Nick Pawlowski
Structured Uncertainty in the Observation Space of Variational Autoencoders
May 25, 2022
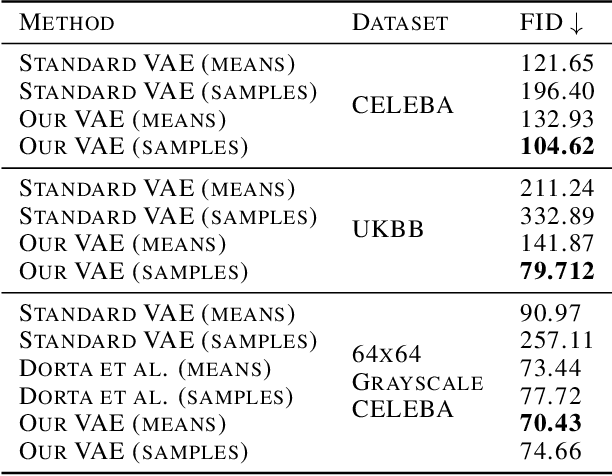
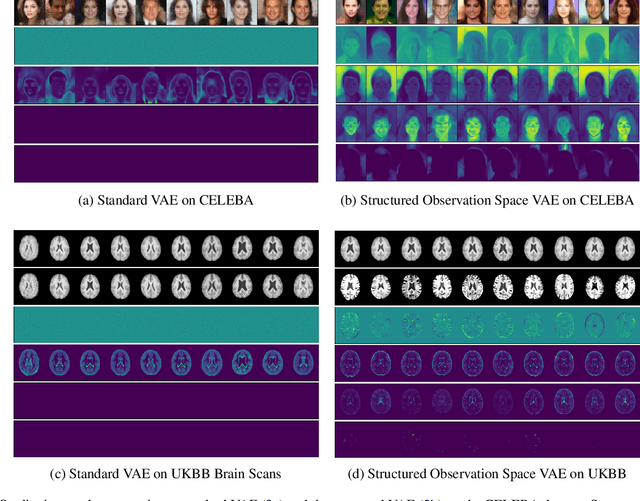

Abstract:Variational autoencoders (VAEs) are a popular class of deep generative models with many variants and a wide range of applications. Improvements upon the standard VAE mostly focus on the modelling of the posterior distribution over the latent space and the properties of the neural network decoder. In contrast, improving the model for the observational distribution is rarely considered and typically defaults to a pixel-wise independent categorical or normal distribution. In image synthesis, sampling from such distributions produces spatially-incoherent results with uncorrelated pixel noise, resulting in only the sample mean being somewhat useful as an output prediction. In this paper, we aim to stay true to VAE theory by improving the samples from the observational distribution. We propose an alternative model for the observation space, encoding spatial dependencies via a low-rank parameterisation. We demonstrate that this new observational distribution has the ability to capture relevant covariance between pixels, resulting in spatially-coherent samples. In contrast to pixel-wise independent distributions, our samples seem to contain semantically meaningful variations from the mean allowing the prediction of multiple plausible outputs with a single forward pass.
Deep End-to-end Causal Inference
Feb 04, 2022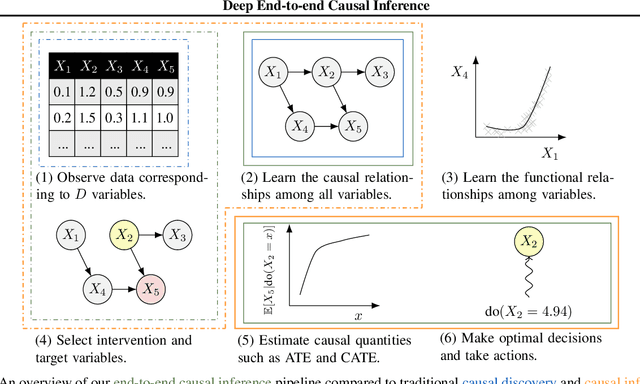
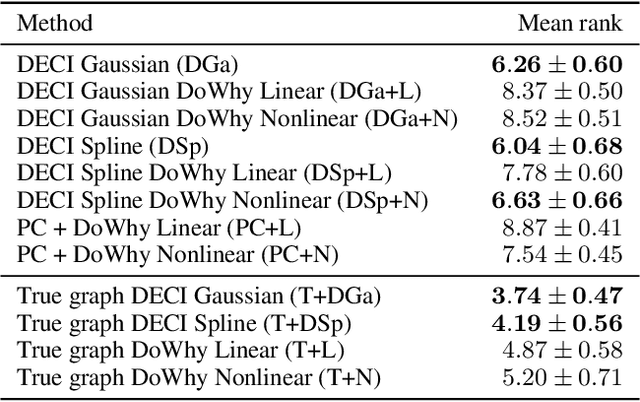
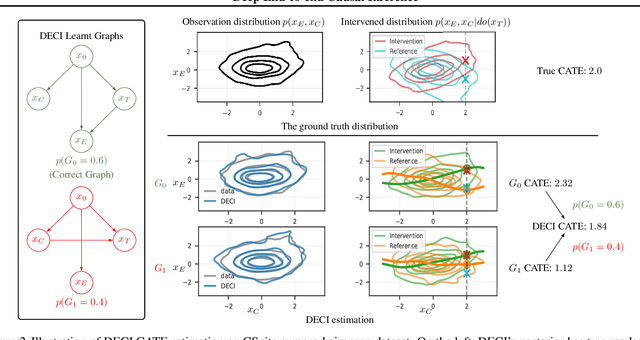
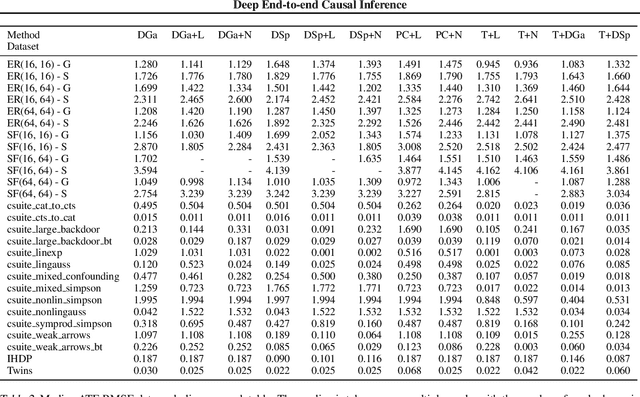
Abstract:Causal inference is essential for data-driven decision making across domains such as business engagement, medical treatment or policy making. However, research on causal discovery and inference has evolved separately, and the combination of the two domains is not trivial. In this work, we develop Deep End-to-end Causal Inference (DECI), a single flow-based method that takes in observational data and can perform both causal discovery and inference, including conditional average treatment effect (CATE) estimation. We provide a theoretical guarantee that DECI can recover the ground truth causal graph under mild assumptions. In addition, our method can handle heterogeneous, real-world, mixed-type data with missing values, allowing for both continuous and discrete treatment decisions. Moreover, the design principle of our method can generalize beyond DECI, providing a general End-to-end Causal Inference (ECI) recipe, which enables different ECI frameworks to be built using existing methods. Our results show the superior performance of DECI when compared to relevant baselines for both causal discovery and (C)ATE estimation in over a thousand experiments on both synthetic datasets and other causal machine learning benchmark datasets.
Does Your Dermatology Classifier Know What It Doesn't Know? Detecting the Long-Tail of Unseen Conditions
Apr 08, 2021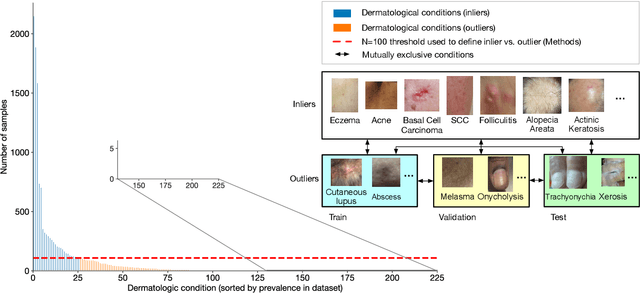
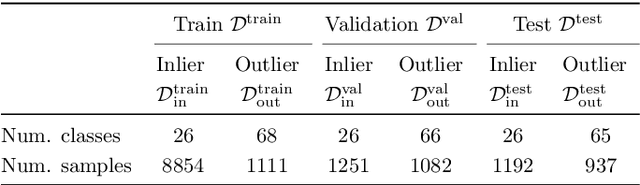
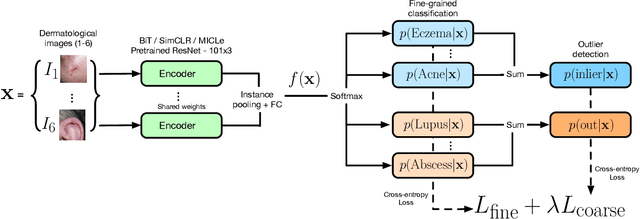
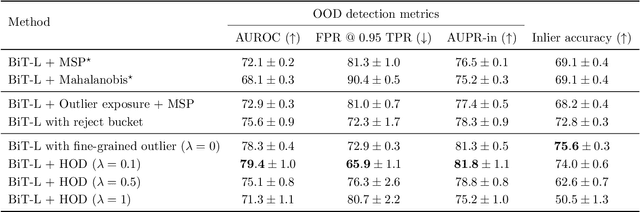
Abstract:We develop and rigorously evaluate a deep learning based system that can accurately classify skin conditions while detecting rare conditions for which there is not enough data available for training a confident classifier. We frame this task as an out-of-distribution (OOD) detection problem. Our novel approach, hierarchical outlier detection (HOD) assigns multiple abstention classes for each training outlier class and jointly performs a coarse classification of inliers vs. outliers, along with fine-grained classification of the individual classes. We demonstrate the effectiveness of the HOD loss in conjunction with modern representation learning approaches (BiT, SimCLR, MICLe) and explore different ensembling strategies for further improving the results. We perform an extensive subgroup analysis over conditions of varying risk levels and different skin types to investigate how the OOD detection performance changes over each subgroup and demonstrate the gains of our framework in comparison to baselines. Finally, we introduce a cost metric to approximate downstream clinical impact. We use this cost metric to compare the proposed method against a baseline system, thereby making a stronger case for the overall system effectiveness in a real-world deployment scenario.
Deep Structural Causal Models for Tractable Counterfactual Inference
Jun 11, 2020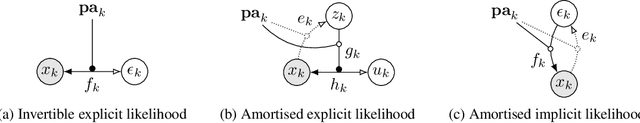
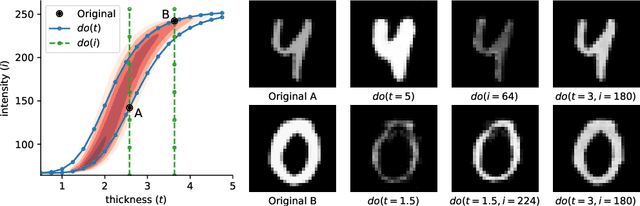
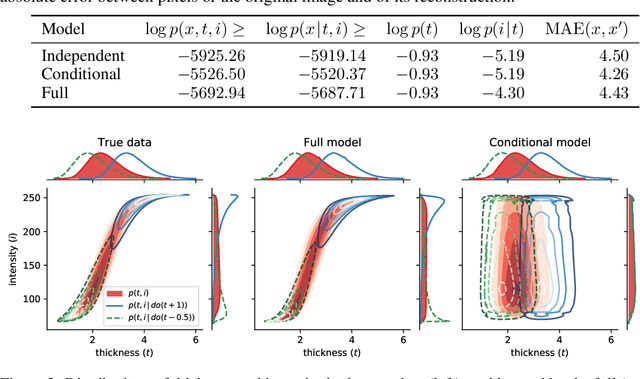
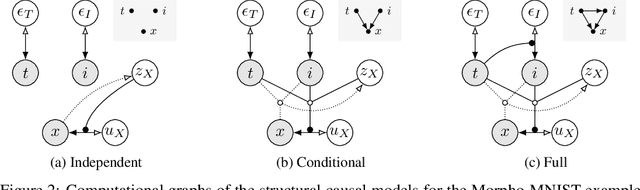

Abstract:We formulate a general framework for building structural causal models (SCMs) with deep learning components. The proposed approach employs normalising flows and variational inference to enable tractable inference of exogenous noise variables - a crucial step for counterfactual inference that is missing from existing deep causal learning methods. Our framework is validated on a synthetic dataset built on MNIST as well as on a real-world medical dataset of brain MRI scans. Our experimental results indicate that we can successfully train deep SCMs that are capable of all three levels of Pearl's ladder of causation: association, intervention, and counterfactuals, giving rise to a powerful new approach for answering causal questions in imaging applications and beyond. The code for all our experiments is available at https://github.com/biomedia-mira/deepscm.
Stochastic Segmentation Networks: Modelling Spatially Correlated Aleatoric Uncertainty
Jun 10, 2020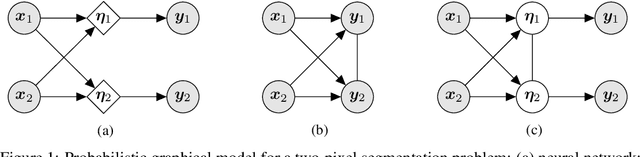
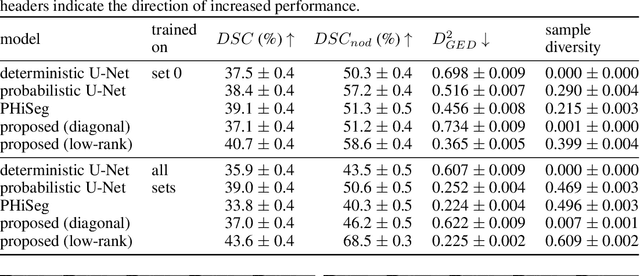

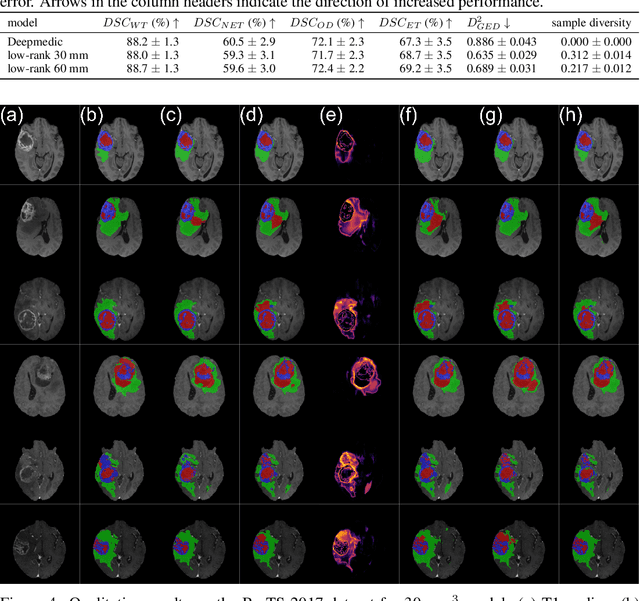
Abstract:In image segmentation, there is often more than one plausible solution for a given input. In medical imaging, for example, experts will often disagree about the exact location of object boundaries. Estimating this inherent uncertainty and predicting multiple plausible hypotheses is of great interest in many applications, yet this ability is lacking in most current deep learning methods. In this paper, we introduce stochastic segmentation networks (SSNs), an efficient probabilistic method for modelling aleatoric uncertainty with any image segmentation network architecture. In contrast to approaches that produce pixel-wise estimates, SSNs model joint distributions over entire label maps and thus can generate multiple spatially coherent hypotheses for a single image. By using a low-rank multivariate normal distribution over the logit space to model the probability of the label map given the image, we obtain a spatially consistent probability distribution that can be efficiently computed by a neural network without any changes to the underlying architecture. We tested our method on the segmentation of real-world medical data, including lung nodules in 2D CT and brain tumours in 3D multimodal MRI scans. SSNs outperform state-of-the-art for modelling correlated uncertainty in ambiguous images while being much simpler, more flexible, and more efficient.
An Explicit Local and Global Representation Disentanglement Framework with Applications in Deep Clustering and Unsupervised Object Detection
Feb 24, 2020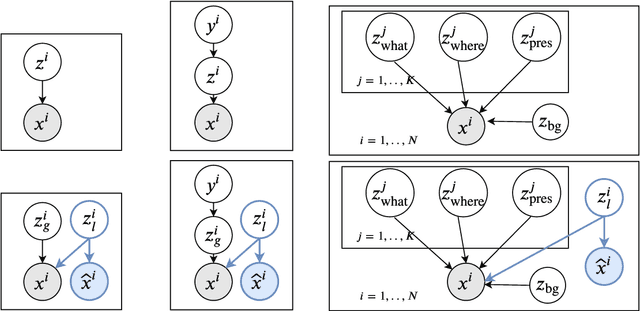

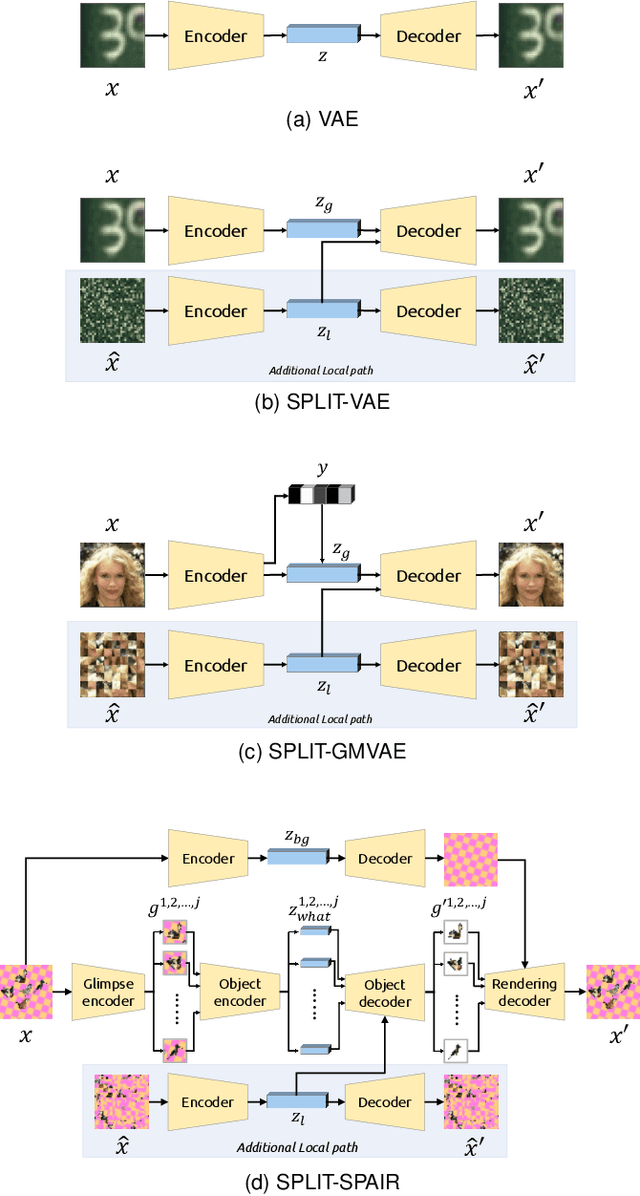
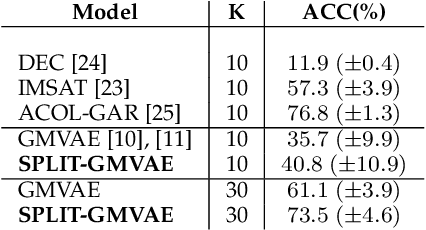
Abstract:Visual data can be understood at different levels of granularity, where global features correspond to semantic-level information and local features correspond to texture patterns. In this work, we propose a framework, called SPLIT, which allows us to disentangle local and global information into two separate sets of latent variables within the variational autoencoder (VAE) framework. Our framework adds generative assumption to the VAE by requiring a subset of the latent variables to generate an auxiliary set of observable data. This additional generative assumption primes the latent variables to local information and encourages the other latent variables to represent global information. We examine three different flavours of VAEs with different generative assumptions. We show that the framework can effectively disentangle local and global information within these models leads to improved representation, with better clustering and unsupervised object detection benchmarks. Finally, we establish connections between SPLIT and recent research in cognitive neuroscience regarding the disentanglement in human visual perception. The code for our experiments is at https://github.com/51616/split-vae .
Representation Disentanglement for Multi-task Learning with application to Fetal Ultrasound
Aug 21, 2019
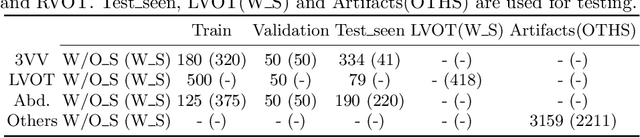
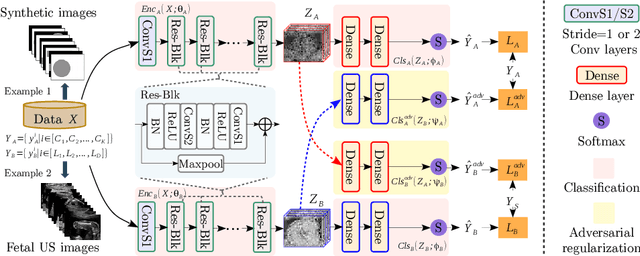
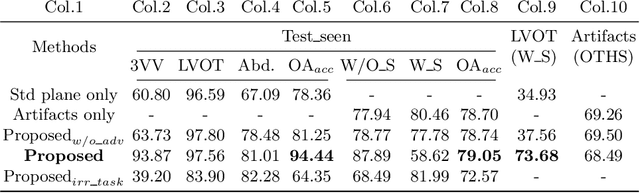
Abstract:One of the biggest challenges for deep learning algorithms in medical image analysis is the indiscriminate mixing of image properties, e.g. artifacts and anatomy. These entangled image properties lead to a semantically redundant feature encoding for the relevant task and thus lead to poor generalization of deep learning algorithms. In this paper we propose a novel representation disentanglement method to extract semantically meaningful and generalizable features for different tasks within a multi-task learning framework. Deep neural networks are utilized to ensure that the encoded features are maximally informative with respect to relevant tasks, while an adversarial regularization encourages these features to be disentangled and minimally informative about irrelevant tasks. We aim to use the disentangled representations to generalize the applicability of deep neural networks. We demonstrate the advantages of the proposed method on synthetic data as well as fetal ultrasound images. Our experiments illustrate that our method is capable of learning disentangled internal representations. It outperforms baseline methods in multiple tasks, especially on images with new properties, e.g. previously unseen artifacts in fetal ultrasound.
Needles in Haystacks: On Classifying Tiny Objects in Large Images
Aug 16, 2019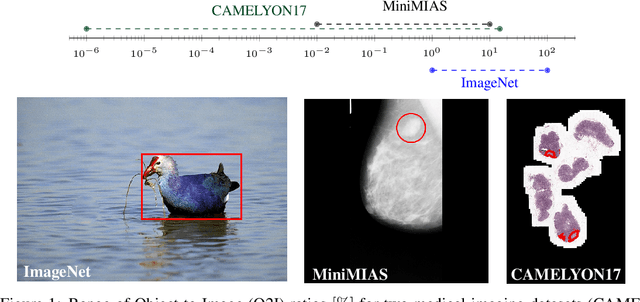
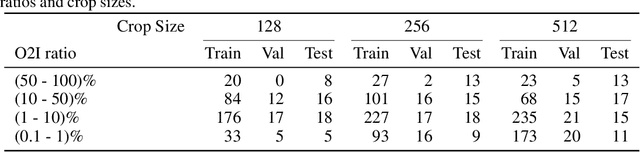
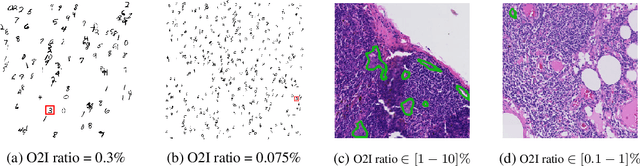
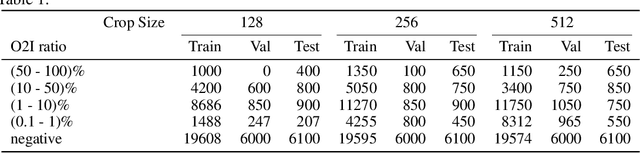
Abstract:In some computer vision domains, such as medical or hyperspectral imaging, we care about the classification of tiny objects in large images. However, most Convolutional Neural Networks (CNNs) for image classification were developed and analyzed using biased datasets that contain large objects, most often, in central image positions. To assess whether classical CNN architectures work well for tiny object classification we build a comprehensive testbed containing two datasets: one derived from MNIST digits and other from histopathology images. This testbed allows us to perform controlled experiments to stress-test CNN architectures using a broad spectrum of signal-to-noise ratios. Our observations suggest that: (1) There exists a limit to signal-to-noise below which CNNs fail to generalize and that this limit is affected by dataset size - more data leading to better performances; however, the amount of training data required for the model to generalize scales rapidly with the inverse of the object-to-image ratio (2) in general, higher capacity models exhibit better generalization; (3) when knowing the approximate object sizes, adapting receptive field is beneficial; and (4) for very small signal-to-noise ratio the choice of global pooling operation affects optimization, whereas for relatively large signal-to-noise values, all tested global pooling operations exhibit similar performance.
Is Texture Predictive for Age and Sex in Brain MRI?
Jul 25, 2019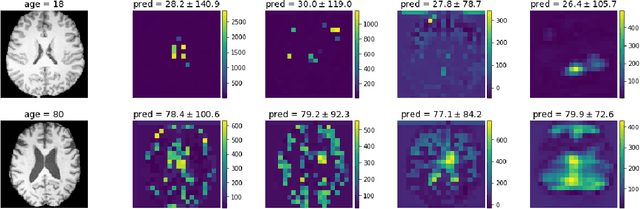
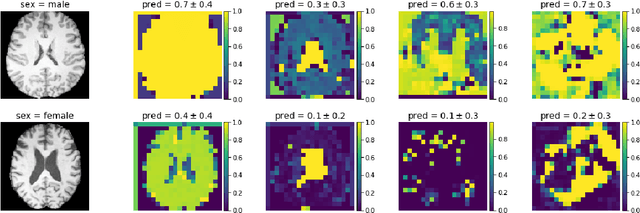
Abstract:Deep learning builds the foundation for many medical image analysis tasks where neuralnetworks are often designed to have a large receptive field to incorporate long spatialdependencies. Recent work has shown that large receptive fields are not always necessaryfor computer vision tasks on natural images. We explore whether this translates to certainmedical imaging tasks such as age and sex prediction from a T1-weighted brain MRI scans.
Deep Generative Models in the Real-World: An Open Challenge from Medical Imaging
Jun 14, 2018
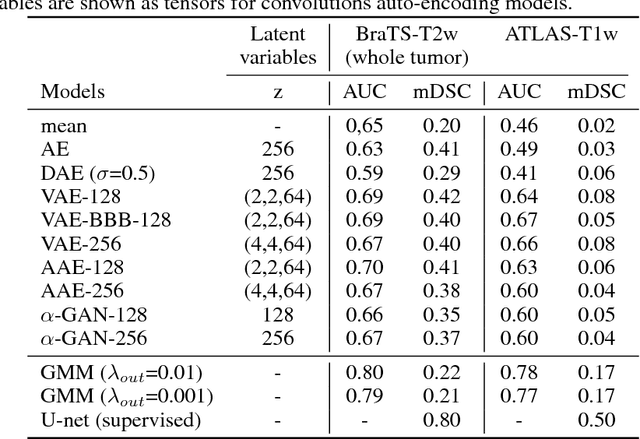
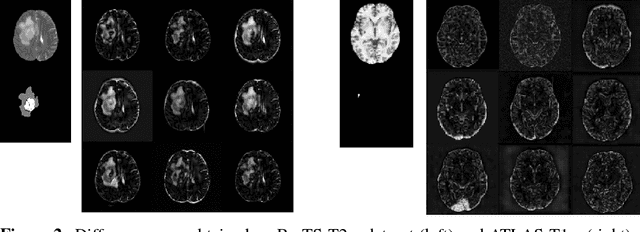
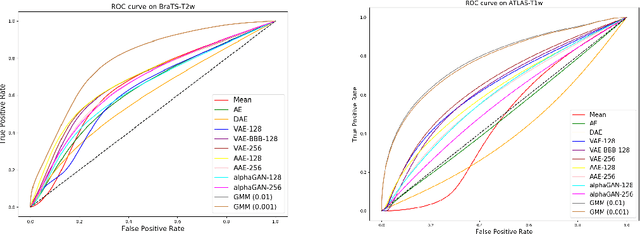
Abstract:Recent advances in deep learning led to novel generative modeling techniques that achieve unprecedented quality in generated samples and performance in learning complex distributions in imaging data. These new models in medical image computing have important applications that form clinically relevant and very challenging unsupervised learning problems. In this paper, we explore the feasibility of using state-of-the-art auto-encoder-based deep generative models, such as variational and adversarial auto-encoders, for one such task: abnormality detection in medical imaging. We utilize typical, publicly available datasets with brain scans from healthy subjects and patients with stroke lesions and brain tumors. We use the data from healthy subjects to train different auto-encoder based models to learn the distribution of healthy images and detect pathologies as outliers. Models that can better learn the data distribution should be able to detect outliers more accurately. We evaluate the detection performance of deep generative models and compare them with non-deep learning based approaches to provide a benchmark of the current state of research. We conclude that abnormality detection is a challenging task for deep generative models and large room exists for improvement. In order to facilitate further research, we aim to provide carefully pre-processed imaging data available to the research community.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge