Naveen Kumar
Machine Learning Fleet Efficiency: Analyzing and Optimizing Large-Scale Google TPU Systems with ML Productivity Goodput
Feb 10, 2025



Abstract:Recent years have seen the emergence of machine learning (ML) workloads deployed in warehouse-scale computing (WSC) settings, also known as ML fleets. As the computational demands placed on ML fleets have increased due to the rise of large models and growing demand for ML applications, it has become increasingly critical to measure and improve the efficiency of such systems. However, there is not yet an established methodology to characterize ML fleet performance and identify potential performance optimizations accordingly. This paper presents a large-scale analysis of an ML fleet based on Google's TPUs, introducing a framework to capture fleet-wide efficiency, systematically evaluate performance characteristics, and identify optimization strategies for the fleet. We begin by defining an ML fleet, outlining its components, and analyzing an example Google ML fleet in production comprising thousands of accelerators running diverse workloads. Our study reveals several critical insights: first, ML fleets extend beyond the hardware layer, with model, data, framework, compiler, and scheduling layers significantly impacting performance; second, the heterogeneous nature of ML fleets poses challenges in characterizing individual workload performance; and third, traditional utilization-based metrics prove insufficient for ML fleet characterization. To address these challenges, we present the "ML Productivity Goodput" (MPG) metric to measure ML fleet efficiency. We show how to leverage this metric to characterize the fleet across the ML system stack. We also present methods to identify and optimize performance bottlenecks using MPG, providing strategies for managing warehouse-scale ML systems in general. Lastly, we demonstrate quantitative evaluations from applying these methods to a real ML fleet for internal-facing Google TPU workloads, where we observed tangible improvements.
Multi-objective Binary Differential Approach with Parameter Tuning for Discovering Business Process Models: MoD-ProM
Jun 25, 2024Abstract:Process discovery approaches analyze the business data to automatically uncover structured information, known as a process model. The quality of a process model is measured using quality dimensions -- completeness (replay fitness), preciseness, simplicity, and generalization. Traditional process discovery algorithms usually output a single process model. A single model may not accurately capture the observed behavior and overfit the training data. We have formed the process discovery problem in a multi-objective framework that yields several candidate solutions for the end user who can pick a suitable model based on the local environmental constraints (possibly varying). We consider the Binary Differential Evolution approach in a multi-objective framework for the task of process discovery. The proposed method employs dichotomous crossover/mutation operators. The parameters are tuned using Grey relational analysis combined with the Taguchi approach. {We have compared the proposed approach with the well-known single-objective algorithms and state-of-the-art multi-objective evolutionary algorithm -- Non-dominated Sorting Genetic Algorithm (NSGA-II).} Additional comparison via computing a weighted average of the quality dimensions is also undertaken. Results show that the proposed algorithm is computationally efficient and produces diversified candidate solutions that score high on the fitness functions. It is shown that the process models generated by the proposed approach are superior to or at least as good as those generated by the state-of-the-art algorithms.
A cross-talk robust multichannel VAD model for multiparty agent interactions trained using synthetic re-recordings
Feb 15, 2024Abstract:In this work, we propose a novel cross-talk rejection framework for a multi-channel multi-talker setup for a live multiparty interactive show. Our far-field audio setup is required to be hands-free during live interaction and comprises four adjacent talkers with directional microphones in the same space. Such setups often introduce heavy cross-talk between channels, resulting in reduced automatic speech recognition (ASR) and natural language understanding (NLU) performance. To address this problem, we propose voice activity detection (VAD) model for all talkers using multichannel information, which is then used to filter audio for downstream tasks. We adopt a synthetic training data generation approach through playback and re-recording for such scenarios, simulating challenging speech overlap conditions. We train our models on this synthetic data and demonstrate that our approach outperforms single-channel VAD models and energy-based multi-channel VAD algorithm in various acoustic environments. In addition to VAD results, we also present multiparty ASR evaluation results to highlight the impact of using our VAD model for filtering audio in downstream tasks by significantly reducing the insertion error.
Biomarker Gene Identification for Breast Cancer Classification
Nov 29, 2021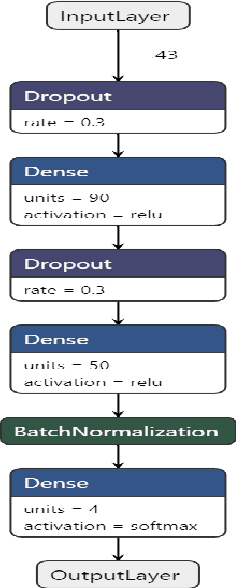
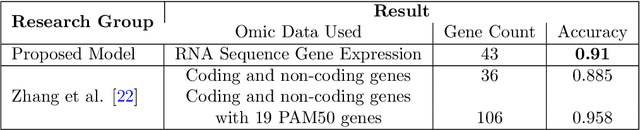

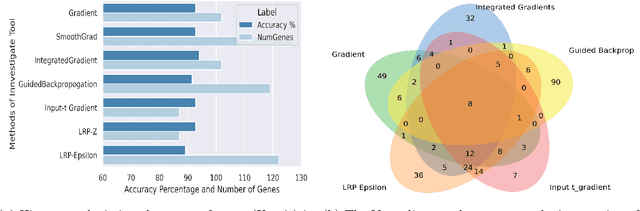
Abstract:BACKGROUND: Breast cancer has emerged as one of the most prevalent cancers among women leading to a high mortality rate. Due to the heterogeneous nature of breast cancer, there is a need to identify differentially expressed genes associated with breast cancer subtypes for its timely diagnosis and treatment. OBJECTIVE: To identify a small gene set for each of the four breast cancer subtypes that could act as its signature, the paper proposes a novel algorithm for gene signature identification. METHODS: The present work uses interpretable AI methods to investigate the predictions made by the deep neural network employed for subtype classification to identify biomarkers using the TCGA breast cancer RNA Sequence data. RESULTS: The proposed algorithm led to the discovery of a set of 43 differentially expressed gene signatures. We achieved a competitive average 10-fold accuracy of 0.91, using neural network classifier. Further, gene set analysis revealed several relevant pathways, such as GRB7 events in ERBB2 and p53 signaling pathway. Using the Pearson correlation matrix, we noted that the subtype-specific genes are correlated within each subtype. CONCLUSIONS: The proposed technique enables us to find a concise and clinically relevant gene signature set.
Deep Learning Based Model for Breast Cancer Subtype Classification
Nov 09, 2021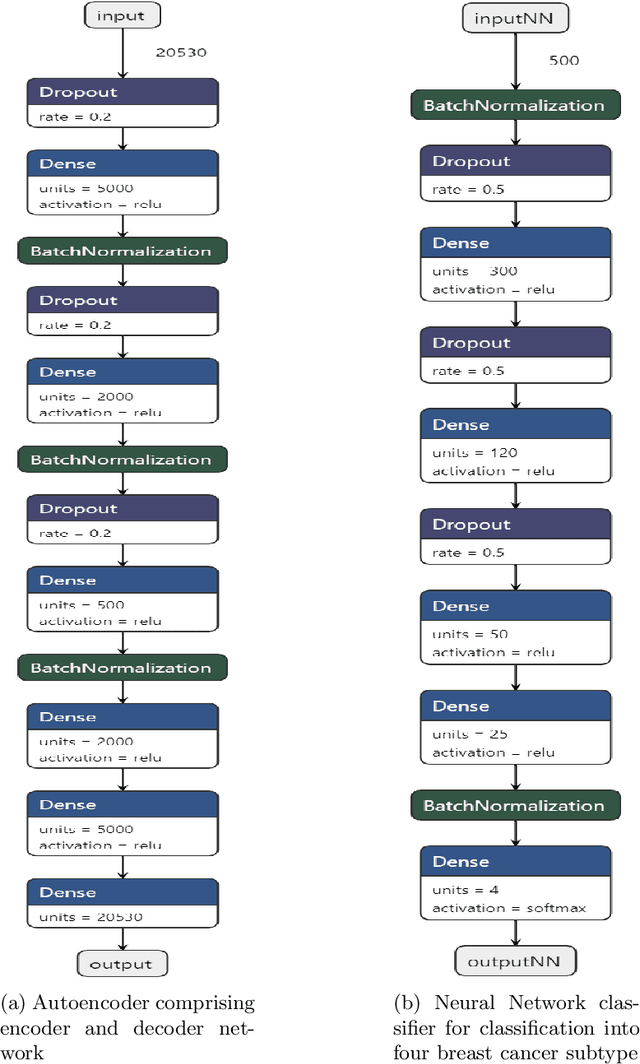
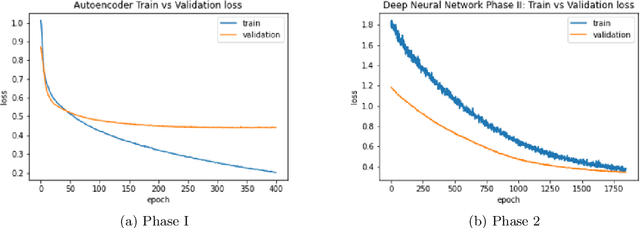
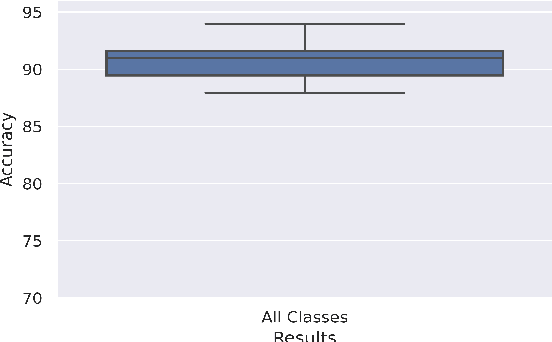
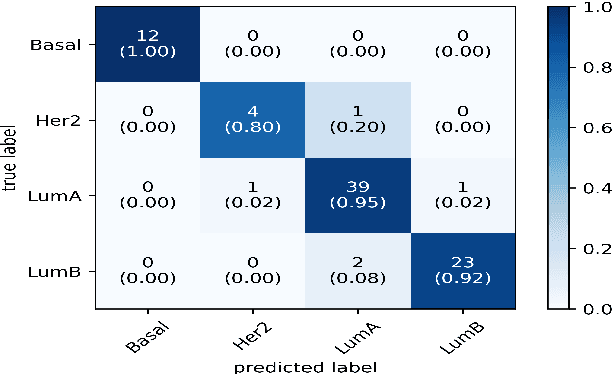
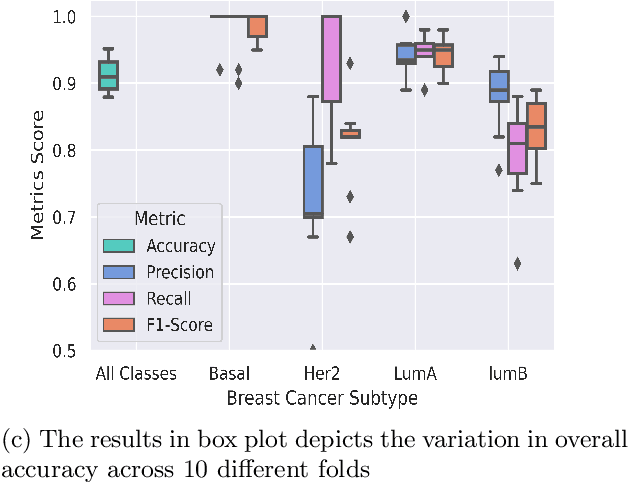
Abstract:Breast cancer has long been a prominent cause of mortality among women. Diagnosis, therapy, and prognosis are now possible, thanks to the availability of RNA sequencing tools capable of recording gene expression data. Molecular subtyping being closely related to devising clinical strategy and prognosis, this paper focuses on the use of gene expression data for the classification of breast cancer into four subtypes, namely, Basal, Her2, LumA, and LumB. In stage 1, we suggested a deep learning-based model that uses an autoencoder to reduce dimensionality. The size of the feature set is reduced from 20,530 gene expression values to 500 by using an autoencoder. This encoded representation is passed to the deep neural network of the second stage for the classification of patients into four molecular subtypes of breast cancer. By deploying the combined network of stages 1 and 2, we have been able to attain a mean 10-fold test accuracy of 0.907 on the TCGA breast cancer dataset. The proposed framework is fairly robust throughout 10 different runs, as shown by the boxplot for classification accuracy. Compared to related work reported in the literature, we have achieved a competitive outcome. In conclusion, the proposed two-stage deep learning-based model is able to accurately classify four breast cancer subtypes, highlighting the autoencoder's capacity to deduce the compact representation and the neural network classifier's ability to correctly label breast cancer patients.
Exploring the limits of Concurrency in ML Training on Google TPUs
Nov 07, 2020
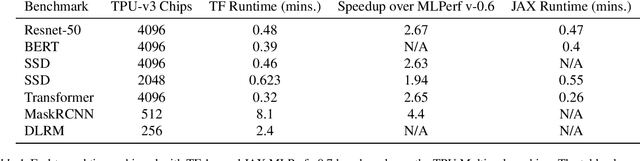
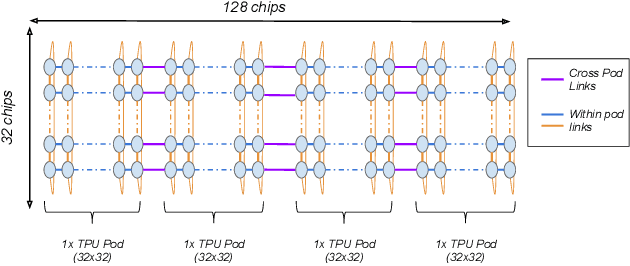
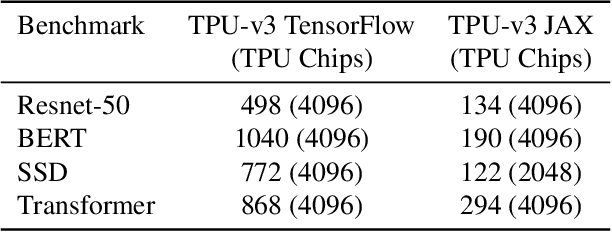
Abstract:Recent results in language understanding using neural networks have required training hardware of unprecedentedscale, with thousands of chips cooperating on a single training run. This paper presents techniques to scaleML models on the Google TPU Multipod, a mesh with 4096 TPU-v3 chips. We discuss model parallelism toovercome scaling limitations from the fixed batch size in data parallelism, communication/collective optimizations,distributed evaluation of training metrics, and host input processing scaling optimizations. These techniques aredemonstrated in both the TensorFlow and JAX programming frameworks. We also present performance resultsfrom the recent Google submission to the MLPerf-v0.7 benchmark contest, achieving record training times from16 to 28 seconds in four MLPerf models on the Google TPU-v3 Multipod machine.
COV-ELM classifier: An Extreme Learning Machine based identification of COVID-19 using Chest X-Ray Images
Aug 15, 2020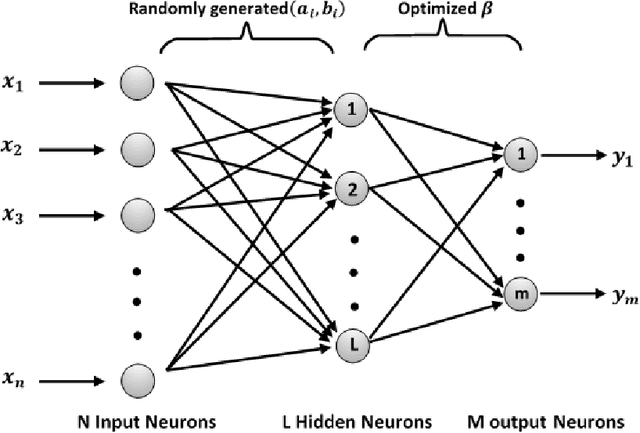
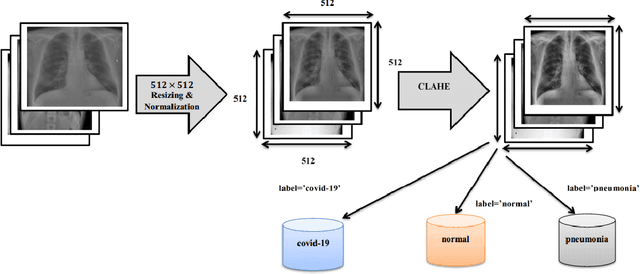
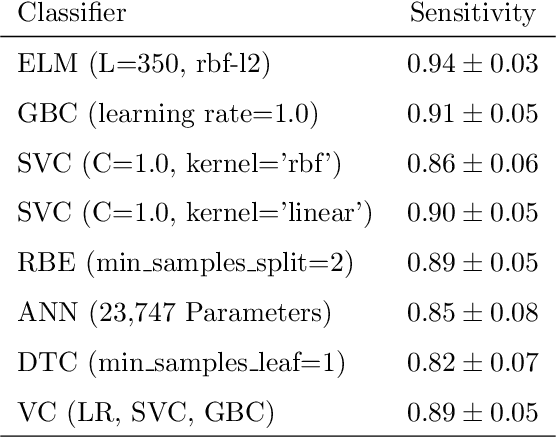
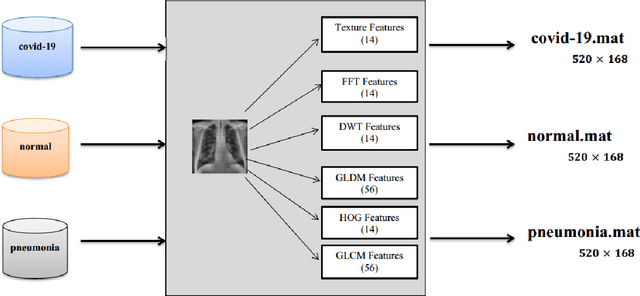
Abstract:Coronaviruses constitute a family of virus that gives rise to respiratory diseases. Coronavirus disease 2019 (COVID-19) is an infectious disease caused by a newly discovered coronavirus also termed as Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Due to its rapid spread, WHO has declared COVID-19 outbreak a pandemic on 11th March 2020. Reverse transcription-polymerase chain reaction (RT-PCR) test is popularly used worldwide for the detection of COVID-19. However, due to the high false-negative rate of RT-PCR test, chest X-ray (CXR) imaging is emerging as a feasible alternative for the detection of COVID-19. In this work, we propose a multiclass classification model COV-ELM, based on the extreme learning machine which classifies the CXR images into one of the three classes, namely COVID-19, normal, and pneumonia. The choice of ELM in this work has been motivated by its significantly short training time as compared to conventional gradient-based learning algorithms. After some preprocessing, we extract a pool of features based on texture and frequency. This pool of features serves as an input to the ELM and a 10-fold cross-validation method is employed to evaluate the proposed model. For experimentation, we use chest X-ray (CXR) images from three publicly available sources. The results of applying COV-ELM on test data are quite promising. The COV-ELM achieved a macro average F1-score of 0.95 and the overall sensitivity of ${0.94 \pm 0.02}$ at 95% confidence interval. When compared to state-of-the-art machine learning algorithms, the COV-ELM is found to outperform in a three-class classification scenario. The main advantage of COV-ELM is that its training time being quite low, as bigger and diverse datasets become available, it can be quickly retrained as compared to its gradient-based competitor models.
Quantized deep learning models on low-power edge devices for robotic systems
Nov 30, 2019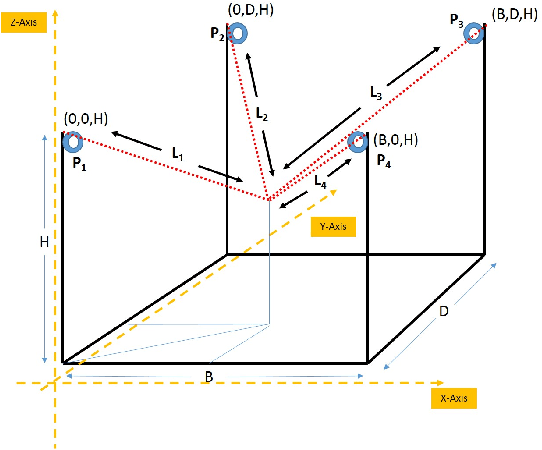
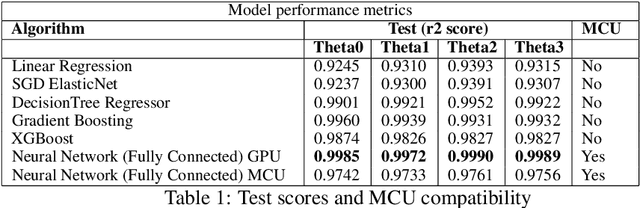
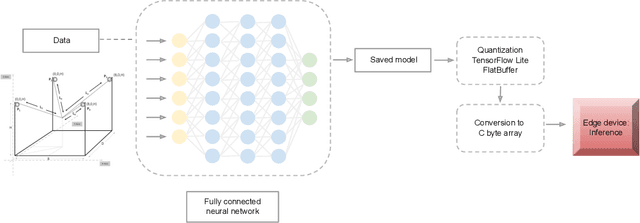
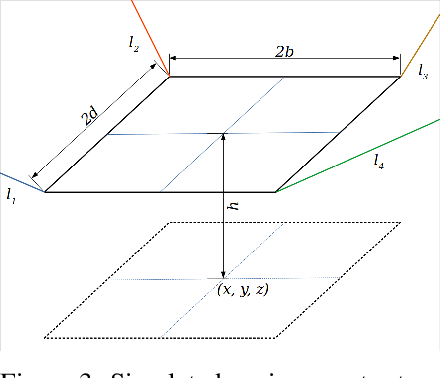
Abstract:In this work, we present a quantized deep neural network deployed on a low-power edge device, inferring learned motor-movements of a suspended robot in a defined space. This serves as the fundamental building block for the original setup, a robotic system for farms or greenhouses aimed at a wide range of agricultural tasks. Deep learning on edge devices and its implications could have a substantial impact on farming systems in the developing world, leading not only to sustainable food production and income, but also increased data privacy and autonomy.
MLPerf Training Benchmark
Oct 30, 2019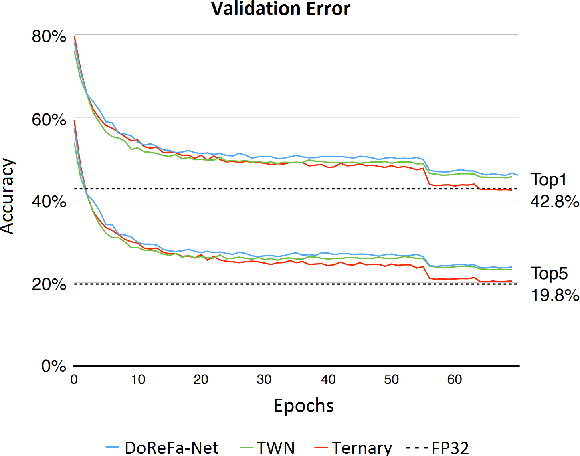
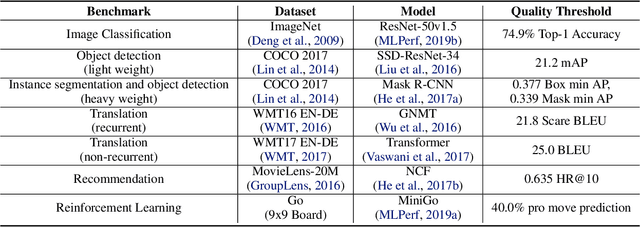
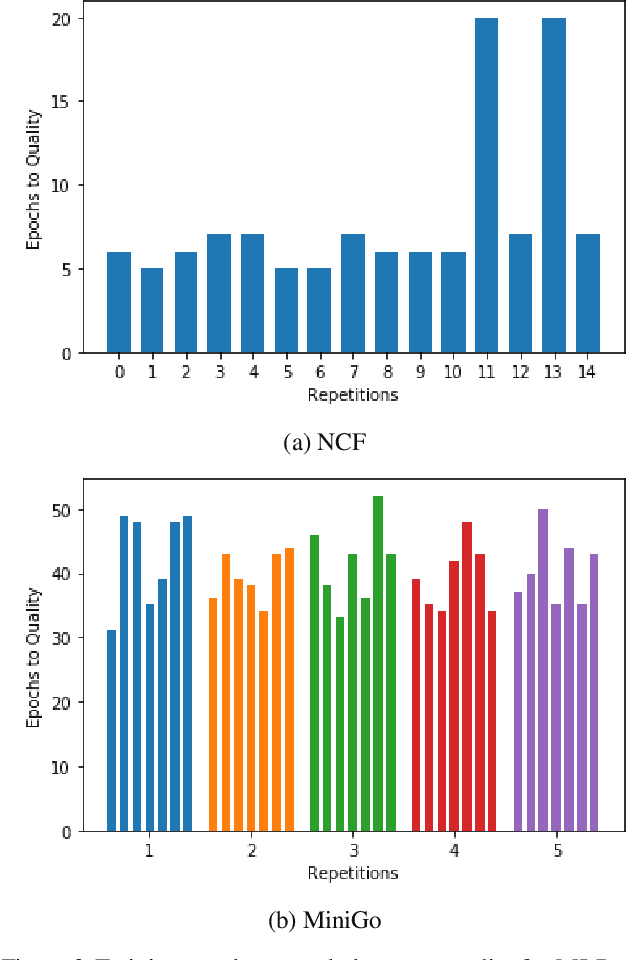
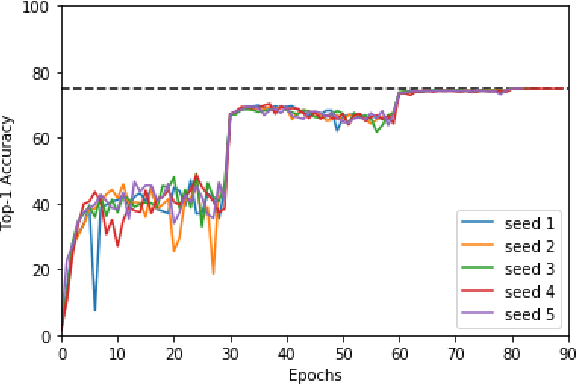
Abstract:Machine learning is experiencing an explosion of software and hardware solutions, and needs industry-standard performance benchmarks to drive design and enable competitive evaluation. However, machine learning training presents a number of unique challenges to benchmarking that do not exist in other domains: (1) some optimizations that improve training throughput actually increase time to solution, (2) training is stochastic and time to solution has high variance, and (3) the software and hardware systems are so diverse that they cannot be fairly benchmarked with the same binary, code, or even hyperparameters. We present MLPerf, a machine learning benchmark that overcomes these challenges. We quantitatively evaluate the efficacy of MLPerf in driving community progress on performance and scalability across two rounds of results from multiple vendors.
Incremental Online Spoken Language Understanding
Oct 23, 2019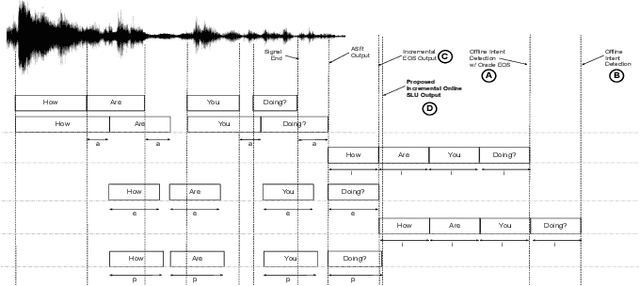
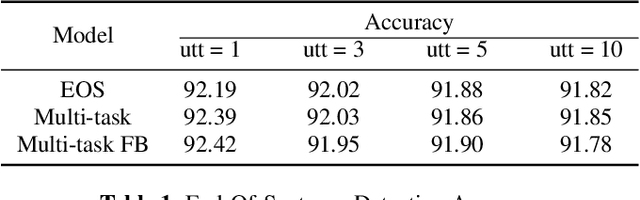
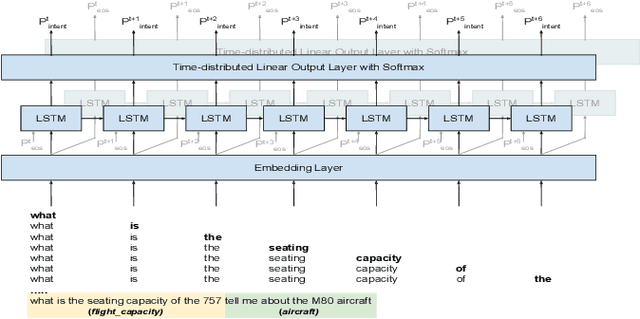
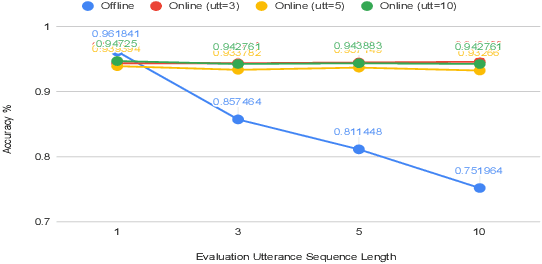
Abstract:Spoken Language Understanding (SLU) typically comprises of an automatic speech recognition (ASR) followed by a natural language understanding (NLU) module. The two modules process signals in a blocking sequential fashion, i.e., the NLU often has to wait for the ASR to finish processing on an utterance basis, potentially leading to high latencies that render the spoken interaction less natural. In this paper, we propose recurrent neural network (RNN) based incremental processing towards the SLU task of intent detection. The proposed methodology offers lower latencies than a typical SLU system, without any significant reduction in system accuracy. We introduce and analyze different recurrent neural network architectures for incremental and online processing of the ASR transcripts and compare it to the existing offline systems. A lexical End-of-Sentence (EOS) detector is proposed for segmenting the stream of transcript into sentences for intent classification. Intent detection experiments are conducted on benchmark ATIS dataset modified to emulate a continuous incremental stream of words with no utterance demarcation. We also analyze the prospects of early intent detection, before EOS, with our proposed system.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge