Youngser Park
An Analysis of Euclidean vs. Graph-Based Framing for Bilingual Lexicon Induction from Word Embedding Spaces
Sep 26, 2021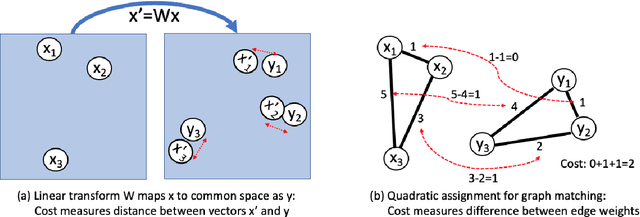

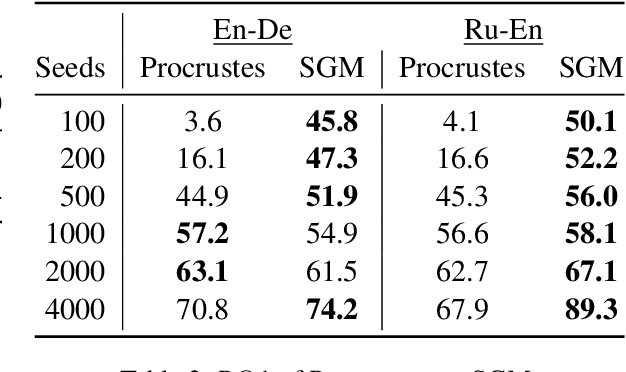
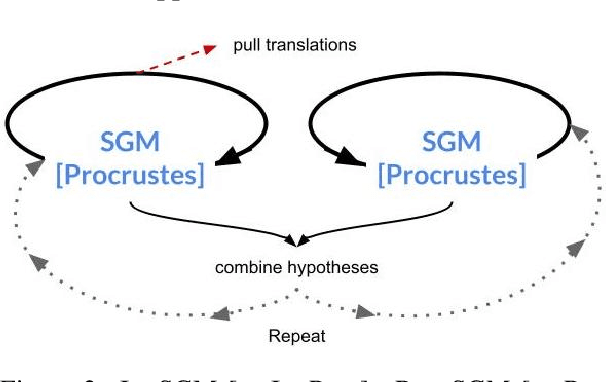
Abstract:Much recent work in bilingual lexicon induction (BLI) views word embeddings as vectors in Euclidean space. As such, BLI is typically solved by finding a linear transformation that maps embeddings to a common space. Alternatively, word embeddings may be understood as nodes in a weighted graph. This framing allows us to examine a node's graph neighborhood without assuming a linear transform, and exploits new techniques from the graph matching optimization literature. These contrasting approaches have not been compared in BLI so far. In this work, we study the behavior of Euclidean versus graph-based approaches to BLI under differing data conditions and show that they complement each other when combined. We release our code at https://github.com/kellymarchisio/euc-v-graph-bli.
Leveraging semantically similar queries for ranking via combining representations
Jun 23, 2021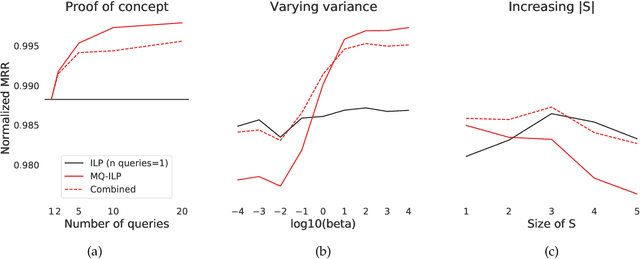
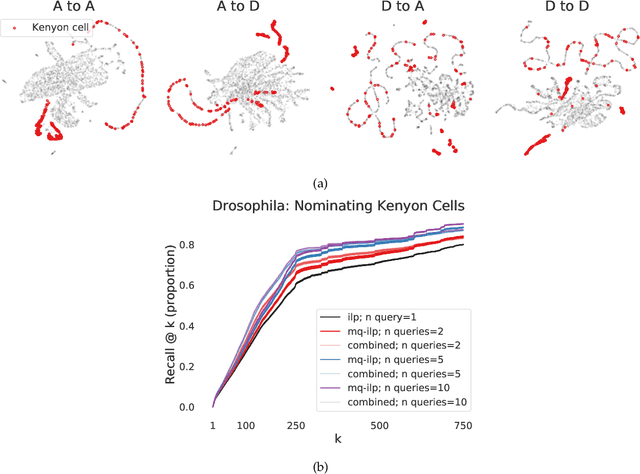
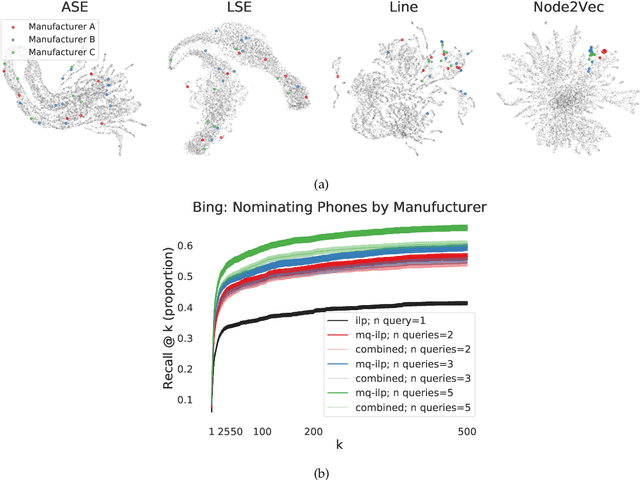
Abstract:In modern ranking problems, different and disparate representations of the items to be ranked are often available. It is sensible, then, to try to combine these representations to improve ranking. Indeed, learning to rank via combining representations is both principled and practical for learning a ranking function for a particular query. In extremely data-scarce settings, however, the amount of labeled data available for a particular query can lead to a highly variable and ineffective ranking function. One way to mitigate the effect of the small amount of data is to leverage information from semantically similar queries. Indeed, as we demonstrate in simulation settings and real data examples, when semantically similar queries are available it is possible to gainfully use them when ranking with respect to a particular query. We describe and explore this phenomenon in the context of the bias-variance trade off and apply it to the data-scarce settings of a Bing navigational graph and the Drosophila larva connectome.
Dynamic Silos: Modularity in intra-organizational communication networks during the Covid-19 pandemic
May 01, 2021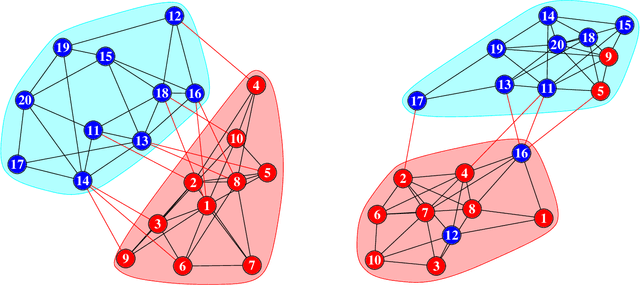
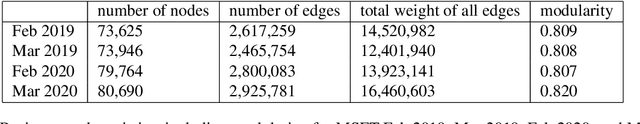
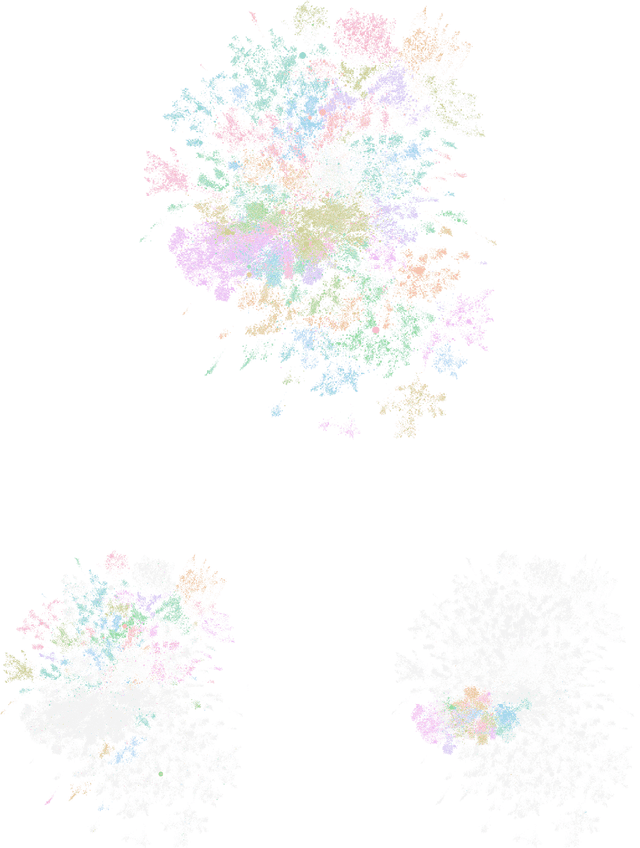
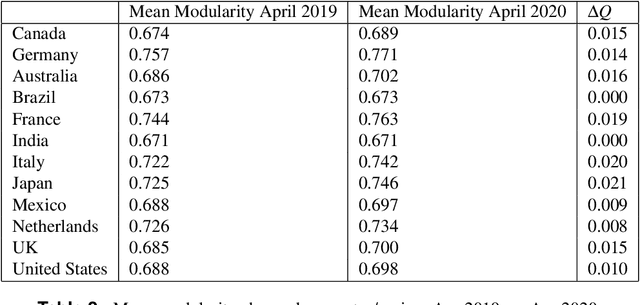
Abstract:Workplace communications around the world were drastically altered by Covid-19, work-from-home orders, and the rise of remote work. We analyze aggregated, anonymized metadata from over 360 billion emails within over 4000 organizations worldwide to examine changes in network community structures from 2019 through 2020. We find that, during 2020, organizations around the world became more siloed, evidenced by increased modularity. This shift was concurrent with decreased stability, indicating that organizational siloes had less stable membership. We provide initial insights into the implications of these network changes -- which we term dynamic silos -- for organizational performance and innovation.
Learning to rank via combining representations
May 20, 2020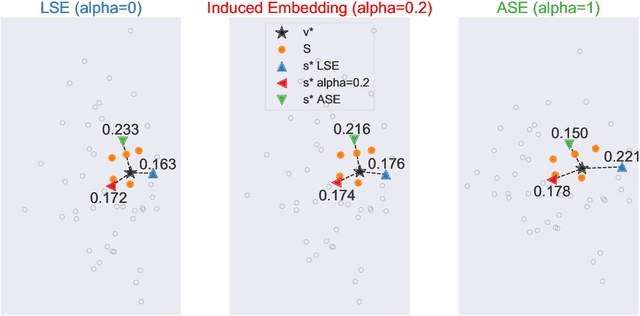
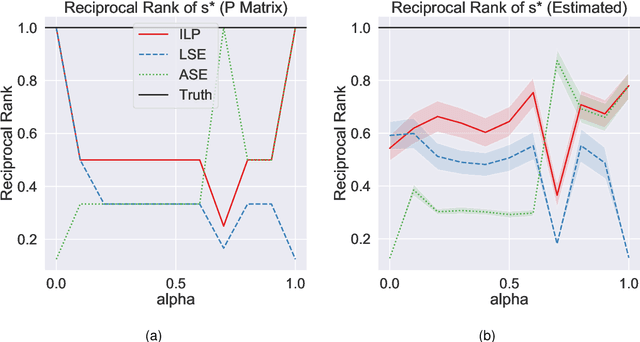
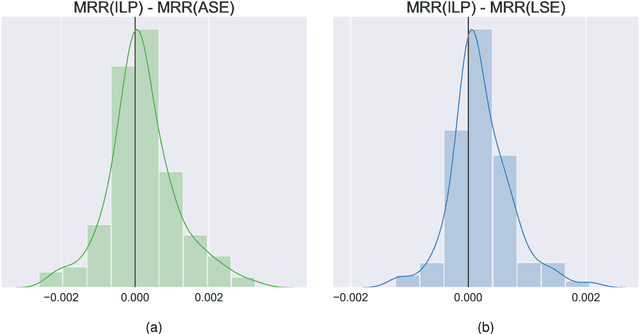
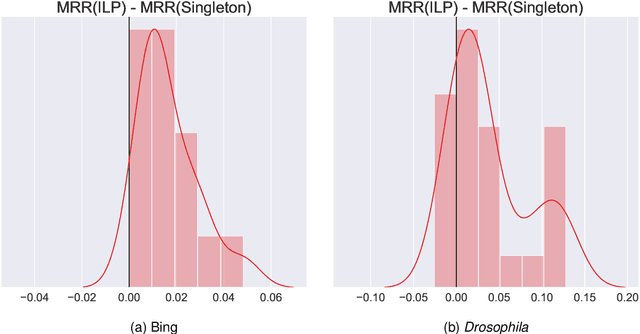
Abstract:Learning to rank -- producing a ranked list of items specific to a query and with respect to a set of supervisory items -- is a problem of general interest. The setting we consider is one in which no analytic description of what constitutes a good ranking is available. Instead, we have a collection of representations and supervisory information consisting of a (target item, interesting items set) pair. We demonstrate -- analytically, in simulation, and in real data examples -- that learning to rank via combining representations using an integer linear program is effective when the supervision is as light as "these few items are similar to your item of interest." While this nomination task is of general interest, for specificity we present our methodology from the perspective of vertex nomination in graphs. The methodology described herein is model agnostic.
Vertex Nomination, Consistent Estimation, and Adversarial Modification
May 15, 2019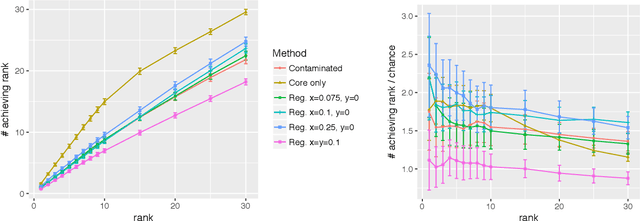
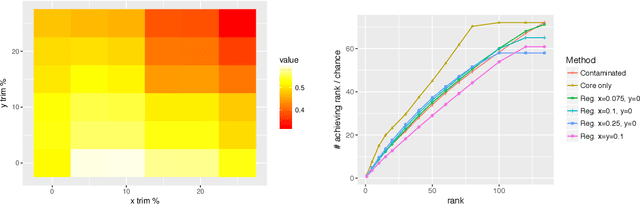
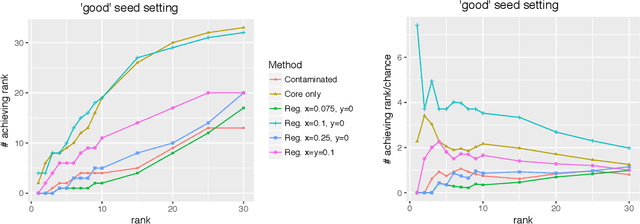
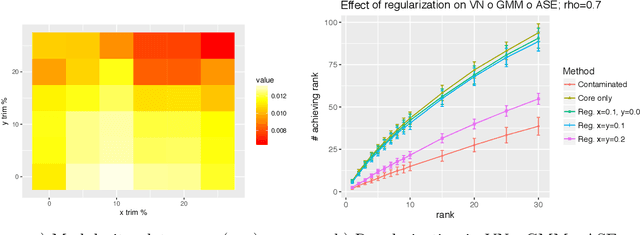
Abstract:Given a pair of graphs $G_1$ and $G_2$ and a vertex set of interest in $G_1$, the vertex nomination problem seeks to find the corresponding vertices of interest in $G_2$ (if they exist) and produce a rank list of the vertices in $G_2$, with the corresponding vertices of interest in $G_2$ concentrating, ideally, at the top of the rank list. In this paper we study the effect of an adversarial contamination model on the performance of a spectral graph embedding-based vertex nomination scheme. In both real and simulated examples, we demonstrate that this vertex nomination scheme performs effectively in the uncontaminated setting; adversarial network contamination adversely impacts the performance of our VN scheme; and network regularization successfully mitigates the impact of the contamination. In addition to furthering the theoretic basis of consistency in vertex nomination, the adversarial noise model posited herein is grounded in theoretical developments that allow us to frame the role of an adversary in terms of maximal vertex nomination consistency classes.
Simultaneous Dimensionality and Complexity Model Selection for Spectral Graph Clustering
Apr 05, 2019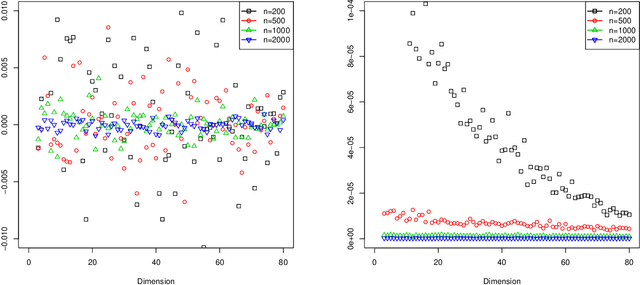
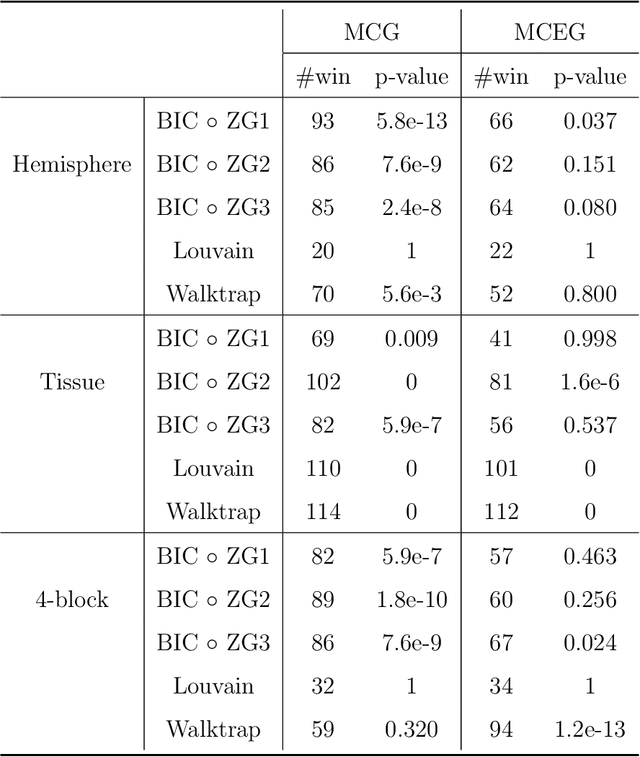
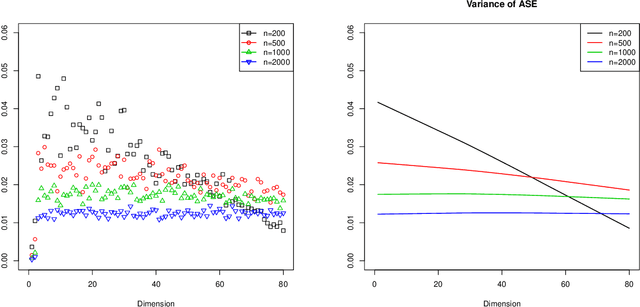
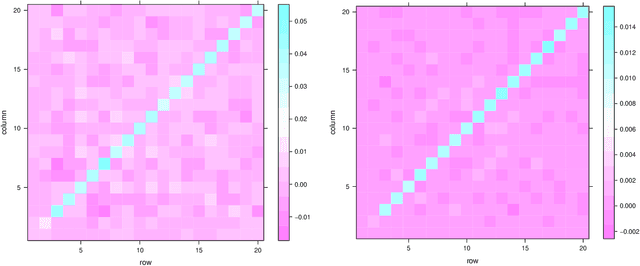
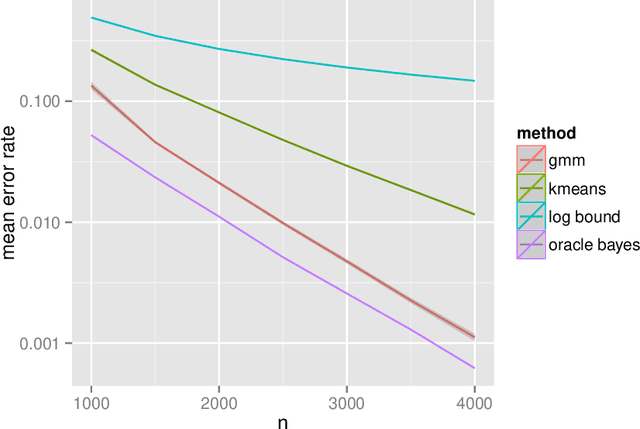
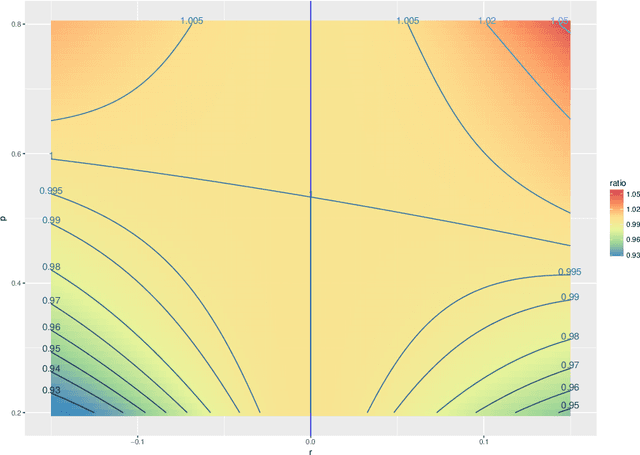
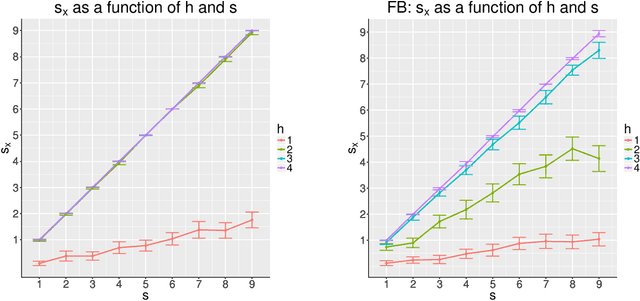
Abstract:Our problem of interest is to cluster vertices of a graph by identifying its underlying community structure. Among various vertex clustering approaches, spectral clustering is one of the most popular methods, because it is easy to implement while often outperforming traditional clustering algorithms. However, there are two inherent model selection problems in spectral clustering, namely estimating the embedding dimension and number of clusters. This paper attempts to address the issue by establishing a novel model selection framework specifically for vertex clustering on graphs under a stochastic block model. The first contribution is a probabilistic model which approximates the distribution of the extended spectral embedding of a graph. The model is constructed based on a theoretical result of asymptotic normality of the informative part of the embedding, and on a simulation result of limiting behavior of the redundant part of the embedding. The second contribution is a simultaneous model selection framework. In contrast with the traditional approaches, our model selection procedure estimates embedding dimension and number of clusters simultaneously. Based on our proposed distributional model, a theorem on the consistency of the estimates of model parameters is stated and proven. The theorem provides a statistical support for the validity of our method. Heuristic algorithms via the simultaneous model selection framework for vertex clustering are proposed, with good performance shown in the experiment on synthetic data and on the real application of connectome analysis.
On a 'Two Truths' Phenomenon in Spectral Graph Clustering
Sep 07, 2018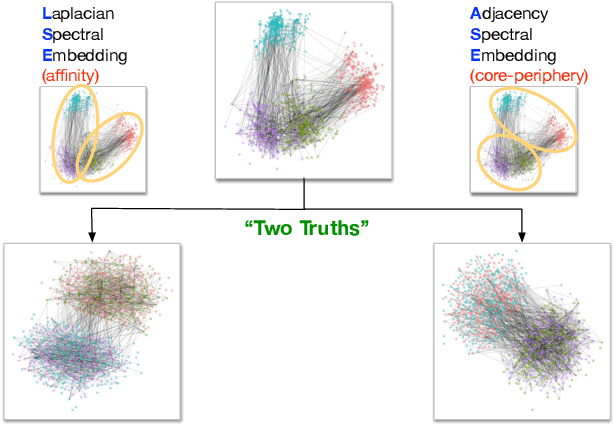
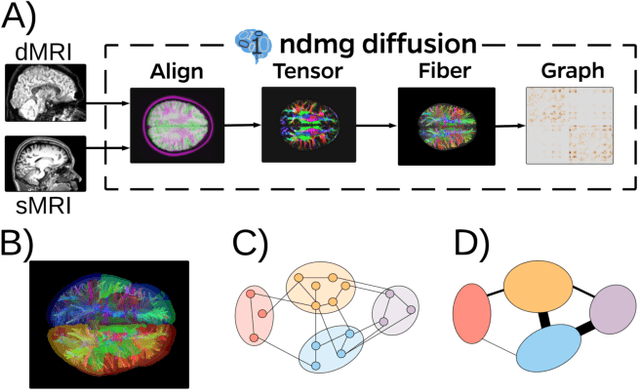
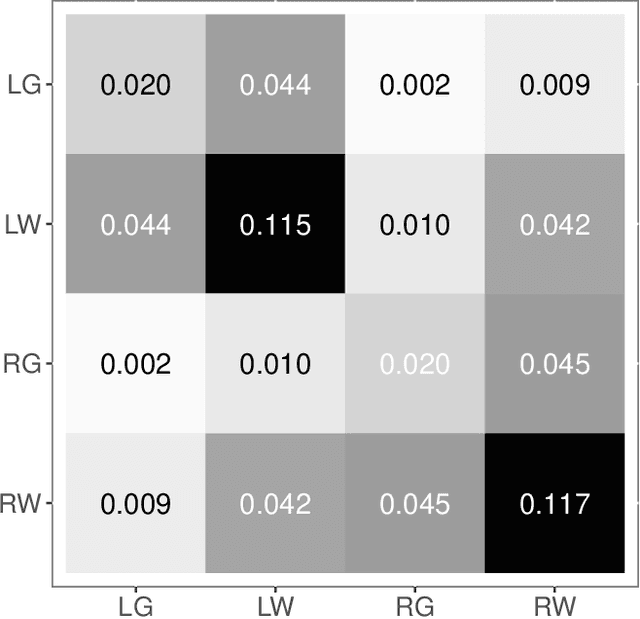
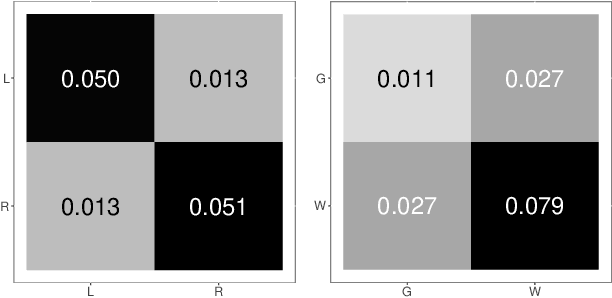
Abstract:Clustering is concerned with coherently grouping observations without any explicit concept of true groupings. Spectral graph clustering - clustering the vertices of a graph based on their spectral embedding - is commonly approached via K-means (or, more generally, Gaussian mixture model) clustering composed with either Laplacian or Adjacency spectral embedding (LSE or ASE). Recent theoretical results provide new understanding of the problem and solutions, and lead us to a 'Two Truths' LSE vs. ASE spectral graph clustering phenomenon convincingly illustrated here via a diffusion MRI connectome data set: the different embedding methods yield different clustering results, with LSE capturing left hemisphere/right hemisphere affinity structure and ASE capturing gray matter/white matter core-periphery structure.
Matched Filters for Noisy Induced Subgraph Detection
Mar 06, 2018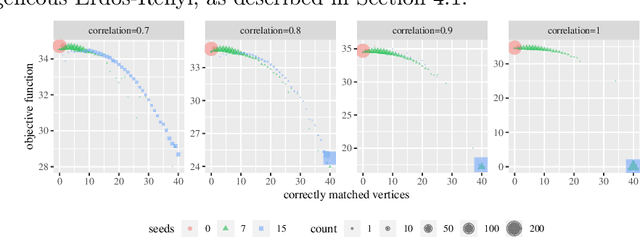

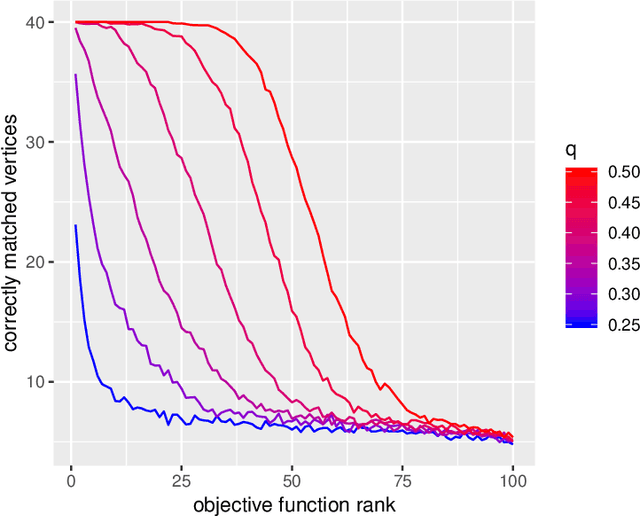
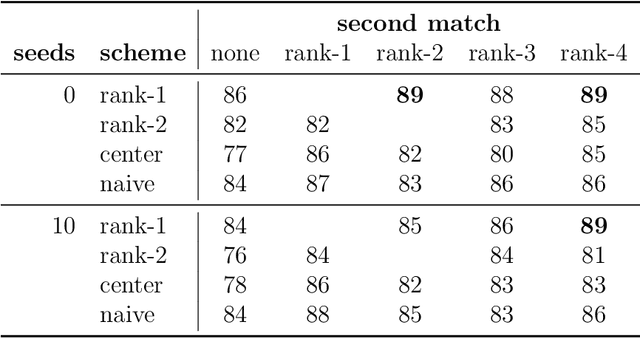
Abstract:We consider the problem of finding the vertex correspondence between two graphs with different number of vertices where the smaller graph is still potentially large. We propose a solution to this problem via a graph matching matched filter: padding the smaller graph in different ways and then using graph matching methods to align it to the larger network. Under a statistical model for correlated pairs of graphs, which yields a noisy copy of the small graph within the larger graph, the resulting optimization problem can be guaranteed to recover the true vertex correspondence between the networks, though there are currently no efficient algorithms for solving this problem. We consider an approach that exploits a partially known correspondence and show via varied simulations and applications to the Drosophila connectome that in practice this approach can achieve good performance.
Statistical inference on random dot product graphs: a survey
Sep 16, 2017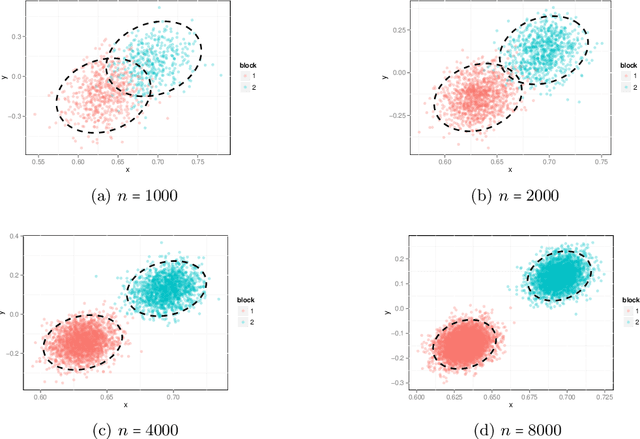

Abstract:The random dot product graph (RDPG) is an independent-edge random graph that is analytically tractable and, simultaneously, either encompasses or can successfully approximate a wide range of random graphs, from relatively simple stochastic block models to complex latent position graphs. In this survey paper, we describe a comprehensive paradigm for statistical inference on random dot product graphs, a paradigm centered on spectral embeddings of adjacency and Laplacian matrices. We examine the analogues, in graph inference, of several canonical tenets of classical Euclidean inference: in particular, we summarize a body of existing results on the consistency and asymptotic normality of the adjacency and Laplacian spectral embeddings, and the role these spectral embeddings can play in the construction of single- and multi-sample hypothesis tests for graph data. We investigate several real-world applications, including community detection and classification in large social networks and the determination of functional and biologically relevant network properties from an exploratory data analysis of the Drosophila connectome. We outline requisite background and current open problems in spectral graph inference.
* An expository survey paper on a comprehensive paradigm for inference for random dot product graphs, centered on graph adjacency and Laplacian spectral embeddings. Paper outlines requisite background; summarizes theory, methodology, and applications from previous and ongoing work; and closes with a discussion of several open problems
Vertex Nomination Via Local Neighborhood Matching
Jul 22, 2017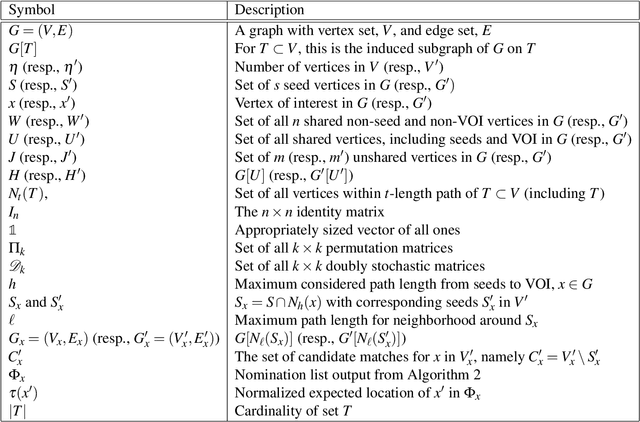
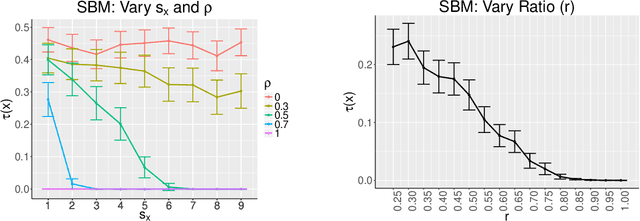
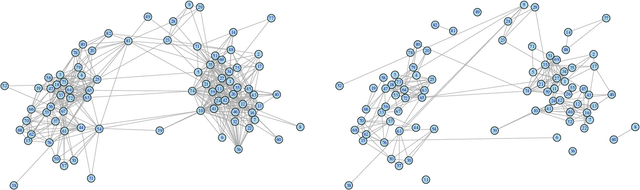
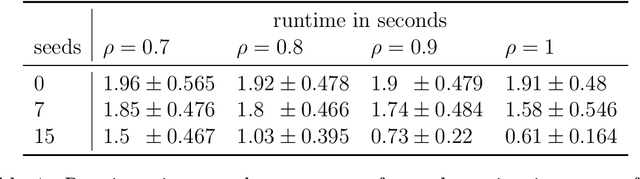
Abstract:Consider two networks on overlapping, non-identical vertex sets. Given vertices of interest in the first network, we seek to identify the corresponding vertices, if any exist, in the second network. While in moderately sized networks graph matching methods can be applied directly to recover the missing correspondences, herein we present a principled methodology appropriate for situations in which the networks are too large for brute-force graph matching. Our methodology identifies vertices in a local neighborhood of the vertices of interest in the first network that have verifiable corresponding vertices in the second network. Leveraging these known correspondences, referred to as seeds, we match the induced subgraphs in each network generated by the neighborhoods of these verified seeds, and rank the vertices of the second network in terms of the most likely matches to the original vertices of interest. We demonstrate the applicability of our methodology through simulations and real data examples.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge