Ziyue Xu
Accounting for Dependencies in Deep Learning Based Multiple Instance Learning for Whole Slide Imaging
Nov 01, 2021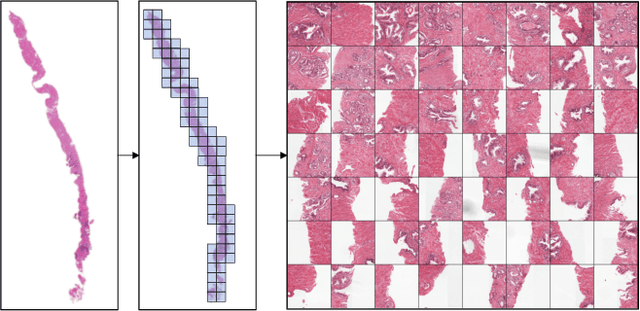
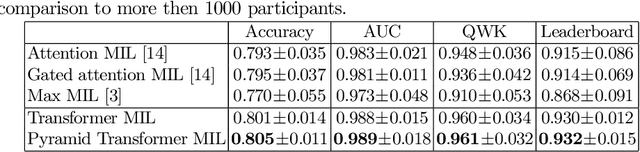
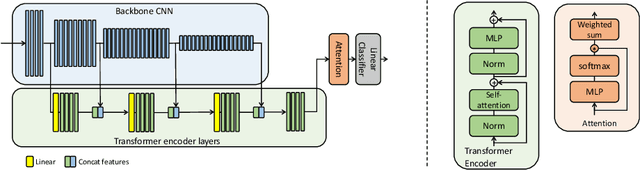
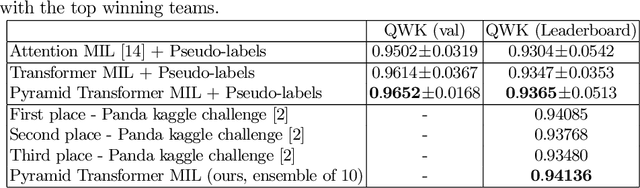
Abstract:Multiple instance learning (MIL) is a key algorithm for classification of whole slide images (WSI). Histology WSIs can have billions of pixels, which create enormous computational and annotation challenges. Typically, such images are divided into a set of patches (a bag of instances), where only bag-level class labels are provided. Deep learning based MIL methods calculate instance features using convolutional neural network (CNN). Our proposed approach is also deep learning based, with the following two contributions: Firstly, we propose to explicitly account for dependencies between instances during training by embedding self-attention Transformer blocks to capture dependencies between instances. For example, a tumor grade may depend on the presence of several particular patterns at different locations in WSI, which requires to account for dependencies between patches. Secondly, we propose an instance-wise loss function based on instance pseudo-labels. We compare the proposed algorithm to multiple baseline methods, evaluate it on the PANDA challenge dataset, the largest publicly available WSI dataset with over 11K images, and demonstrate state-of-the-art results.
Improving Pneumonia Localization via Cross-Attention on Medical Images and Reports
Oct 06, 2021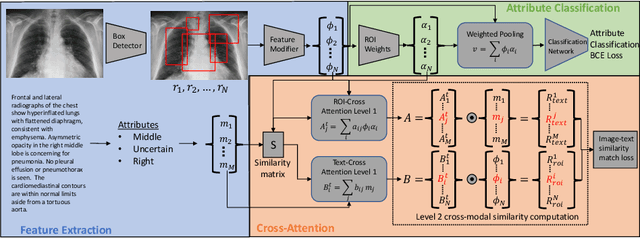
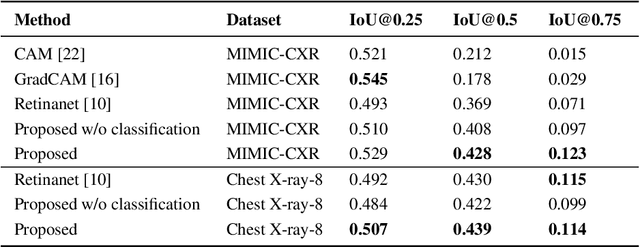
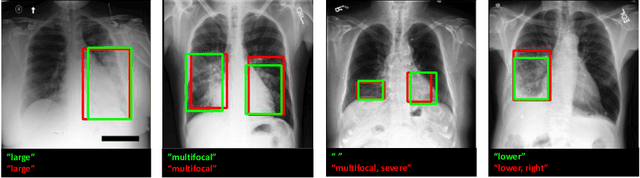
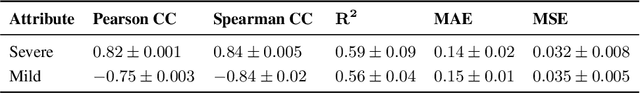
Abstract:Localization and characterization of diseases like pneumonia are primary steps in a clinical pipeline, facilitating detailed clinical diagnosis and subsequent treatment planning. Additionally, such location annotated datasets can provide a pathway for deep learning models to be used for downstream tasks. However, acquiring quality annotations is expensive on human resources and usually requires domain expertise. On the other hand, medical reports contain a plethora of information both about pneumonia characteristics and its location. In this paper, we propose a novel weakly-supervised attention-driven deep learning model that leverages encoded information in medical reports during training to facilitate better localization. Our model also performs classification of attributes that are associated to pneumonia and extracted from medical reports for supervision. Both the classification and localization are trained in conjunction and once trained, the model can be utilized for both the localization and characterization of pneumonia using only the input image. In this paper, we explore and analyze the model using chest X-ray datasets and demonstrate qualitatively and quantitatively that the introduction of textual information improves pneumonia localization. We showcase quantitative results on two datasets, MIMIC-CXR and Chest X-ray-8, and we also showcase severity characterization on the COVID-19 dataset.
Federated Whole Prostate Segmentation in MRI with Personalized Neural Architectures
Jul 16, 2021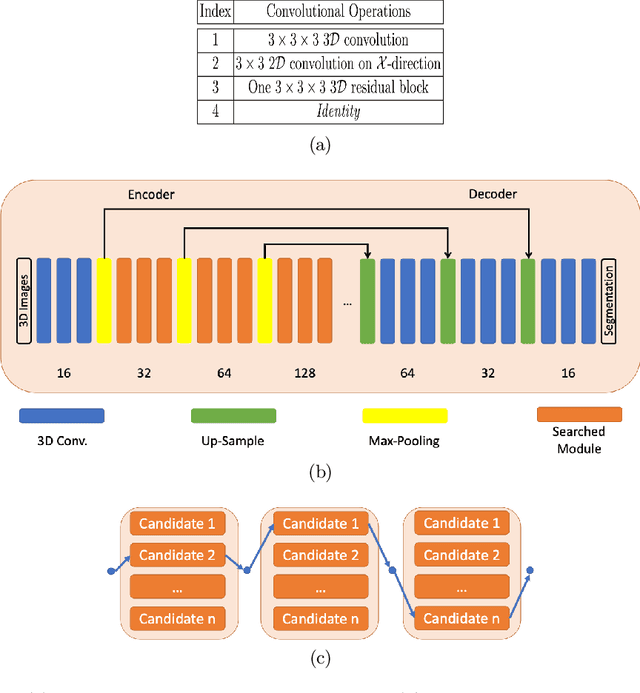
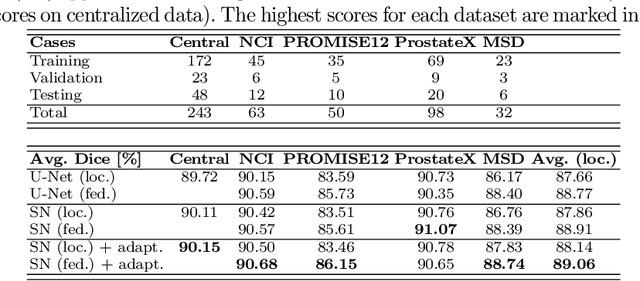
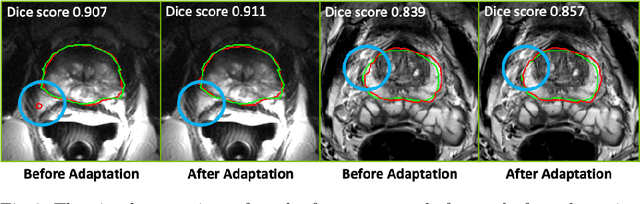
Abstract:Building robust deep learning-based models requires diverse training data, ideally from several sources. However, these datasets cannot be combined easily because of patient privacy concerns or regulatory hurdles, especially if medical data is involved. Federated learning (FL) is a way to train machine learning models without the need for centralized datasets. Each FL client trains on their local data while only sharing model parameters with a global server that aggregates the parameters from all clients. At the same time, each client's data can exhibit differences and inconsistencies due to the local variation in the patient population, imaging equipment, and acquisition protocols. Hence, the federated learned models should be able to adapt to the local particularities of a client's data. In this work, we combine FL with an AutoML technique based on local neural architecture search by training a "supernet". Furthermore, we propose an adaptation scheme to allow for personalized model architectures at each FL client's site. The proposed method is evaluated on four different datasets from 3D prostate MRI and shown to improve the local models' performance after adaptation through selecting an optimal path through the AutoML supernet.
FedGL: Federated Graph Learning Framework with Global Self-Supervision
May 07, 2021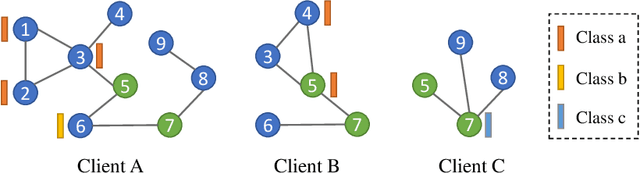
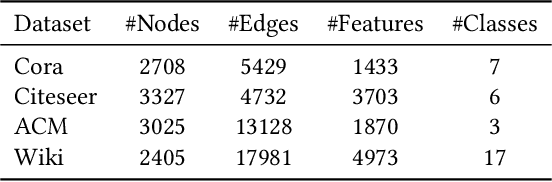
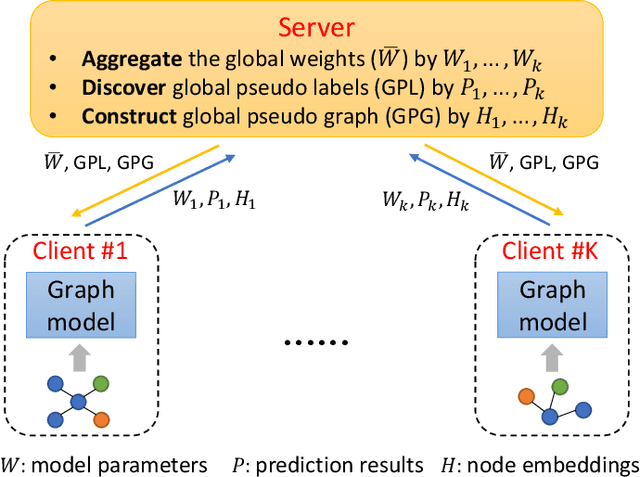
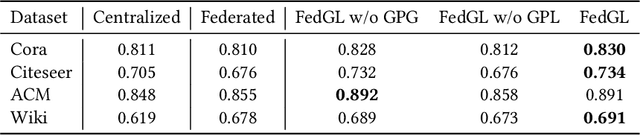
Abstract:Graph data are ubiquitous in the real world. Graph learning (GL) tries to mine and analyze graph data so that valuable information can be discovered. Existing GL methods are designed for centralized scenarios. However, in practical scenarios, graph data are usually distributed in different organizations, i.e., the curse of isolated data islands. To address this problem, we incorporate federated learning into GL and propose a general Federated Graph Learning framework FedGL, which is capable of obtaining a high-quality global graph model while protecting data privacy by discovering the global self-supervision information during the federated training. Concretely, we propose to upload the prediction results and node embeddings to the server for discovering the global pseudo label and global pseudo graph, which are distributed to each client to enrich the training labels and complement the graph structure respectively, thereby improving the quality of each local model. Moreover, the global self-supervision enables the information of each client to flow and share in a privacy-preserving manner, thus alleviating the heterogeneity and utilizing the complementarity of graph data among different clients. Finally, experimental results show that FedGL significantly outperforms baselines on four widely used graph datasets.
Information Bottleneck Attribution for Visual Explanations of Diagnosis and Prognosis
Apr 07, 2021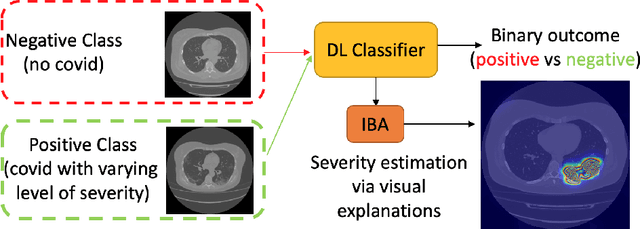
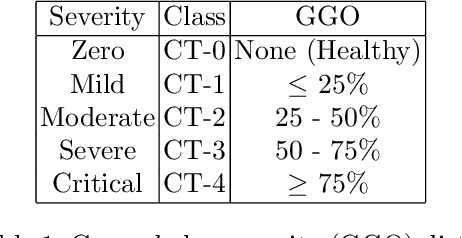
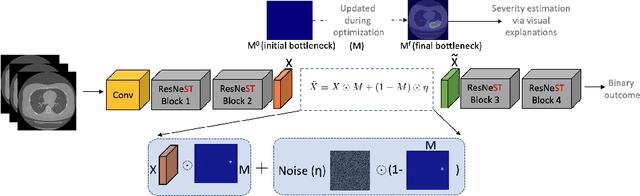
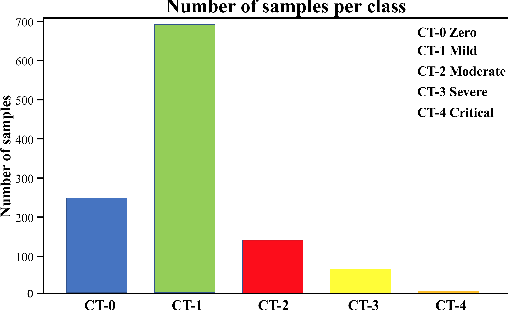
Abstract:Visual explanation methods have an important role in the prognosis of the patients where the annotated data is limited or not available. There have been several attempts to use gradient-based attribution methods to localize pathology from medical scans without using segmentation labels. This research direction has been impeded by the lack of robustness and reliability. These methods are highly sensitive to the network parameters. In this study, we introduce a robust visual explanation method to address this problem for medical applications. We provide a highly innovative algorithm to quantifying lesions in the lungs caused by the Covid-19 with high accuracy and robustness without using dense segmentation labels. Inspired by the information bottleneck concept, we mask the neural network representation with noise to find out important regions. This approach overcomes the drawbacks of commonly used Grad-Cam and its derived algorithms. The premise behind our proposed strategy is that the information flow is minimized while ensuring the classifier prediction stays similar. Our findings indicate that the bottleneck condition provides a more stable and robust severity estimation than the similar attribution methods.
Self-supervised Image-text Pre-training With Mixed Data In Chest X-rays
Mar 30, 2021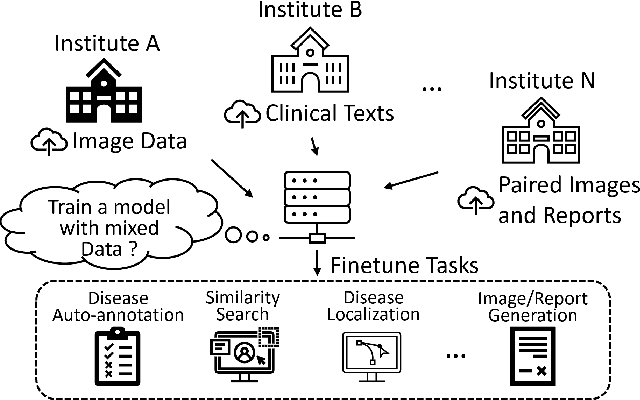
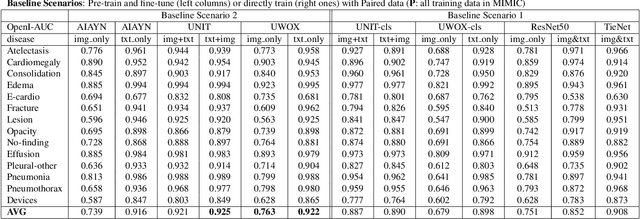
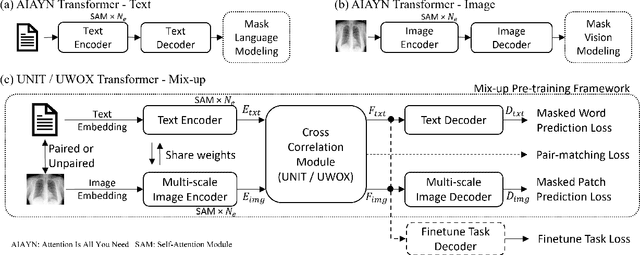
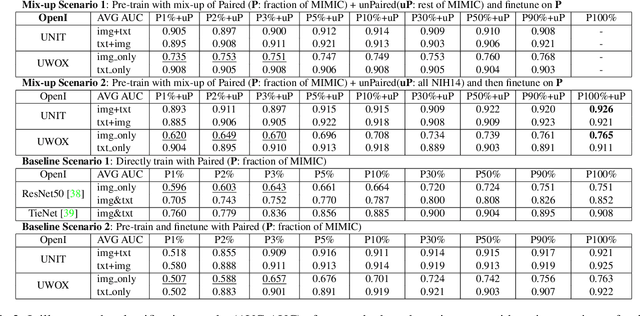
Abstract:Pre-trained models, e.g., from ImageNet, have proven to be effective in boosting the performance of many downstream applications. It is too demanding to acquire large-scale annotations to build such models for medical imaging. Meanwhile, there are numerous clinical data (in the form of images and text reports) stored in the hospital information systems. The paired image-text data from the same patient study could be utilized for the pre-training task in a weakly supervised manner. However, the integrity, accessibility, and amount of such raw data vary across different institutes, e.g., paired vs. unpaired (image-only or text-only). In this work, we introduce an image-text pre-training framework that can learn from these raw data with mixed data inputs, i.e., paired image-text data, a mixture of paired and unpaired data. The unpaired data can be sourced from one or multiple institutes (e.g., images from one institute coupled with texts from another). Specifically, we propose a transformer-based training framework for jointly learning the representation of both the image and text data. In addition to the existing masked language modeling, multi-scale masked vision modeling is introduced as a self-supervised training task for image patch regeneration. We not only demonstrate the feasibility of pre-training across mixed data inputs but also illustrate the benefits of adopting such pre-trained models in 3 chest X-ray applications, i.e., classification, retrieval, and image regeneration. Superior results are reported in comparison to prior art using MIMIC-CXR, NIH14-CXR, and OpenI-CXR datasets.
Federated Semi-Supervised Learning for COVID Region Segmentation in Chest CT using Multi-National Data from China, Italy, Japan
Nov 23, 2020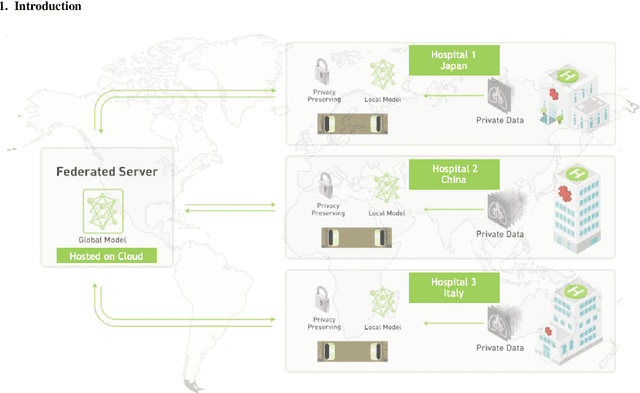
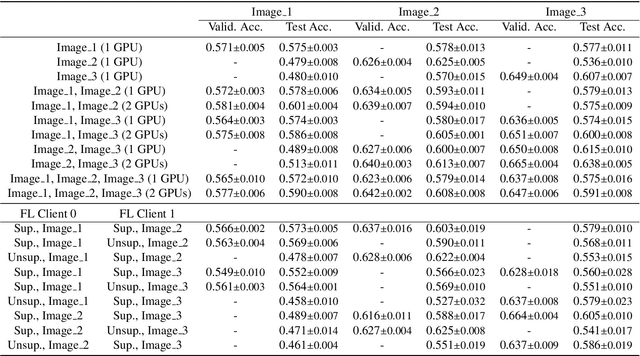
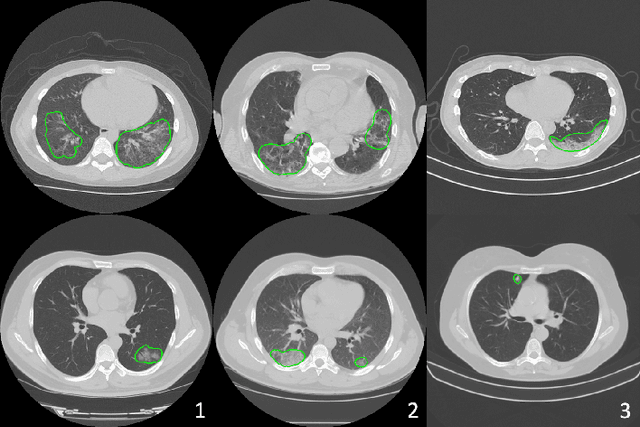
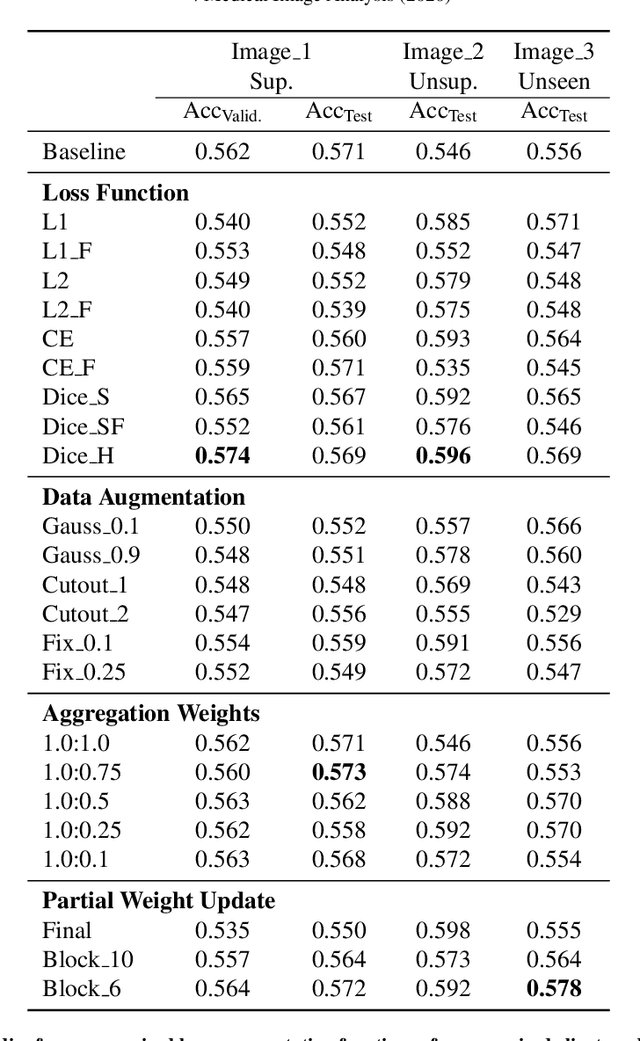
Abstract:The recent outbreak of COVID-19 has led to urgent needs for reliable diagnosis and management of SARS-CoV-2 infection. As a complimentary tool, chest CT has been shown to be able to reveal visual patterns characteristic for COVID-19, which has definite value at several stages during the disease course. To facilitate CT analysis, recent efforts have focused on computer-aided characterization and diagnosis, which has shown promising results. However, domain shift of data across clinical data centers poses a serious challenge when deploying learning-based models. In this work, we attempt to find a solution for this challenge via federated and semi-supervised learning. A multi-national database consisting of 1704 scans from three countries is adopted to study the performance gap, when training a model with one dataset and applying it to another. Expert radiologists manually delineated 945 scans for COVID-19 findings. In handling the variability in both the data and annotations, a novel federated semi-supervised learning technique is proposed to fully utilize all available data (with or without annotations). Federated learning avoids the need for sensitive data-sharing, which makes it favorable for institutions and nations with strict regulatory policy on data privacy. Moreover, semi-supervision potentially reduces the annotation burden under a distributed setting. The proposed framework is shown to be effective compared to fully supervised scenarios with conventional data sharing instead of model weight sharing.
Learning Image Labels On-the-fly for Training Robust Classification Models
Oct 02, 2020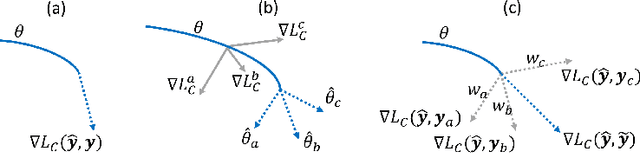
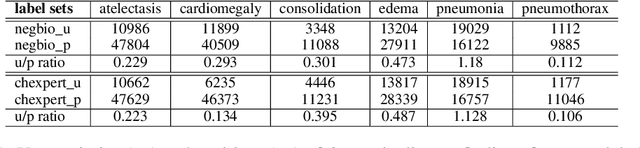
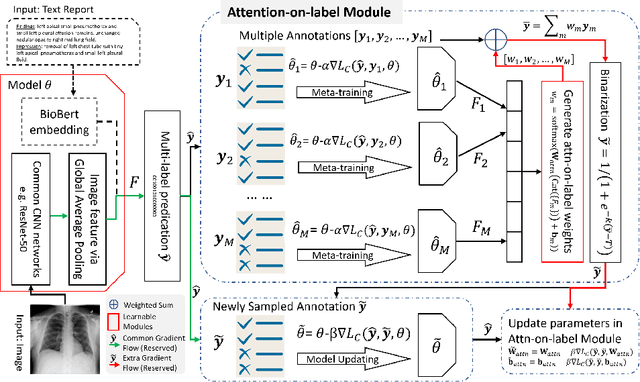
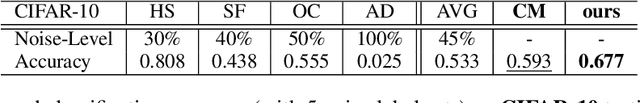
Abstract:Current deep learning paradigms largely benefit from the tremendous amount of annotated data. However, the quality of the annotations often varies among labelers. Multi-observer studies have been conducted to study these annotation variances (by labeling the same data for multiple times) and its effects on critical applications like medical image analysis. This process indeed adds an extra burden to the already tedious annotation work that usually requires professional training and expertise in the specific domains. On the other hand, automated annotation methods based on NLP algorithms have recently shown promise as a reasonable alternative, relying on the existing diagnostic reports of those images that are widely available in the clinical system. Compared to human labelers, different algorithms provide labels with varying qualities that are even noisier. In this paper, we show how noisy annotations (e.g., from different algorithm-based labelers) can be utilized together and mutually benefit the learning of classification tasks. Specifically, the concept of attention-on-label is introduced to sample better label sets on-the-fly as the training data. A meta-training based label-sampling module is designed to attend the labels that benefit the model learning the most through additional back-propagation processes. We apply the attention-on-label scheme on the classification task of a synthetic noisy CIFAR-10 dataset to prove the concept, and then demonstrate superior results (3-5% increase on average in multiple disease classification AUCs) on the chest x-ray images from a hospital-scale dataset (MIMIC-CXR) and hand-labeled dataset (OpenI) in comparison to regular training paradigms.
Going to Extremes: Weakly Supervised Medical Image Segmentation
Sep 25, 2020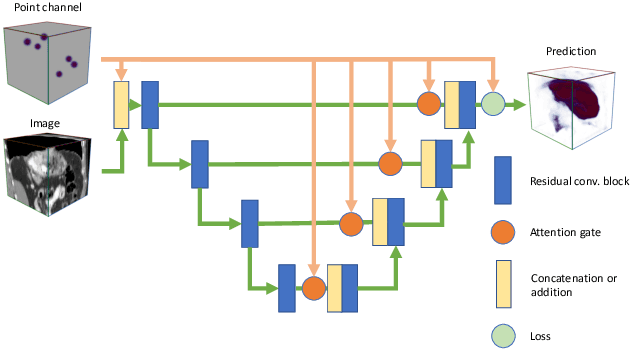
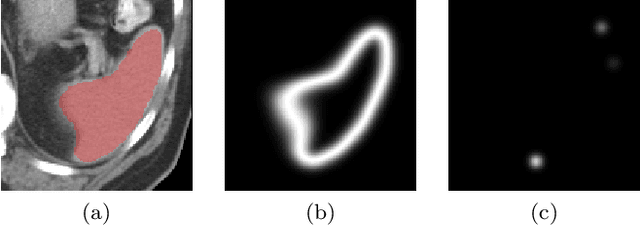
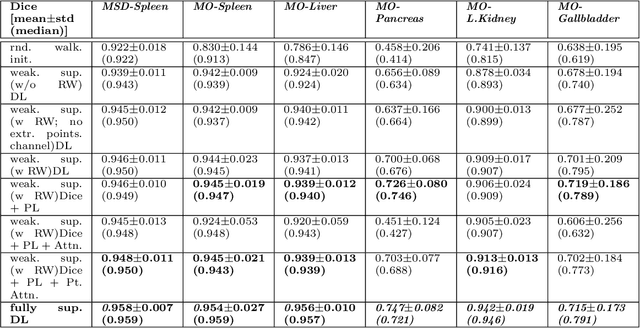
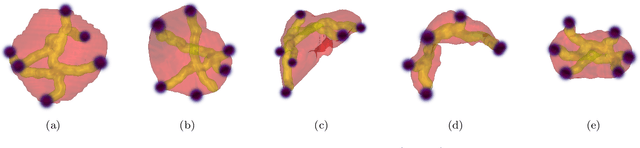

Abstract:Medical image annotation is a major hurdle for developing precise and robust machine learning models. Annotation is expensive, time-consuming, and often requires expert knowledge, particularly in the medical field. Here, we suggest using minimal user interaction in the form of extreme point clicks to train a segmentation model which, in effect, can be used to speed up medical image annotation. An initial segmentation is generated based on the extreme points utilizing the random walker algorithm. This initial segmentation is then used as a noisy supervision signal to train a fully convolutional network that can segment the organ of interest, based on the provided user clicks. Through experimentation on several medical imaging datasets, we show that the predictions of the network can be refined using several rounds of training with the prediction from the same weakly annotated data. Further improvements are shown utilizing the clicked points within a custom-designed loss and attention mechanism. Our approach has the potential to speed up the process of generating new training datasets for the development of new machine learning and deep learning-based models for, but not exclusively, medical image analysis.
ScribbleBox: Interactive Annotation Framework for Video Object Segmentation
Aug 22, 2020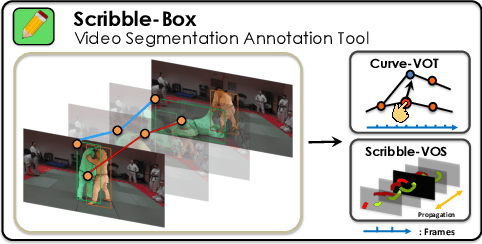
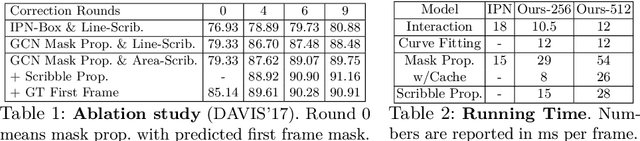
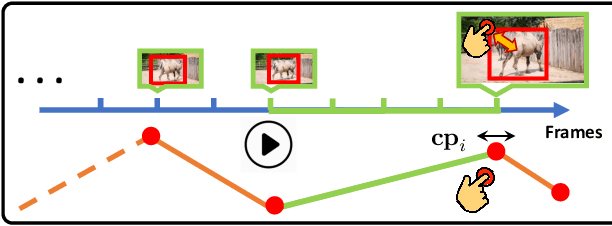
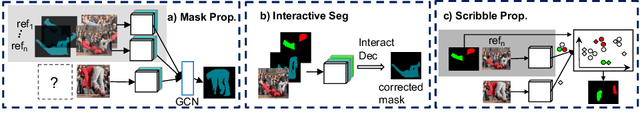
Abstract:Manually labeling video datasets for segmentation tasks is extremely time consuming. In this paper, we introduce ScribbleBox, a novel interactive framework for annotating object instances with masks in videos. In particular, we split annotation into two steps: annotating objects with tracked boxes, and labeling masks inside these tracks. We introduce automation and interaction in both steps. Box tracks are annotated efficiently by approximating the trajectory using a parametric curve with a small number of control points which the annotator can interactively correct. Our approach tolerates a modest amount of noise in the box placements, thus typically only a few clicks are needed to annotate tracked boxes to a sufficient accuracy. Segmentation masks are corrected via scribbles which are efficiently propagated through time. We show significant performance gains in annotation efficiency over past work. We show that our ScribbleBox approach reaches 88.92% J&F on DAVIS2017 with 9.14 clicks per box track, and 4 frames of scribble annotation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge