Yiyu Shi
Federated Self-Supervised Contrastive Learning and Masked Autoencoder for Dermatological Disease Diagnosis
Aug 24, 2022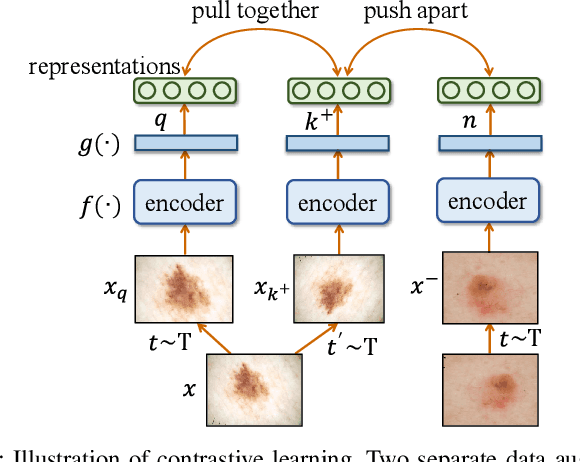
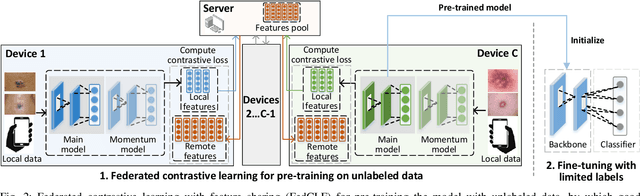
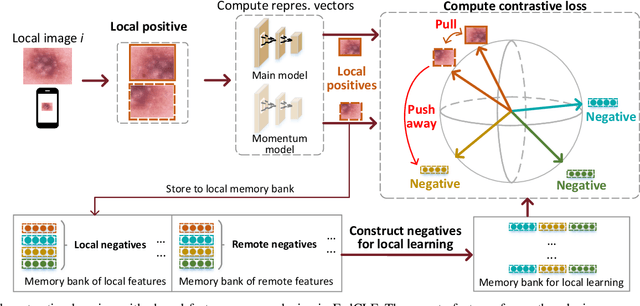
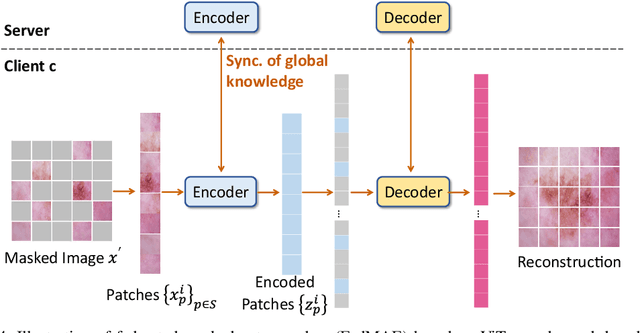
Abstract:In dermatological disease diagnosis, the private data collected by mobile dermatology assistants exist on distributed mobile devices of patients. Federated learning (FL) can use decentralized data to train models while keeping data local. Existing FL methods assume all the data have labels. However, medical data often comes without full labels due to high labeling costs. Self-supervised learning (SSL) methods, contrastive learning (CL) and masked autoencoders (MAE), can leverage the unlabeled data to pre-train models, followed by fine-tuning with limited labels. However, combining SSL and FL has unique challenges. For example, CL requires diverse data but each device only has limited data. For MAE, while Vision Transformer (ViT) based MAE has higher accuracy over CNNs in centralized learning, MAE's performance in FL with unlabeled data has not been investigated. Besides, the ViT synchronization between the server and clients is different from traditional CNNs. Therefore, special synchronization methods need to be designed. In this work, we propose two federated self-supervised learning frameworks for dermatological disease diagnosis with limited labels. The first one features lower computation costs, suitable for mobile devices. The second one features high accuracy and fits high-performance servers. Based on CL, we proposed federated contrastive learning with feature sharing (FedCLF). Features are shared for diverse contrastive information without sharing raw data for privacy. Based on MAE, we proposed FedMAE. Knowledge split separates the global and local knowledge learned from each client. Only global knowledge is aggregated for higher generalization performance. Experiments on dermatological disease datasets show superior accuracy of the proposed frameworks over state-of-the-arts.
Achieving Fairness in Dermatological Disease Diagnosis through Automatic Weight Adjusting Federated Learning and Personalization
Aug 23, 2022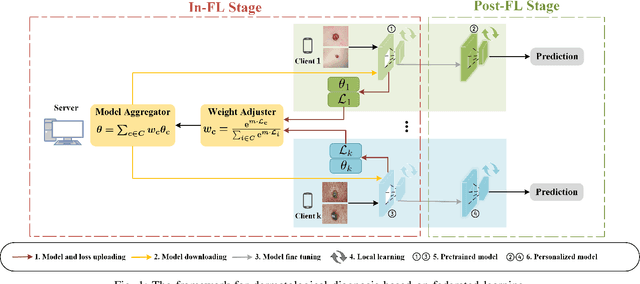
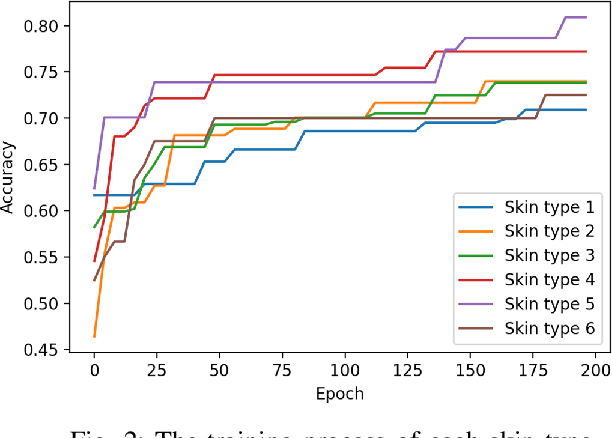
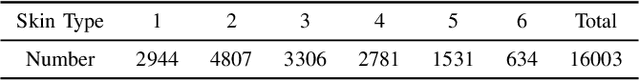
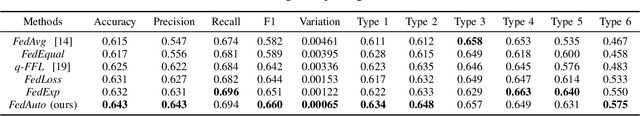
Abstract:Dermatological diseases pose a major threat to the global health, affecting almost one-third of the world's population. Various studies have demonstrated that early diagnosis and intervention are often critical to prognosis and outcome. To this end, the past decade has witnessed the rapid evolvement of deep learning based smartphone apps, which allow users to conveniently and timely identify issues that have emerged around their skins. In order to collect sufficient data needed by deep learning and at the same time protect patient privacy, federated learning is often used, where individual clients aggregate a global model while keeping datasets local. However, existing federated learning frameworks are mostly designed to optimize the overall performance, while common dermatological datasets are heavily imbalanced. When applying federated learning to such datasets, significant disparities in diagnosis accuracy may occur. To address such a fairness issue, this paper proposes a fairness-aware federated learning framework for dermatological disease diagnosis. The framework is divided into two stages: In the first in-FL stage, clients with different skin types are trained in a federated learning process to construct a global model for all skin types. An automatic weight aggregator is used in this process to assign higher weights to the client with higher loss, and the intensity of the aggregator is determined by the level of difference between losses. In the latter post-FL stage, each client fine-tune its personalized model based on the global model in the in-FL stage. To achieve better fairness, models from different epochs are selected for each client to keep the accuracy difference of different skin types within 0.05. Experiments indicate that our proposed framework effectively improves both fairness and accuracy compared with the state-of-the-art.
Distributed Contrastive Learning for Medical Image Segmentation
Aug 07, 2022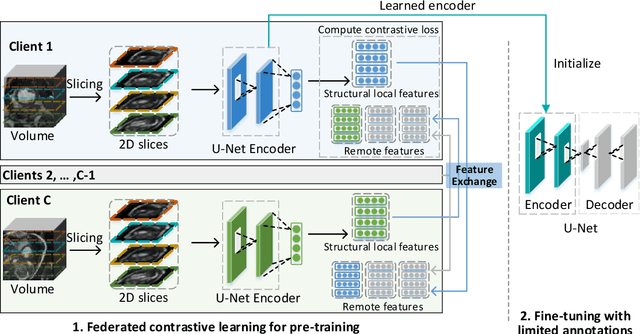
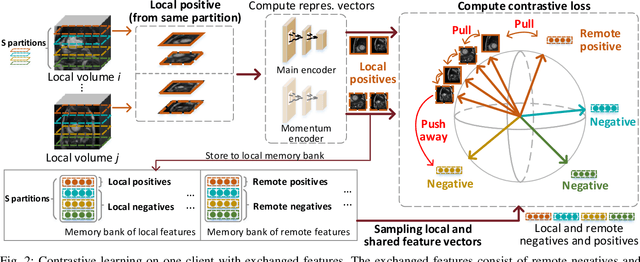
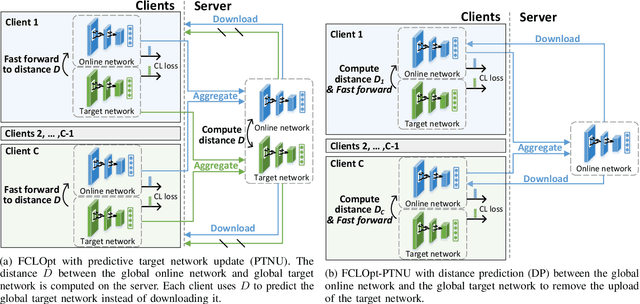
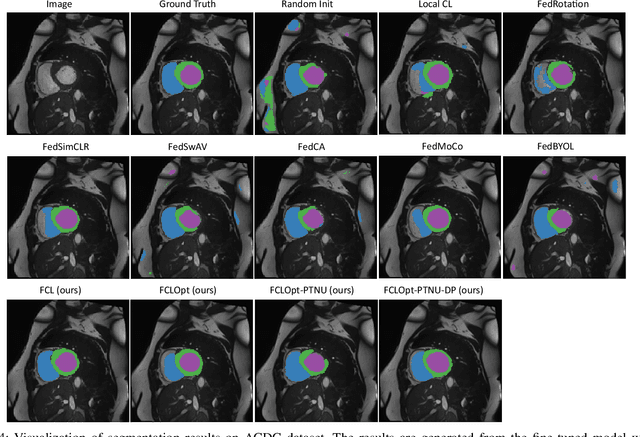
Abstract:Supervised deep learning needs a large amount of labeled data to achieve high performance. However, in medical imaging analysis, each site may only have a limited amount of data and labels, which makes learning ineffective. Federated learning (FL) can learn a shared model from decentralized data. But traditional FL requires fully-labeled data for training, which is very expensive to obtain. Self-supervised contrastive learning (CL) can learn from unlabeled data for pre-training, followed by fine-tuning with limited annotations. However, when adopting CL in FL, the limited data diversity on each site makes federated contrastive learning (FCL) ineffective. In this work, we propose two federated self-supervised learning frameworks for volumetric medical image segmentation with limited annotations. The first one features high accuracy and fits high-performance servers with high-speed connections. The second one features lower communication costs, suitable for mobile devices. In the first framework, features are exchanged during FCL to provide diverse contrastive data to each site for effective local CL while keeping raw data private. Global structural matching aligns local and remote features for a unified feature space among different sites. In the second framework, to reduce the communication cost for feature exchanging, we propose an optimized method FCLOpt that does not rely on negative samples. To reduce the communications of model download, we propose the predictive target network update (PTNU) that predicts the parameters of the target network. Based on PTNU, we propose the distance prediction (DP) to remove most of the uploads of the target network. Experiments on a cardiac MRI dataset show the proposed two frameworks substantially improve the segmentation and generalization performance compared with state-of-the-art techniques.
PAN: Pulse Ansatz on NISQ Machines
Aug 02, 2022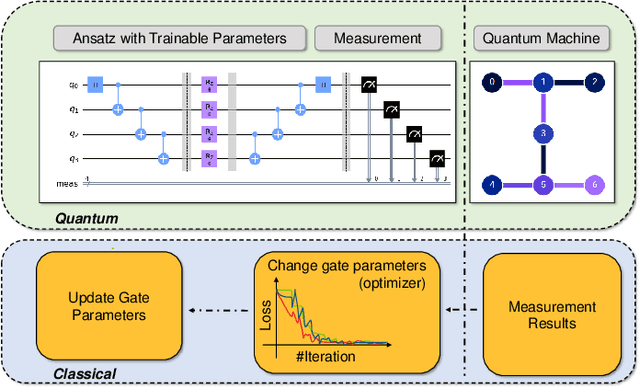
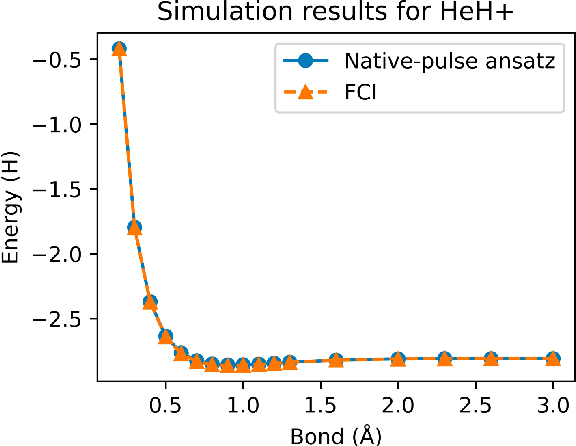
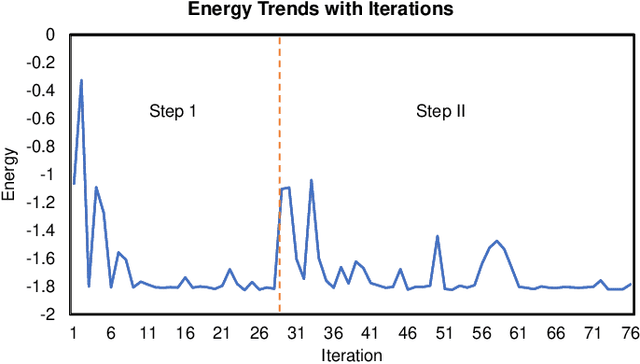
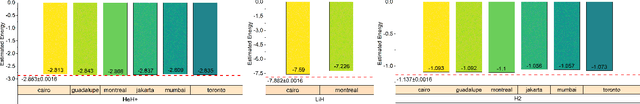
Abstract:Variational quantum algorithms (VQAs) have demonstrated great potentials in the NISQ era. In the workflow of VQA, the parameters of ansatz are iteratively updated to approximate the desired quantum states. We have seen various efforts to draft better ansatz with less gates. In quantum computers, the gate ansatz will eventually be transformed into control signals such as microwave pulses on transmons. And the control pulses need elaborate calibration to minimize the errors such as over-rotation and under-rotation. In the case of VQAs, this procedure will introduce redundancy, but the variational properties of VQAs can naturally handle problems of over-rotation and under-rotation by updating the amplitude and frequency parameters. Therefore, we propose PAN, a native-pulse ansatz generator framework for VQAs. We generate native-pulse ansatz with trainable parameters for amplitudes and frequencies. In our proposed PAN, we are tuning parametric pulses, which are natively supported on NISQ computers. Considering that parameter-shift rules do not hold for native-pulse ansatz, we need to deploy non-gradient optimizers. To constrain the number of parameters sent to the optimizer, we adopt a progressive way to generate our native-pulse ansatz. Experiments are conducted on both simulators and quantum devices to validate our methods. When adopted on NISQ machines, PAN obtained improved the performance with decreased latency by an average of 86%. PAN is able to achieve 99.336% and 96.482% accuracy for VQE tasks on H2 and HeH+ respectively, even with considerable noises in NISQ machines.
Contrastive Image Synthesis and Self-supervised Feature Adaptation for Cross-Modality Biomedical Image Segmentation
Jul 27, 2022
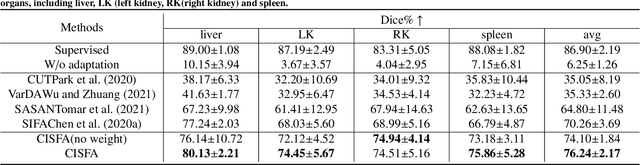
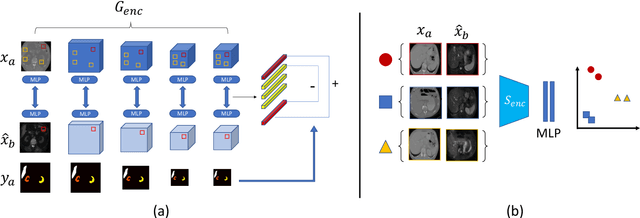
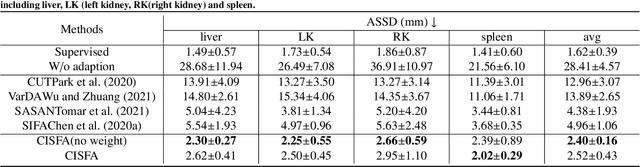
Abstract:This work presents a novel framework CISFA (Contrastive Image synthesis and Self-supervised Feature Adaptation)that builds on image domain translation and unsupervised feature adaptation for cross-modality biomedical image segmentation. Different from existing works, we use a one-sided generative model and add a weighted patch-wise contrastive loss between sampled patches of the input image and the corresponding synthetic image, which serves as shape constraints. Moreover, we notice that the generated images and input images share similar structural information but are in different modalities. As such, we enforce contrastive losses on the generated images and the input images to train the encoder of a segmentation model to minimize the discrepancy between paired images in the learned embedding space. Compared with existing works that rely on adversarial learning for feature adaptation, such a method enables the encoder to learn domain-independent features in a more explicit way. We extensively evaluate our methods on segmentation tasks containing CT and MRI images for abdominal cavities and whole hearts. Experimental results show that the proposed framework not only outputs synthetic images with less distortion of organ shapes, but also outperforms state-of-the-art domain adaptation methods by a large margin.
Computing-In-Memory Neural Network Accelerators for Safety-Critical Systems: Can Small Device Variations Be Disastrous?
Jul 15, 2022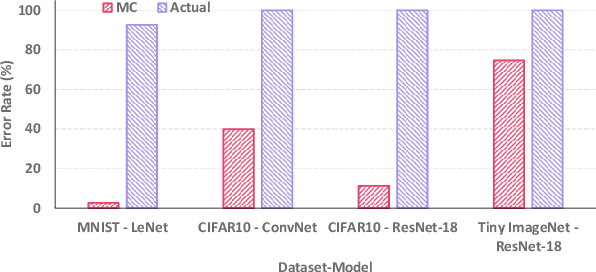



Abstract:Computing-in-Memory (CiM) architectures based on emerging non-volatile memory (NVM) devices have demonstrated great potential for deep neural network (DNN) acceleration thanks to their high energy efficiency. However, NVM devices suffer from various non-idealities, especially device-to-device variations due to fabrication defects and cycle-to-cycle variations due to the stochastic behavior of devices. As such, the DNN weights actually mapped to NVM devices could deviate significantly from the expected values, leading to large performance degradation. To address this issue, most existing works focus on maximizing average performance under device variations. This objective would work well for general-purpose scenarios. But for safety-critical applications, the worst-case performance must also be considered. Unfortunately, this has been rarely explored in the literature. In this work, we formulate the problem of determining the worst-case performance of CiM DNN accelerators under the impact of device variations. We further propose a method to effectively find the specific combination of device variation in the high-dimensional space that leads to the worst-case performance. We find that even with very small device variations, the accuracy of a DNN can drop drastically, causing concerns when deploying CiM accelerators in safety-critical applications. Finally, we show that surprisingly none of the existing methods used to enhance average DNN performance in CiM accelerators are very effective when extended to enhance the worst-case performance, and further research down the road is needed to address this problem.
RT-DNAS: Real-time Constrained Differentiable Neural Architecture Search for 3D Cardiac Cine MRI Segmentation
Jun 13, 2022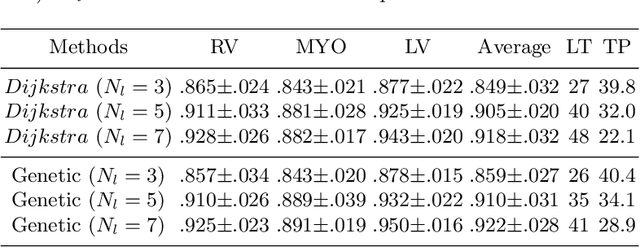

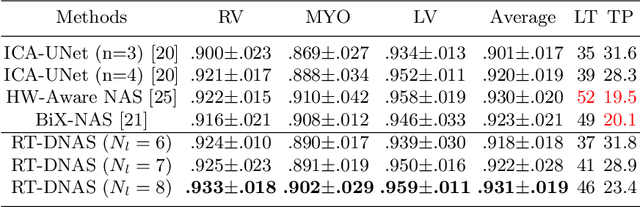
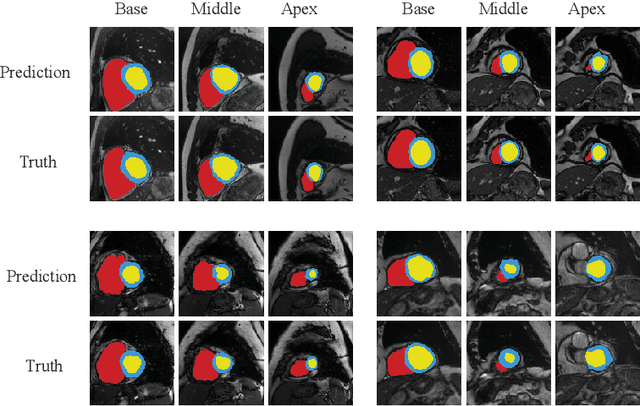
Abstract:Accurately segmenting temporal frames of cine magnetic resonance imaging (MRI) is a crucial step in various real-time MRI guided cardiac interventions. To achieve fast and accurate visual assistance, there are strict requirements on the maximum latency and minimum throughput of the segmentation framework. State-of-the-art neural networks on this task are mostly hand-crafted to satisfy these constraints while achieving high accuracy. On the other hand, while existing literature have demonstrated the power of neural architecture search (NAS) in automatically identifying the best neural architectures for various medical applications, they are mostly guided by accuracy, sometimes with computation complexity, and the importance of real-time constraints are overlooked. A major challenge is that such constraints are non-differentiable and are thus not compatible with the widely used differentiable NAS frameworks. In this paper, we present a strategy that directly handles real-time constraints in a differentiable NAS framework named RT-DNAS. Experiments on extended 2017 MICCAI ACDC dataset show that compared with state-of-the-art manually and automatically designed architectures, RT-DNAS is able to identify ones with better accuracy while satisfying the real-time constraints.
OTFPF: Optimal Transport-Based Feature Pyramid Fusion Network for Brain Age Estimation with 3D Overlapped ConvNeXt
May 11, 2022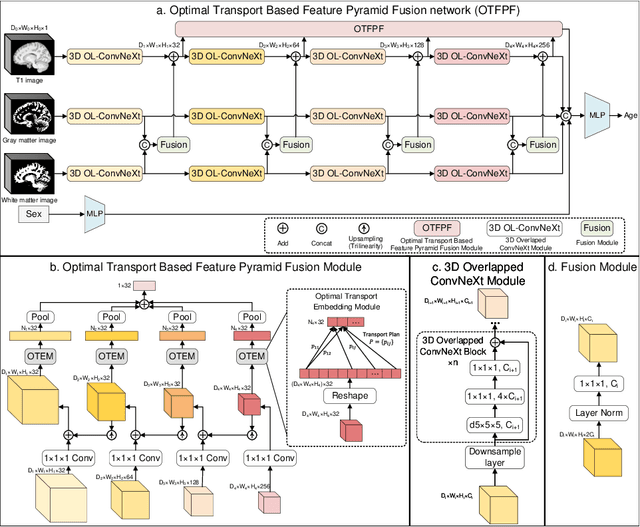
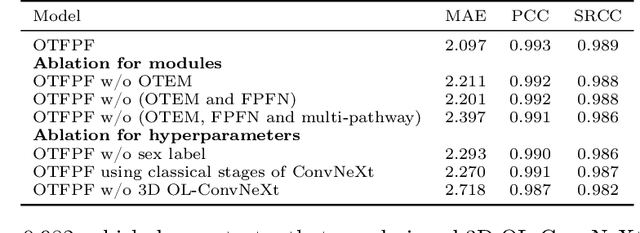
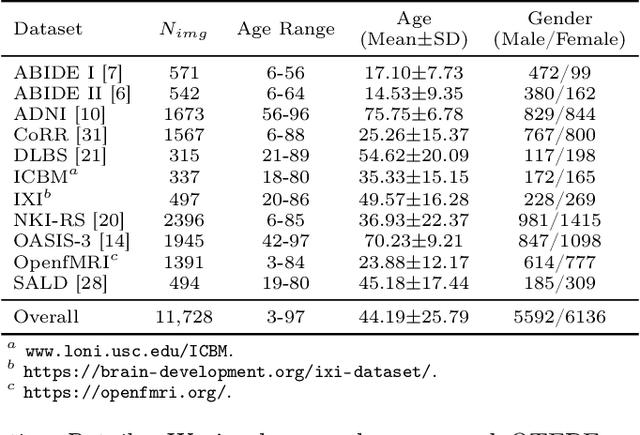
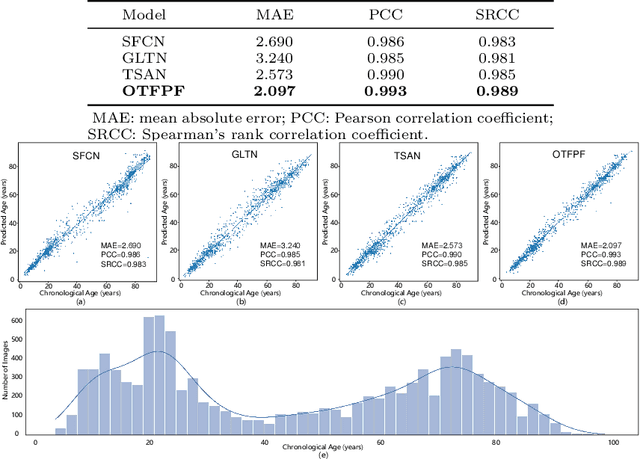

Abstract:Chronological age of healthy brain is able to be predicted using deep neural networks from T1-weighted magnetic resonance images (T1 MRIs), and the predicted brain age could serve as an effective biomarker for detecting aging-related diseases or disorders. In this paper, we propose an end-to-end neural network architecture, referred to as optimal transport based feature pyramid fusion (OTFPF) network, for the brain age estimation with T1 MRIs. The OTFPF consists of three types of modules: Optimal Transport based Feature Pyramid Fusion (OTFPF) module, 3D overlapped ConvNeXt (3D OL-ConvNeXt) module and fusion module. These modules strengthen the OTFPF network's understanding of each brain's semi-multimodal and multi-level feature pyramid information, and significantly improve its estimation performances. Comparing with recent state-of-the-art models, the proposed OTFPF converges faster and performs better. The experiments with 11,728 MRIs aged 3-97 years show that OTFPF network could provide accurate brain age estimation, yielding mean absolute error (MAE) of 2.097, Pearson's correlation coefficient (PCC) of 0.993 and Spearman's rank correlation coefficient (SRCC) of 0.989, between the estimated and chronological ages. Widespread quantitative experiments and ablation experiments demonstrate the superiority and rationality of OTFPF network. The codes and implement details will be released on GitHub: https://github.com/ZJU-Brain/OTFPF after final decision.
Federated Contrastive Learning for Volumetric Medical Image Segmentation
Apr 23, 2022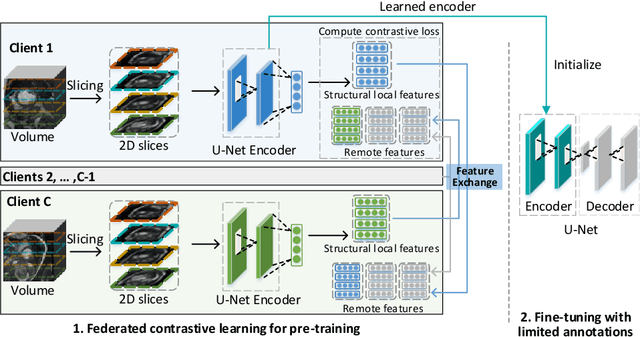
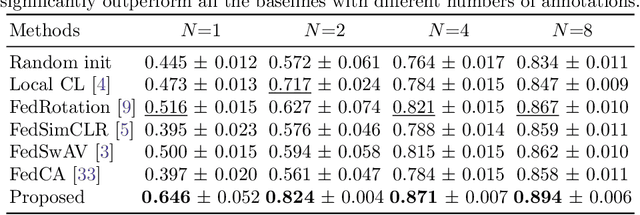
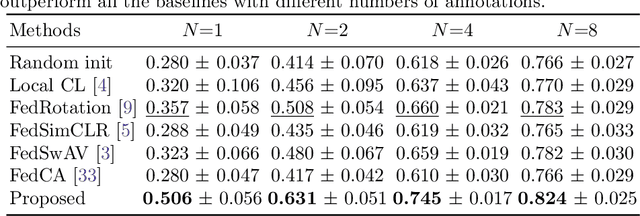
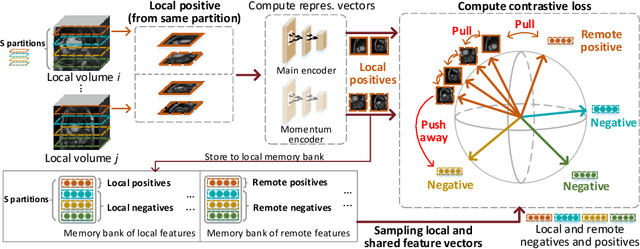

Abstract:Supervised deep learning needs a large amount of labeled data to achieve high performance. However, in medical imaging analysis, each site may only have a limited amount of data and labels, which makes learning ineffective. Federated learning (FL) can help in this regard by learning a shared model while keeping training data local for privacy. Traditional FL requires fully-labeled data for training, which is inconvenient or sometimes infeasible to obtain due to high labeling cost and the requirement of expertise. Contrastive learning (CL), as a self-supervised learning approach, can effectively learn from unlabeled data to pre-train a neural network encoder, followed by fine-tuning for downstream tasks with limited annotations. However, when adopting CL in FL, the limited data diversity on each client makes federated contrastive learning (FCL) ineffective. In this work, we propose an FCL framework for volumetric medical image segmentation with limited annotations. More specifically, we exchange the features in the FCL pre-training process such that diverse contrastive data are provided to each site for effective local CL while keeping raw data private. Based on the exchanged features, global structural matching further leverages the structural similarity to align local features to the remote ones such that a unified feature space can be learned among different sites. Experiments on a cardiac MRI dataset show the proposed framework substantially improves the segmentation performance compared with state-of-the-art techniques.
A Semi-Decoupled Approach to Fast and Optimal Hardware-Software Co-Design of Neural Accelerators
Mar 25, 2022
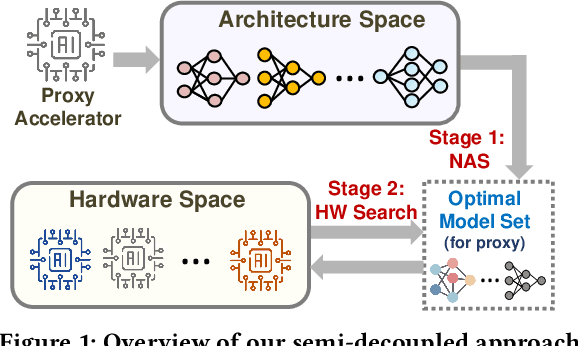
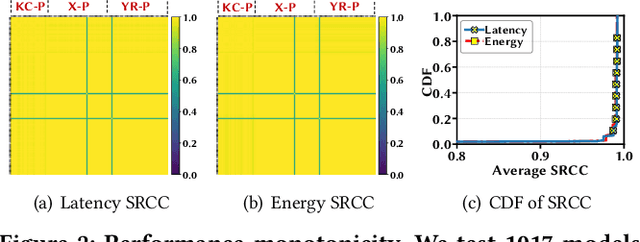
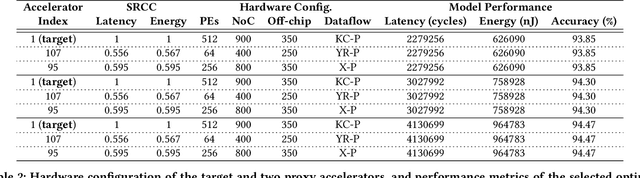
Abstract:In view of the performance limitations of fully-decoupled designs for neural architectures and accelerators, hardware-software co-design has been emerging to fully reap the benefits of flexible design spaces and optimize neural network performance. Nonetheless, such co-design also enlarges the total search space to practically infinity and presents substantial challenges. While the prior studies have been focusing on improving the search efficiency (e.g., via reinforcement learning), they commonly rely on co-searches over the entire architecture-accelerator design space. In this paper, we propose a \emph{semi}-decoupled approach to reduce the size of the total design space by orders of magnitude, yet without losing optimality. We first perform neural architecture search to obtain a small set of optimal architectures for one accelerator candidate. Importantly, this is also the set of (close-to-)optimal architectures for other accelerator designs based on the property that neural architectures' ranking orders in terms of inference latency and energy consumption on different accelerator designs are highly similar. Then, instead of considering all the possible architectures, we optimize the accelerator design only in combination with this small set of architectures, thus significantly reducing the total search cost. We validate our approach by conducting experiments on various architecture spaces for accelerator designs with different dataflows. Our results highlight that we can obtain the optimal design by only navigating over the reduced search space. The source code of this work is at \url{https://github.com/Ren-Research/CoDesign}.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge