results with supervised DL-MRI trained on fully-sampled data.
Dense Recurrent Neural Networks for Inverse Problems: History-Cognizant Unrolling of Optimization Algorithms
Dec 16, 2019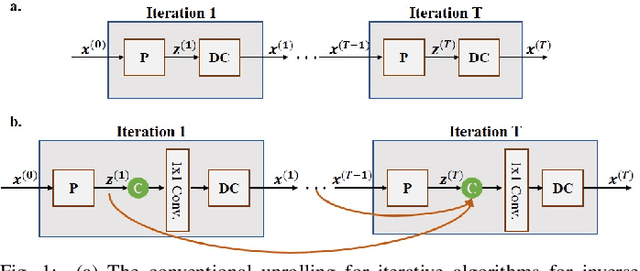
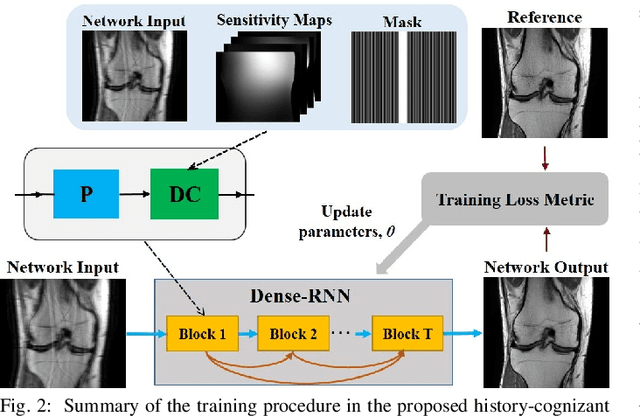
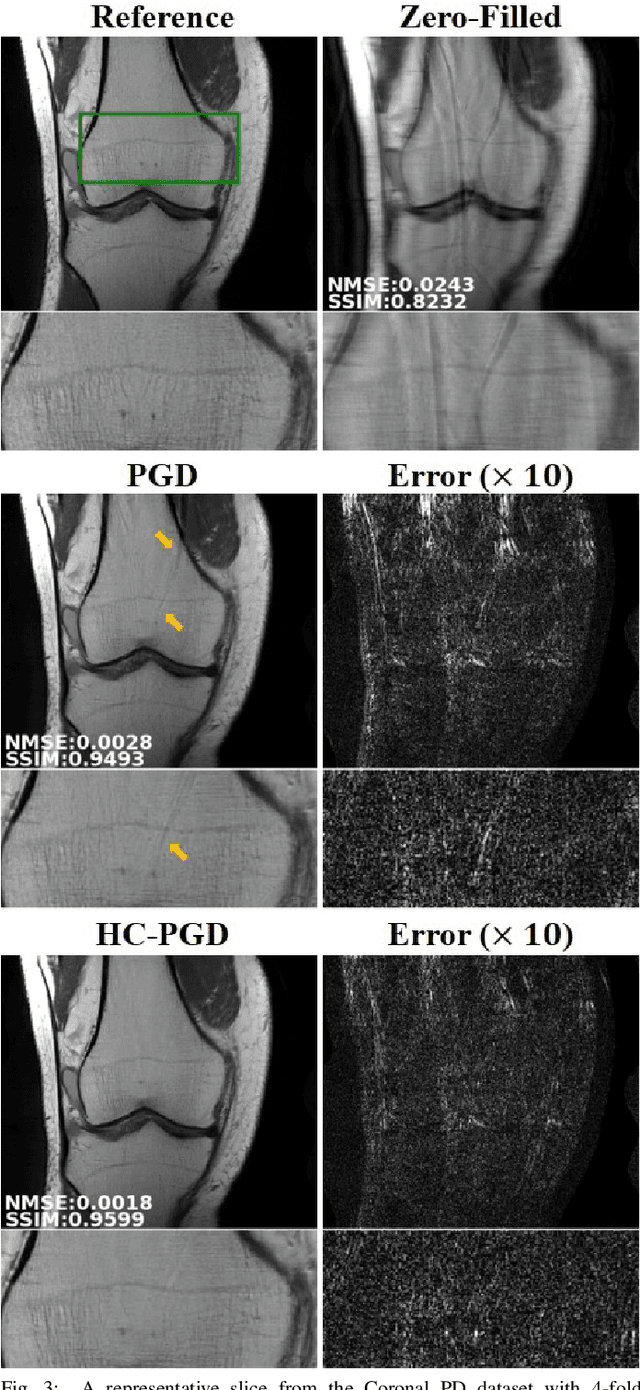
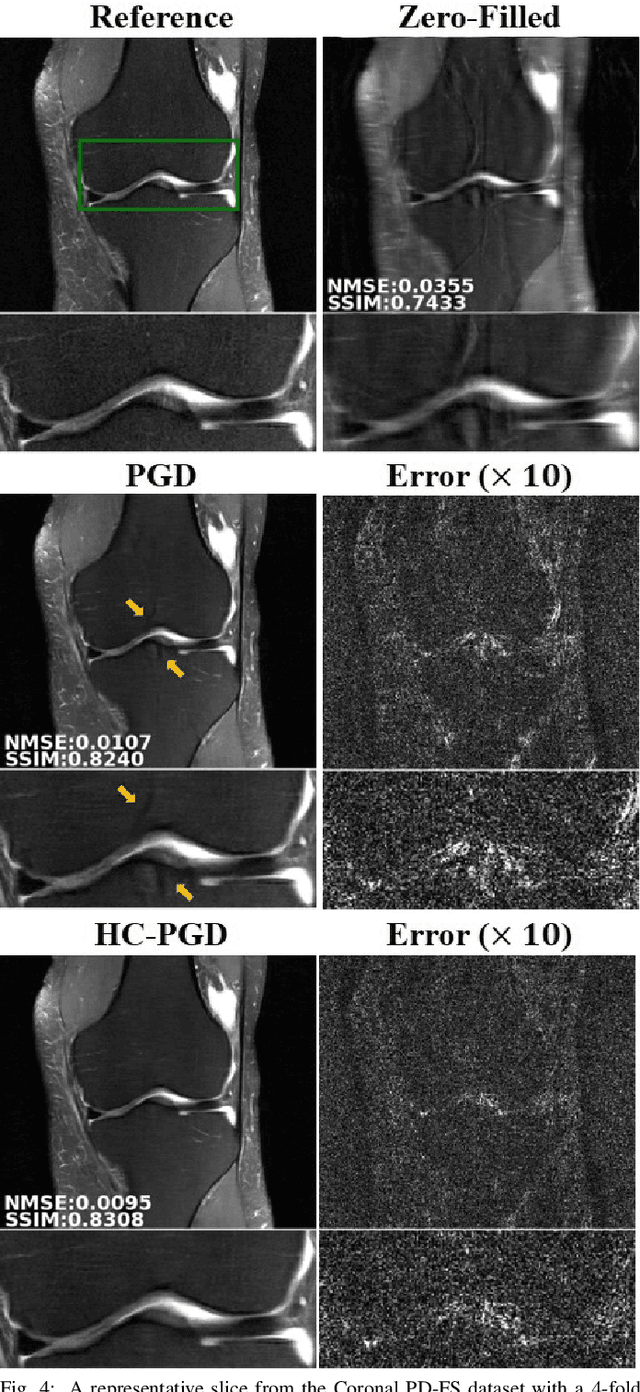
Abstract:Inverse problems in medical imaging applications incorporate domain-specific knowledge about the forward encoding operator in a regularized reconstruction framework. Recently physics-driven deep learning (DL) methods have been proposed to use neural networks for data-driven regularization. These methods unroll iterative optimization algorithms to solve the inverse problem objective function, by alternating between domain-specific data consistency and data-driven regularization via neural networks. The whole unrolled network is then trained end-to-end to learn the parameters of the network. Due to simplicity of data consistency updates with gradient descent steps, proximal gradient descent (PGD) is a common approach to unroll physics-driven DL reconstruction methods. However, PGD methods have slow convergence rates, necessitating a higher number of unrolled iterations, leading to memory issues in training and slower reconstruction times in testing. Inspired by efficient variants of PGD methods that use a history of the previous iterates, we propose a history-cognizant unrolling of the optimization algorithm with dense connections across iterations for improved performance. In our approach, the gradient descent steps are calculated at a trainable combination of the outputs of all the previous regularization units. We also apply this idea to unrolling variable splitting methods with quadratic relaxation. Our results in reconstruction of the fastMRI knee dataset show that the proposed history-cognizant approach reduces residual aliasing artifacts compared to its conventional unrolled counterpart without requiring extra computational power or increasing reconstruction time.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge