Michael Moor
RadAgent: A tool-using AI agent for stepwise interpretation of chest computed tomography
Apr 16, 2026Abstract:Vision-language models (VLM) have markedly advanced AI-driven interpretation and reporting of complex medical imaging, such as computed tomography (CT). Yet, existing methods largely relegate clinicians to passive observers of final outputs, offering no interpretable reasoning trace for them to inspect, validate, or refine. To address this, we introduce RadAgent, a tool-using AI agent that generates CT reports through a stepwise and interpretable process. Each resulting report is accompanied by a fully inspectable trace of intermediate decisions and tool interactions, allowing clinicians to examine how the reported findings are derived. In our experiments, we observe that RadAgent improves Chest CT report generation over its 3D VLM counterpart, CT-Chat, across three dimensions. Clinical accuracy improves by 6.0 points (36.4% relative) in macro-F1 and 5.4 points (19.6% relative) in micro-F1. Robustness under adversarial conditions improves by 24.7 points (41.9% relative). Furthermore, RadAgent achieves 37.0% in faithfulness, a new capability entirely absent in its 3D VLM counterpart. By structuring the interpretation of chest CT as an explicit, tool-augmented and iterative reasoning trace, RadAgent brings us closer toward transparent and reliable AI for radiology.
Process Reward Agents for Steering Knowledge-Intensive Reasoning
Apr 10, 2026Abstract:Reasoning in knowledge-intensive domains remains challenging as intermediate steps are often not locally verifiable: unlike math or code, evaluating step correctness may require synthesizing clues across large external knowledge sources. As a result, subtle errors can propagate through reasoning traces, potentially never to be detected. Prior work has proposed process reward models (PRMs), including retrieval-augmented variants, but these methods operate post hoc, scoring completed trajectories, which prevents their integration into dynamic inference procedures. Here, we introduce Process Reward Agents (PRA), a test-time method for providing domain-grounded, online, step-wise rewards to a frozen policy. In contrast to prior retrieval-augmented PRMs, PRA enables search-based decoding to rank and prune candidate trajectories at every generation step. Experiments on multiple medical reasoning benchmarks demonstrate that PRA consistently outperforms strong baselines, achieving 80.8% accuracy on MedQA with Qwen3-4B, a new state of the art at the 4B scale. Importantly, PRA generalizes to unseen frozen policy models ranging from 0.5B to 8B parameters, improving their accuracy by up to 25.7% without any policy model updates. More broadly, PRA suggests a paradigm in which frozen reasoners are decoupled from domain-specific reward modules, allowing the deployment of new backbones in complex domains without retraining.
Meta-RL Induces Exploration in Language Agents
Dec 18, 2025Abstract:Reinforcement learning (RL) has enabled the training of large language model (LLM) agents to interact with the environment and to solve multi-turn long-horizon tasks. However, the RL-trained agents often struggle in tasks that require active exploration and fail to efficiently adapt from trial-and-error experiences. In this paper, we present LaMer, a general Meta-RL framework that enables LLM agents to actively explore and learn from the environment feedback at test time. LaMer consists of two key components: (i) a cross-episode training framework to encourage exploration and long-term rewards optimization; and (ii) in-context policy adaptation via reflection, allowing the agent to adapt their policy from task feedback signal without gradient update. Experiments across diverse environments show that LaMer significantly improves performance over RL baselines, with 11%, 14%, and 19% performance gains on Sokoban, MineSweeper and Webshop, respectively. Moreover, LaMer also demonstrates better generalization to more challenging or previously unseen tasks compared to the RL-trained agents. Overall, our results demonstrate that Meta-RL provides a principled approach to induce exploration in language agents, enabling more robust adaptation to novel environments through learned exploration strategies.
Enhancing Radiology Report Generation and Visual Grounding using Reinforcement Learning
Dec 11, 2025Abstract:Recent advances in vision-language models (VLMs) have improved Chest X-ray (CXR) interpretation in multiple aspects. However, many medical VLMs rely solely on supervised fine-tuning (SFT), which optimizes next-token prediction without evaluating answer quality. In contrast, reinforcement learning (RL) can incorporate task-specific feedback, and its combination with explicit intermediate reasoning ("thinking") has demonstrated substantial gains on verifiable math and coding tasks. To investigate the effects of RL and thinking in a CXR VLM, we perform large-scale SFT on CXR data to build an updated RadVLM based on Qwen3-VL, followed by a cold-start SFT stage that equips the model with basic thinking ability. We then apply Group Relative Policy Optimization (GRPO) with clinically grounded, task-specific rewards for report generation and visual grounding, and run matched RL experiments on both domain-specific and general-domain Qwen3-VL variants, with and without thinking. Across these settings, we find that while strong SFT remains crucial for high base performance, RL provides additional gains on both tasks, whereas explicit thinking does not appear to further improve results. Under a unified evaluation pipeline, the RL-optimized RadVLM models outperform their baseline counterparts and reach state-of-the-art performance on both report generation and grounding, highlighting clinically aligned RL as a powerful complement to SFT for medical VLMs.
Agentic Systems in Radiology: Design, Applications, Evaluation, and Challenges
Oct 10, 2025Abstract:Building agents, systems that perceive and act upon their environment with a degree of autonomy, has long been a focus of AI research. This pursuit has recently become vastly more practical with the emergence of large language models (LLMs) capable of using natural language to integrate information, follow instructions, and perform forms of "reasoning" and planning across a wide range of tasks. With its multimodal data streams and orchestrated workflows spanning multiple systems, radiology is uniquely suited to benefit from agents that can adapt to context and automate repetitive yet complex tasks. In radiology, LLMs and their multimodal variants have already demonstrated promising performance for individual tasks such as information extraction and report summarization. However, using LLMs in isolation underutilizes their potential to support complex, multi-step workflows where decisions depend on evolving context from multiple information sources. Equipping LLMs with external tools and feedback mechanisms enables them to drive systems that exhibit a spectrum of autonomy, ranging from semi-automated workflows to more adaptive agents capable of managing complex processes. This review examines the design of such LLM-driven agentic systems, highlights key applications, discusses evaluation methods for planning and tool use, and outlines challenges such as error cascades, tool-use efficiency, and health IT integration.
SMMILE: An Expert-Driven Benchmark for Multimodal Medical In-Context Learning
Jun 26, 2025



Abstract:Multimodal in-context learning (ICL) remains underexplored despite significant potential for domains such as medicine. Clinicians routinely encounter diverse, specialized tasks requiring adaptation from limited examples, such as drawing insights from a few relevant prior cases or considering a constrained set of differential diagnoses. While multimodal large language models (MLLMs) have shown advances in medical visual question answering (VQA), their ability to learn multimodal tasks from context is largely unknown. We introduce SMMILE, the first expert-driven multimodal ICL benchmark for medical tasks. Eleven medical experts curated problems, each including a multimodal query and multimodal in-context examples as task demonstrations. SMMILE encompasses 111 problems (517 question-image-answer triplets) covering 6 medical specialties and 13 imaging modalities. We further introduce SMMILE++, an augmented variant with 1038 permuted problems. A comprehensive evaluation of 15 MLLMs demonstrates that most models exhibit moderate to poor multimodal ICL ability in medical tasks. In open-ended evaluations, ICL contributes only 8% average improvement over zero-shot on SMMILE and 9.4% on SMMILE++. We observe a susceptibility for irrelevant in-context examples: even a single noisy or irrelevant example can degrade performance by up to 9.5%. Moreover, example ordering exhibits a recency bias, i.e., placing the most relevant example last can lead to substantial performance improvements by up to 71%. Our findings highlight critical limitations and biases in current MLLMs when learning multimodal medical tasks from context.
Med-PRM: Medical Reasoning Models with Stepwise, Guideline-verified Process Rewards
Jun 13, 2025Abstract:Large language models have shown promise in clinical decision making, but current approaches struggle to localize and correct errors at specific steps of the reasoning process. This limitation is critical in medicine, where identifying and addressing reasoning errors is essential for accurate diagnosis and effective patient care. We introduce Med-PRM, a process reward modeling framework that leverages retrieval-augmented generation to verify each reasoning step against established medical knowledge bases. By verifying intermediate reasoning steps with evidence retrieved from clinical guidelines and literature, our model can precisely assess the reasoning quality in a fine-grained manner. Evaluations on five medical QA benchmarks and two open-ended diagnostic tasks demonstrate that Med-PRM achieves state-of-the-art performance, with improving the performance of base models by up to 13.50% using Med-PRM. Moreover, we demonstrate the generality of Med-PRM by integrating it in a plug-and-play fashion with strong policy models such as Meerkat, achieving over 80\% accuracy on MedQA for the first time using small-scale models of 8 billion parameters. Our code and data are available at: https://med-prm.github.io/
MIRIAD: Augmenting LLMs with millions of medical query-response pairs
Jun 06, 2025



Abstract:LLMs are bound to transform healthcare with advanced decision support and flexible chat assistants. However, LLMs are prone to generate inaccurate medical content. To ground LLMs in high-quality medical knowledge, LLMs have been equipped with external knowledge via RAG, where unstructured medical knowledge is split into small text chunks that can be selectively retrieved and integrated into the LLMs context. Yet, existing RAG pipelines rely on raw, unstructured medical text, which can be noisy, uncurated and difficult for LLMs to effectively leverage. Systematic approaches to organize medical knowledge to best surface it to LLMs are generally lacking. To address these challenges, we introduce MIRIAD, a large-scale, curated corpus of 5,821,948 medical QA pairs, each rephrased from and grounded in a passage from peer-reviewed medical literature using a semi-automated pipeline combining LLM generation, filtering, grounding, and human annotation. Unlike prior medical corpora, which rely on unstructured text, MIRIAD encapsulates web-scale medical knowledge in an operationalized query-response format, which enables more targeted retrieval. Experiments on challenging medical QA benchmarks show that augmenting LLMs with MIRIAD improves accuracy up to 6.7% compared to unstructured RAG baselines with the same source corpus and with the same amount of retrieved text. Moreover, MIRIAD improved the ability of LLMs to detect medical hallucinations by 22.5 to 37% (increase in F1 score). We further introduce MIRIAD-Atlas, an interactive map of MIRIAD spanning 56 medical disciplines, enabling clinical users to visually explore, search, and refine medical knowledge. MIRIAD promises to unlock a wealth of down-stream applications, including medical information retrievers, enhanced RAG applications, and knowledge-grounded chat interfaces, which ultimately enables more reliable LLM applications in healthcare.
Reverse Image Retrieval Cues Parametric Memory in Multimodal LLMs
May 29, 2024


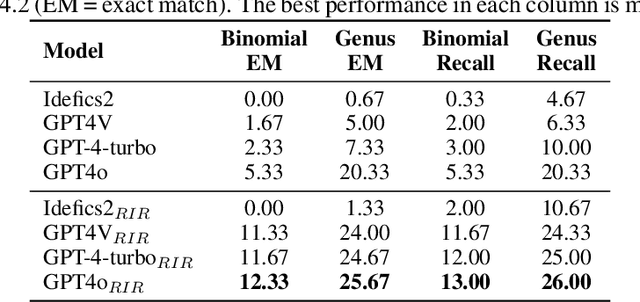
Abstract:Despite impressive advances in recent multimodal large language models (MLLMs), state-of-the-art models such as from the GPT-4 suite still struggle with knowledge-intensive tasks. To address this, we consider Reverse Image Retrieval (RIR) augmented generation, a simple yet effective strategy to augment MLLMs with web-scale reverse image search results. RIR robustly improves knowledge-intensive visual question answering (VQA) of GPT-4V by 37-43%, GPT-4 Turbo by 25-27%, and GPT-4o by 18-20% in terms of open-ended VQA evaluation metrics. To our surprise, we discover that RIR helps the model to better access its own world knowledge. Concretely, our experiments suggest that RIR augmentation helps by providing further visual and textual cues without necessarily containing the direct answer to a query. In addition, we elucidate cases in which RIR can hurt performance and conduct a human evaluation. Finally, we find that the overall advantage of using RIR makes it difficult for an agent that can choose to use RIR to perform better than an approach where RIR is the default setting.
Almanac Copilot: Towards Autonomous Electronic Health Record Navigation
May 14, 2024



Abstract:Clinicians spend large amounts of time on clinical documentation, and inefficiencies impact quality of care and increase clinician burnout. Despite the promise of electronic medical records (EMR), the transition from paper-based records has been negatively associated with clinician wellness, in part due to poor user experience, increased burden of documentation, and alert fatigue. In this study, we present Almanac Copilot, an autonomous agent capable of assisting clinicians with EMR-specific tasks such as information retrieval and order placement. On EHR-QA, a synthetic evaluation dataset of 300 common EHR queries based on real patient data, Almanac Copilot obtains a successful task completion rate of 74% (n = 221 tasks) with a mean score of 2.45 over 3 (95% CI:2.34-2.56). By automating routine tasks and streamlining the documentation process, our findings highlight the significant potential of autonomous agents to mitigate the cognitive load imposed on clinicians by current EMR systems.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge