Nan Wu
Dog nose print matching with dual global descriptor based on Contrastive Learning
Jun 01, 2022

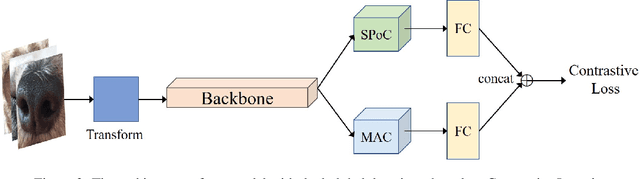

Abstract:Recent studies in biometric-based identification tasks have shown that deep learning methods can achieve better performance. These methods generally extract the global features as descriptor to represent the original image. Nonetheless, it does not perform well for biometric identification under fine-grained tasks. The main reason is that the single image descriptor contains insufficient information to represent image. In this paper, we present a dual global descriptor model, which combines multiple global descriptors to exploit multi level image features. Moreover, we utilize a contrastive loss to enlarge the distance between image representations of confusing classes. The proposed framework achieves the top2 on the CVPR2022 Biometrics Workshop Pet Biometric Challenge. The source code and trained models are publicly available at: https://github.com/flyingsheepbin/pet-biometrics
Neural network topological snake models for locating general phase diagrams
May 19, 2022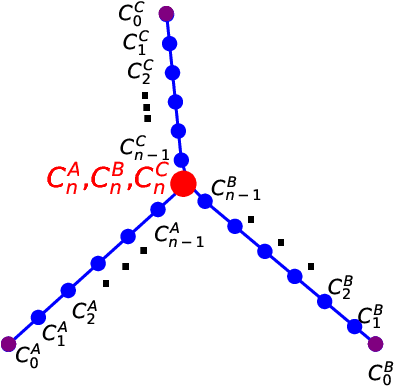
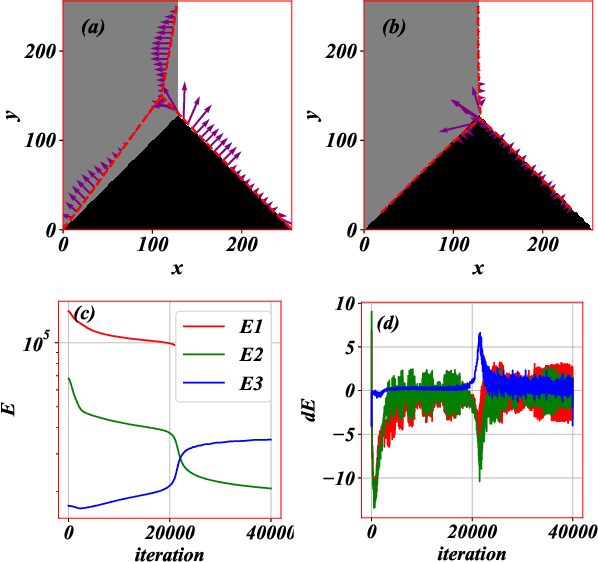
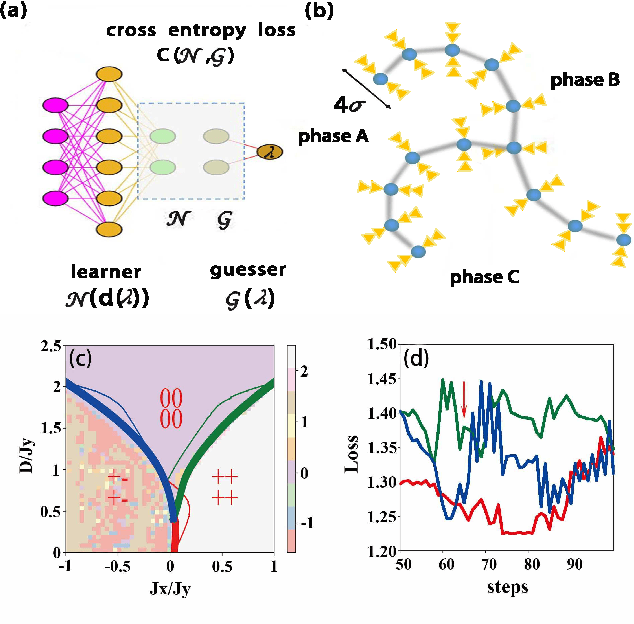
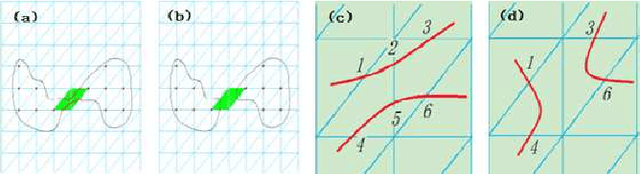
Abstract:Machine learning for locating phase diagram has received intensive research interest in recent years. However, its application in automatically locating phase diagram is limited to single closed phase boundary. In this paper, in order to locate phase diagrams with multiple phases and complex boundaries, we introduce (i) a network-shaped snake model and (ii) a topologically transformable snake with discriminative cooperative networks, respectively. The phase diagrams of both quantum and classical spin-1 model are obtained. Our method is flexible to determine the phase diagram with just snapshots of configurations from the cold-atom or other experiments.
GTNet: A Tree-Based Deep Graph Learning Architecture
Apr 27, 2022
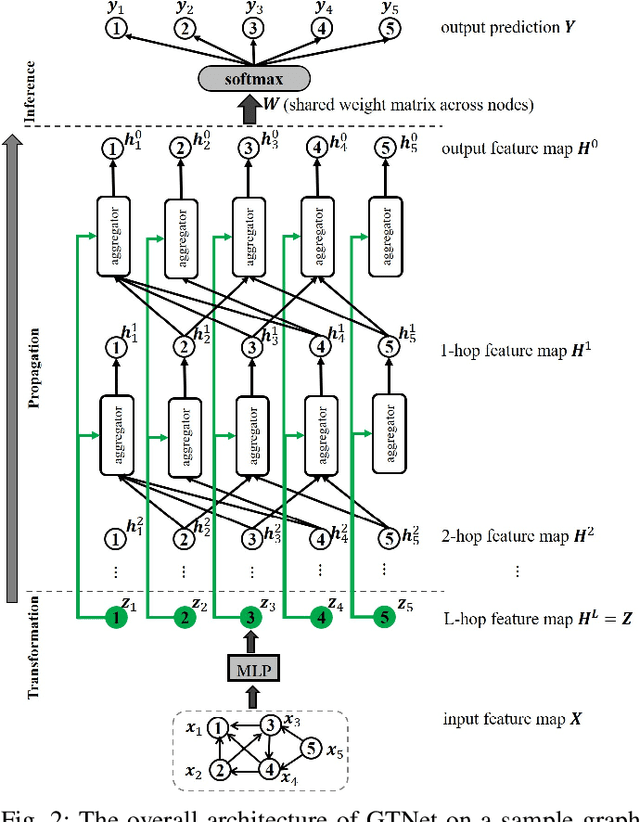
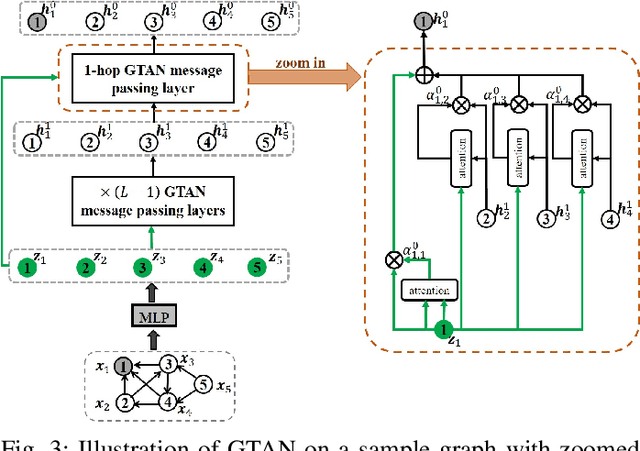
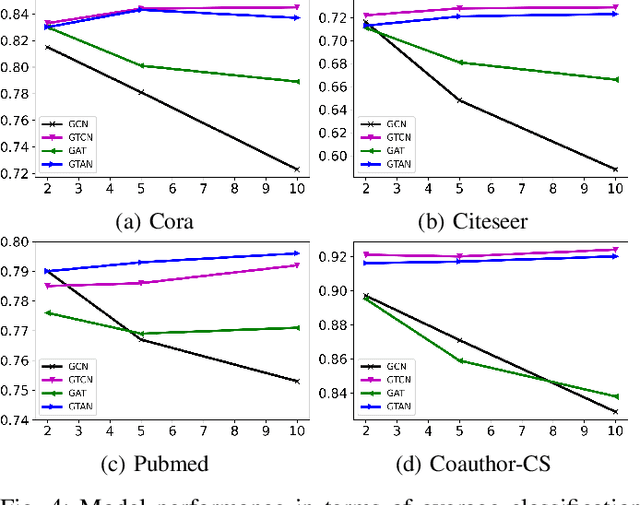
Abstract:We propose Graph Tree Networks (GTNets), a deep graph learning architecture with a new general message passing scheme that originates from the tree representation of graphs. In the tree representation, messages propagate upward from the leaf nodes to the root node, and each node preserves its initial information prior to receiving information from its child nodes (neighbors). We formulate a general propagation rule following the nature of message passing in the tree to update a node's feature by aggregating its initial feature and its neighbor nodes' updated features. Two graph representation learning models are proposed within this GTNet architecture - Graph Tree Attention Network (GTAN) and Graph Tree Convolution Network (GTCN), with experimentally demonstrated state-of-the-art performance on several popular benchmark datasets. Unlike the vanilla Graph Attention Network (GAT) and Graph Convolution Network (GCN) which have the "over-smoothing" issue, the proposed GTAN and GTCN models can go deep as demonstrated by comprehensive experiments and rigorous theoretical analysis.
Characterizing and overcoming the greedy nature of learning in multi-modal deep neural networks
Feb 10, 2022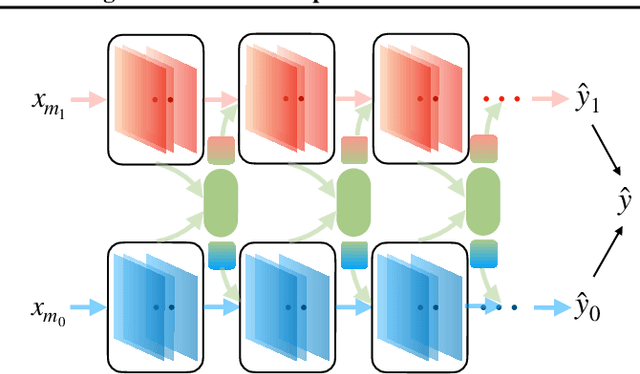
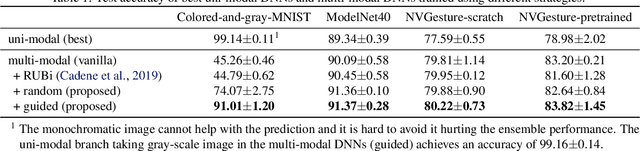

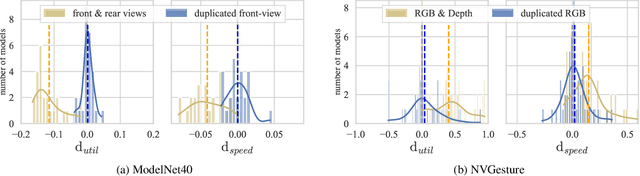
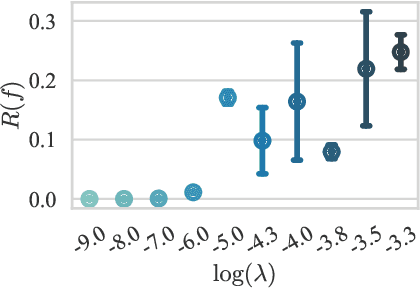
Abstract:We hypothesize that due to the greedy nature of learning in multi-modal deep neural networks, these models tend to rely on just one modality while under-fitting the other modalities. Such behavior is counter-intuitive and hurts the models' generalization, as we observe empirically. To estimate the model's dependence on each modality, we compute the gain on the accuracy when the model has access to it in addition to another modality. We refer to this gain as the conditional utilization rate. In the experiments, we consistently observe an imbalance in conditional utilization rates between modalities, across multiple tasks and architectures. Since conditional utilization rate cannot be computed efficiently during training, we introduce a proxy for it based on the pace at which the model learns from each modality, which we refer to as the conditional learning speed. We propose an algorithm to balance the conditional learning speeds between modalities during training and demonstrate that it indeed addresses the issue of greedy learning. The proposed algorithm improves the model's generalization on three datasets: Colored MNIST, Princeton ModelNet40, and NVIDIA Dynamic Hand Gesture.
Hybrid Graph Models for Logic Optimization via Spatio-Temporal Information
Jan 20, 2022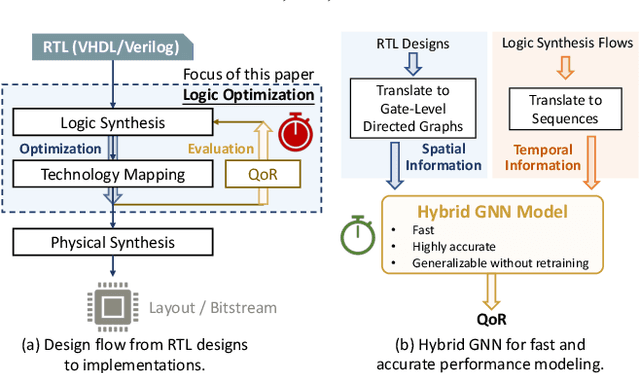
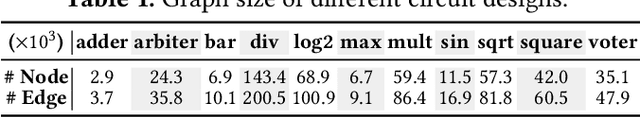

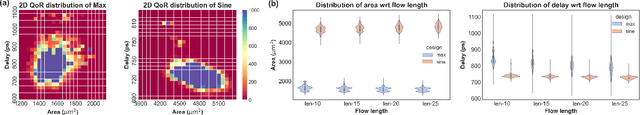

Abstract:Despite the stride made by machine learning (ML) based performance modeling, two major concerns that may impede production-ready ML applications in EDA are stringent accuracy requirements and generalization capability. To this end, we propose hybrid graph neural network (GNN) based approaches towards highly accurate quality-of-result (QoR) estimations with great generalization capability, specifically targeting logic synthesis optimization. The key idea is to simultaneously leverage spatio-temporal information from hardware designs and logic synthesis flows to forecast performance (i.e., delay/area) of various synthesis flows on different designs. The structural characteristics inside hardware designs are distilled and represented by GNNs; the temporal knowledge (i.e., relative ordering of logic transformations) in synthesis flows can be imposed on hardware designs by combining a virtually added supernode or a sequence processing model with conventional GNN models. Evaluation on 3.3 million data points shows that the testing mean absolute percentage error (MAPE) on designs seen and unseen during training are no more than 1.2% and 3.1%, respectively, which are 7-15X lower than existing studies.
High-Level Synthesis Performance Prediction using GNNs: Benchmarking, Modeling, and Advancing
Jan 18, 2022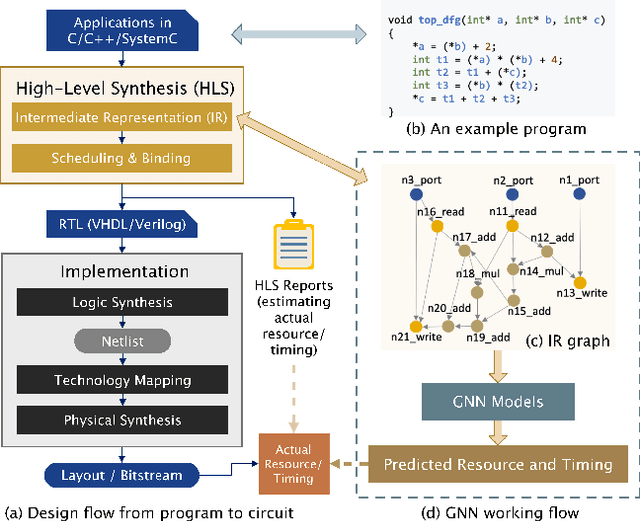
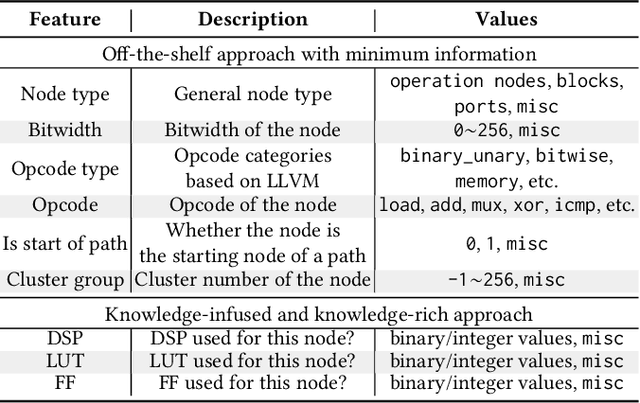
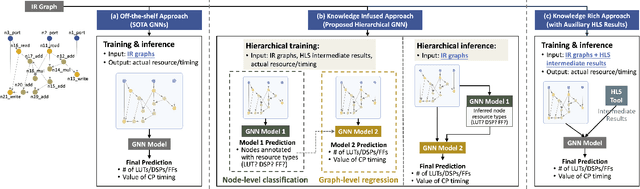
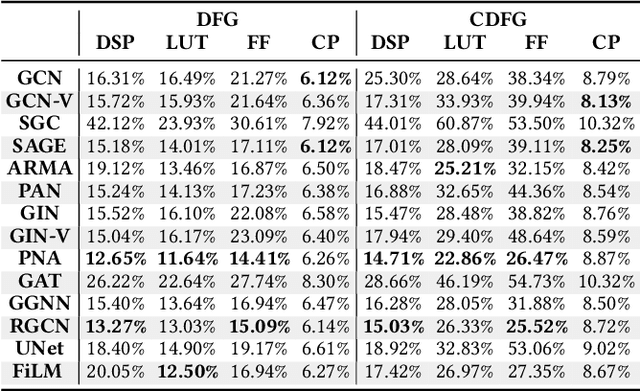
Abstract:Agile hardware development requires fast and accurate circuit quality evaluation from early design stages. Existing work of high-level synthesis (HLS) performance prediction usually needs extensive feature engineering after the synthesis process. To expedite circuit evaluation from as earlier design stage as possible, we propose a rapid and accurate performance modeling, exploiting the representation power of graph neural networks (GNNs) by representing C/C++ programs as graphs. The contribution of this work is three-fold. First, we build a standard benchmark containing 40k C synthesizable programs, which includes both synthetic programs and three sets of real-world HLS benchmarks. Each program is implemented on FPGA to generate ground-truth performance metrics. Second, we formally formulate the HLS performance prediction problem on graphs, and propose multiple modeling strategies with GNNs that leverage different trade-offs between prediction timeliness (early/late prediction) and accuracy. Third, we further propose a novel hierarchical GNN that does not sacrifice timeliness but largely improves prediction accuracy, significantly outperforming HLS tools. We apply extensive evaluations for both synthetic and unseen real-case programs; our proposed predictor largely outperforms HLS by up to 40X and excels existing predictors by 2X to 5X in terms of resource usage and timing prediction.
Realtime 3D Object Detection for Headsets
Jan 15, 2022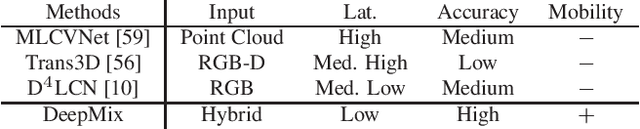
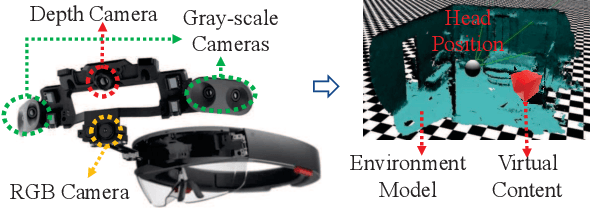
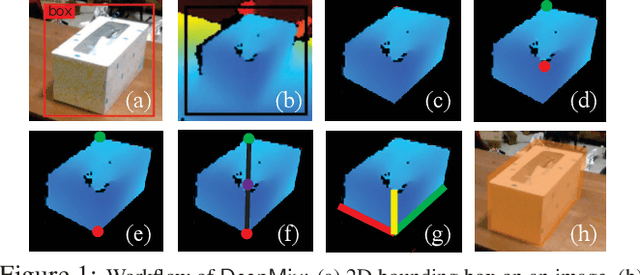

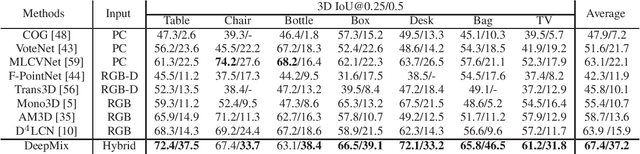
Abstract:Mobile headsets should be capable of understanding 3D physical environments to offer a truly immersive experience for augmented/mixed reality (AR/MR). However, their small form-factor and limited computation resources make it extremely challenging to execute in real-time 3D vision algorithms, which are known to be more compute-intensive than their 2D counterparts. In this paper, we propose DeepMix, a mobility-aware, lightweight, and hybrid3D object detection framework for improving the user experience of AR/MR on mobile headsets. Motivated by our analysis and evaluation of state-of-the-art 3D object detection models, DeepMix intelligently combines edge-assisted 2D object detection and novel, on-device 3D bounding box estimations that leverage depth data captured by headsets. This leads to low end-to-end latency and significantly boosts detection accuracy in mobile scenarios.
SLAM: A Unified Encoder for Speech and Language Modeling via Speech-Text Joint Pre-Training
Oct 20, 2021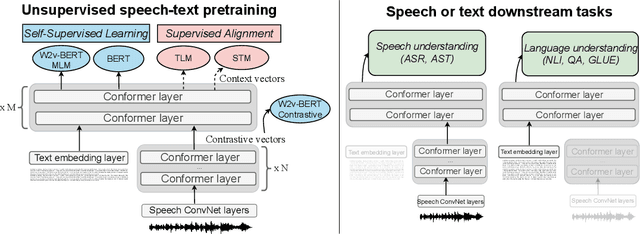
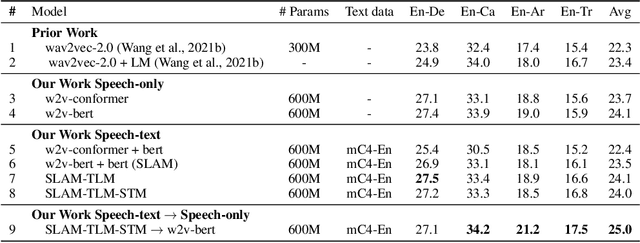
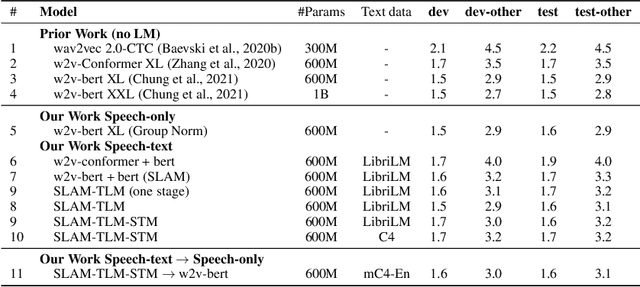
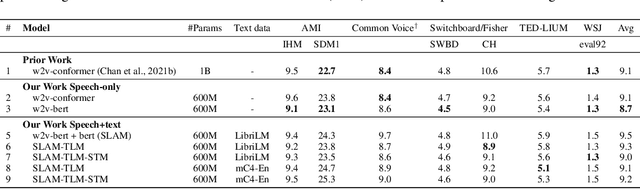
Abstract:Unsupervised pre-training is now the predominant approach for both text and speech understanding. Self-attention models pre-trained on large amounts of unannotated data have been hugely successful when fine-tuned on downstream tasks from a variety of domains and languages. This paper takes the universality of unsupervised language pre-training one step further, by unifying speech and text pre-training within a single model. We build a single encoder with the BERT objective on unlabeled text together with the w2v-BERT objective on unlabeled speech. To further align our model representations across modalities, we leverage alignment losses, specifically Translation Language Modeling (TLM) and Speech Text Matching (STM) that make use of supervised speech-text recognition data. We demonstrate that incorporating both speech and text data during pre-training can significantly improve downstream quality on CoVoST~2 speech translation, by around 1 BLEU compared to single-modality pre-trained models, while retaining close to SotA performance on LibriSpeech and SpeechStew ASR tasks. On four GLUE tasks and text-normalization, we observe evidence of capacity limitations and interference between the two modalities, leading to degraded performance compared to an equivalent text-only model, while still being competitive with BERT. Through extensive empirical analysis we also demonstrate the importance of the choice of objective function for speech pre-training, and the beneficial effect of adding additional supervised signals on the quality of the learned representations.
Cooperative Localization in Massive Networks
Oct 15, 2021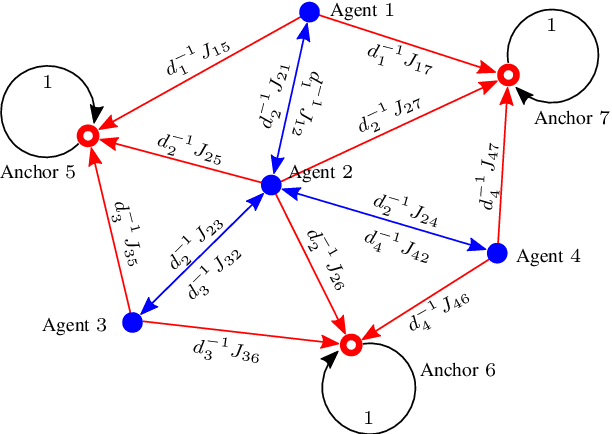
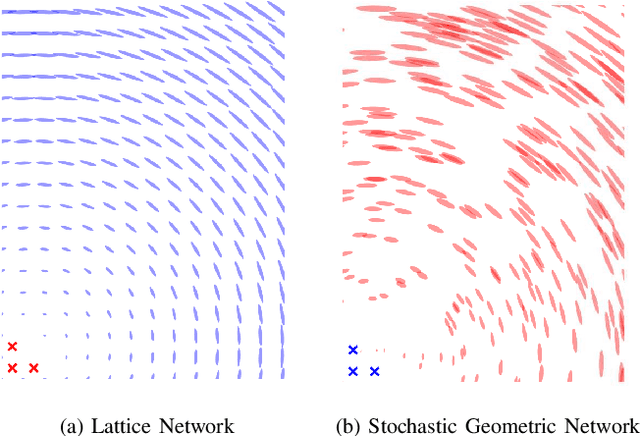
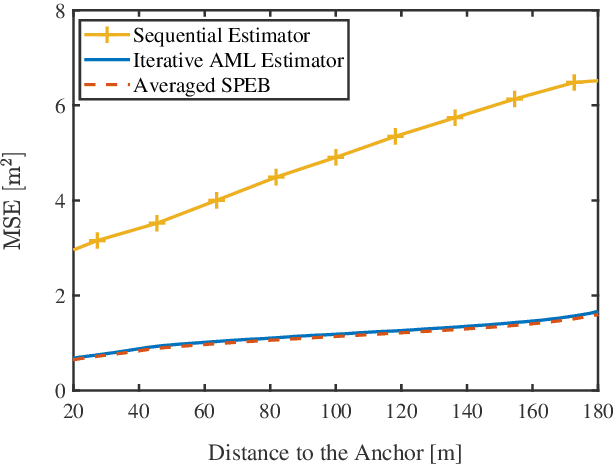
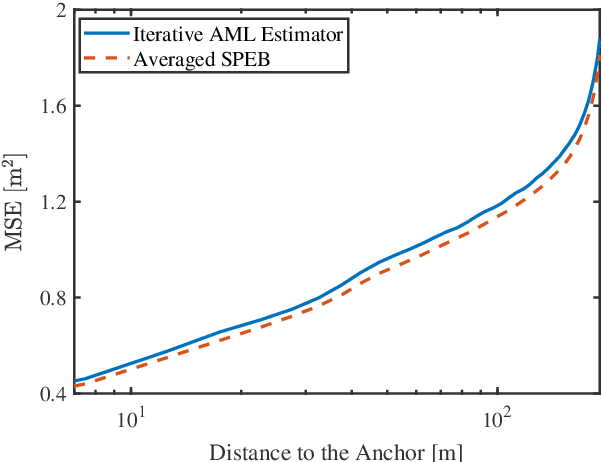
Abstract:Network localization is capable of providing accurate and ubiquitous position information for numerous wireless applications. This paper studies the accuracy of cooperative network localization in large-scale wireless networks. Based on a decomposition of the equivalent Fisher information matrix (EFIM), we develop a random-walk-inspired approach for the analysis of EFIM, and propose a position information routing interpretation of cooperative network localization. Using this approach, we show that in large lattice and stochastic geometric networks, when anchors are uniformly distributed, the average localization error of agents grows logarithmically with the reciprocal of anchor density in an asymptotic regime. The results are further illustrated using numerical examples.
Inferring Manifolds From Noisy Data Using Gaussian Processes
Oct 14, 2021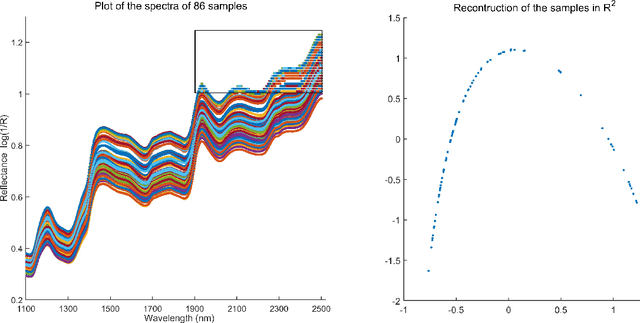
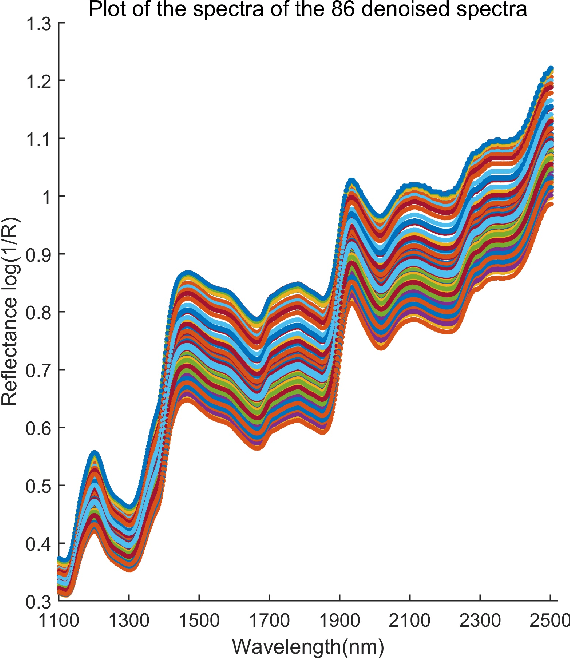
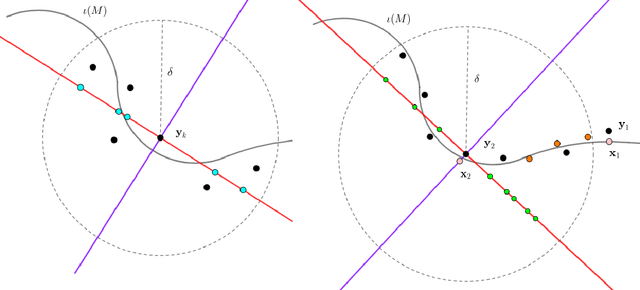
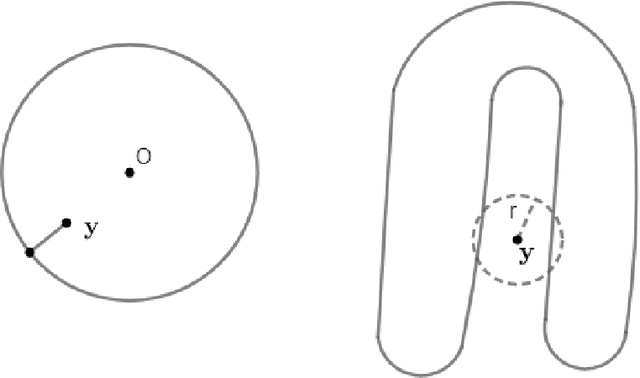
Abstract:In analyzing complex datasets, it is often of interest to infer lower dimensional structure underlying the higher dimensional observations. As a flexible class of nonlinear structures, it is common to focus on Riemannian manifolds. Most existing manifold learning algorithms replace the original data with lower dimensional coordinates without providing an estimate of the manifold in the observation space or using the manifold to denoise the original data. This article proposes a new methodology for addressing these problems, allowing interpolation of the estimated manifold between fitted data points. The proposed approach is motivated by novel theoretical properties of local covariance matrices constructed from noisy samples on a manifold. Our results enable us to turn a global manifold reconstruction problem into a local regression problem, allowing application of Gaussian processes for probabilistic manifold reconstruction. In addition to theory justifying the algorithm, we provide simulated and real data examples to illustrate the performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge