Cyril Zakka
MIRIAD: Augmenting LLMs with millions of medical query-response pairs
Jun 06, 2025



Abstract:LLMs are bound to transform healthcare with advanced decision support and flexible chat assistants. However, LLMs are prone to generate inaccurate medical content. To ground LLMs in high-quality medical knowledge, LLMs have been equipped with external knowledge via RAG, where unstructured medical knowledge is split into small text chunks that can be selectively retrieved and integrated into the LLMs context. Yet, existing RAG pipelines rely on raw, unstructured medical text, which can be noisy, uncurated and difficult for LLMs to effectively leverage. Systematic approaches to organize medical knowledge to best surface it to LLMs are generally lacking. To address these challenges, we introduce MIRIAD, a large-scale, curated corpus of 5,821,948 medical QA pairs, each rephrased from and grounded in a passage from peer-reviewed medical literature using a semi-automated pipeline combining LLM generation, filtering, grounding, and human annotation. Unlike prior medical corpora, which rely on unstructured text, MIRIAD encapsulates web-scale medical knowledge in an operationalized query-response format, which enables more targeted retrieval. Experiments on challenging medical QA benchmarks show that augmenting LLMs with MIRIAD improves accuracy up to 6.7% compared to unstructured RAG baselines with the same source corpus and with the same amount of retrieved text. Moreover, MIRIAD improved the ability of LLMs to detect medical hallucinations by 22.5 to 37% (increase in F1 score). We further introduce MIRIAD-Atlas, an interactive map of MIRIAD spanning 56 medical disciplines, enabling clinical users to visually explore, search, and refine medical knowledge. MIRIAD promises to unlock a wealth of down-stream applications, including medical information retrievers, enhanced RAG applications, and knowledge-grounded chat interfaces, which ultimately enables more reliable LLM applications in healthcare.
SmolVLM: Redefining small and efficient multimodal models
Apr 07, 2025



Abstract:Large Vision-Language Models (VLMs) deliver exceptional performance but require significant computational resources, limiting their deployment on mobile and edge devices. Smaller VLMs typically mirror design choices of larger models, such as extensive image tokenization, leading to inefficient GPU memory usage and constrained practicality for on-device applications. We introduce SmolVLM, a series of compact multimodal models specifically engineered for resource-efficient inference. We systematically explore architectural configurations, tokenization strategies, and data curation optimized for low computational overhead. Through this, we identify key design choices that yield substantial performance gains on image and video tasks with minimal memory footprints. Our smallest model, SmolVLM-256M, uses less than 1GB GPU memory during inference and outperforms the 300-times larger Idefics-80B model, despite an 18-month development gap. Our largest model, at 2.2B parameters, rivals state-of-the-art VLMs consuming twice the GPU memory. SmolVLM models extend beyond static images, demonstrating robust video comprehension capabilities. Our results emphasize that strategic architectural optimizations, aggressive yet efficient tokenization, and carefully curated training data significantly enhance multimodal performance, facilitating practical, energy-efficient deployments at significantly smaller scales.
SmolLM2: When Smol Goes Big -- Data-Centric Training of a Small Language Model
Feb 04, 2025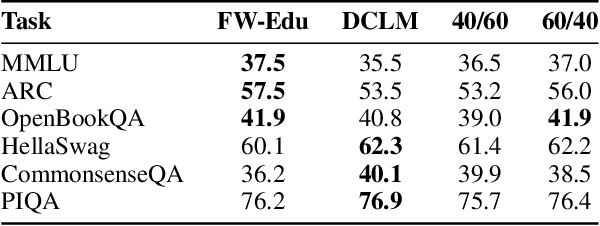

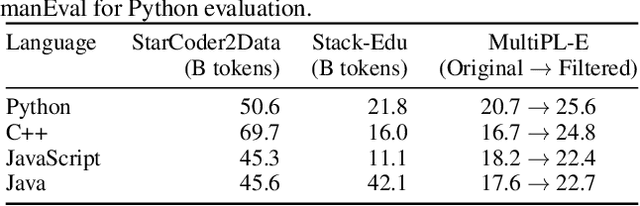

Abstract:While large language models have facilitated breakthroughs in many applications of artificial intelligence, their inherent largeness makes them computationally expensive and challenging to deploy in resource-constrained settings. In this paper, we document the development of SmolLM2, a state-of-the-art "small" (1.7 billion parameter) language model (LM). To attain strong performance, we overtrain SmolLM2 on ~11 trillion tokens of data using a multi-stage training process that mixes web text with specialized math, code, and instruction-following data. We additionally introduce new specialized datasets (FineMath, Stack-Edu, and SmolTalk) at stages where we found existing datasets to be problematically small or low-quality. To inform our design decisions, we perform both small-scale ablations as well as a manual refinement process that updates the dataset mixing rates at each stage based on the performance at the previous stage. Ultimately, we demonstrate that SmolLM2 outperforms other recent small LMs including Qwen2.5-1.5B and Llama3.2-1B. To facilitate future research on LM development as well as applications of small LMs, we release both SmolLM2 as well as all of the datasets we prepared in the course of this project.
Best Practices for Large Language Models in Radiology
Dec 02, 2024



Abstract:At the heart of radiological practice is the challenge of integrating complex imaging data with clinical information to produce actionable insights. Nuanced application of language is key for various activities, including managing requests, describing and interpreting imaging findings in the context of clinical data, and concisely documenting and communicating the outcomes. The emergence of large language models (LLMs) offers an opportunity to improve the management and interpretation of the vast data in radiology. Despite being primarily general-purpose, these advanced computational models demonstrate impressive capabilities in specialized language-related tasks, even without specific training. Unlocking the potential of LLMs for radiology requires basic understanding of their foundations and a strategic approach to navigate their idiosyncrasies. This review, drawing from practical radiology and machine learning expertise and recent literature, provides readers insight into the potential of LLMs in radiology. It examines best practices that have so far stood the test of time in the rapidly evolving landscape of LLMs. This includes practical advice for optimizing LLM characteristics for radiology practices along with limitations, effective prompting, and fine-tuning strategies.
SurGen: Text-Guided Diffusion Model for Surgical Video Generation
Aug 26, 2024



Abstract:Diffusion-based video generation models have made significant strides, producing outputs with improved visual fidelity, temporal coherence, and user control. These advancements hold great promise for improving surgical education by enabling more realistic, diverse, and interactive simulation environments. In this study, we introduce SurGen, a text-guided diffusion model tailored for surgical video synthesis, producing the highest resolution and longest duration videos among existing surgical video generation models. We validate the visual and temporal quality of the outputs using standard image and video generation metrics. Additionally, we assess their alignment to the corresponding text prompts through a deep learning classifier trained on surgical data. Our results demonstrate the potential of diffusion models to serve as valuable educational tools for surgical trainees.
GP-VLS: A general-purpose vision language model for surgery
Jul 27, 2024Abstract:Surgery requires comprehensive medical knowledge, visual assessment skills, and procedural expertise. While recent surgical AI models have focused on solving task-specific problems, there is a need for general-purpose systems that can understand surgical scenes and interact through natural language. This paper introduces GP-VLS, a general-purpose vision language model for surgery that integrates medical and surgical knowledge with visual scene understanding. For comprehensively evaluating general-purpose surgical models, we propose SurgiQual, which evaluates across medical and surgical knowledge benchmarks as well as surgical vision-language questions. To train GP-VLS, we develop six new datasets spanning medical knowledge, surgical textbooks, and vision-language pairs for tasks like phase recognition and tool identification. We show that GP-VLS significantly outperforms existing open- and closed-source models on surgical vision-language tasks, with 8-21% improvements in accuracy across SurgiQual benchmarks. GP-VLS also demonstrates strong performance on medical and surgical knowledge tests compared to open-source alternatives. Overall, GP-VLS provides an open-source foundation for developing AI assistants to support surgeons across a wide range of tasks and scenarios.
MediSyn: Text-Guided Diffusion Models for Broad Medical 2D and 3D Image Synthesis
May 16, 2024Abstract:Diffusion models have recently gained significant traction due to their ability to generate high-fidelity and diverse images and videos conditioned on text prompts. In medicine, this application promises to address the critical challenge of data scarcity, a consequence of barriers in data sharing, stringent patient privacy regulations, and disparities in patient population and demographics. By generating realistic and varying medical 2D and 3D images, these models offer a rich, privacy-respecting resource for algorithmic training and research. To this end, we introduce MediSyn, a pair of instruction-tuned text-guided latent diffusion models with the ability to generate high-fidelity and diverse medical 2D and 3D images across specialties and modalities. Through established metrics, we show significant improvement in broad medical image and video synthesis guided by text prompts.
Almanac Copilot: Towards Autonomous Electronic Health Record Navigation
May 14, 2024



Abstract:Clinicians spend large amounts of time on clinical documentation, and inefficiencies impact quality of care and increase clinician burnout. Despite the promise of electronic medical records (EMR), the transition from paper-based records has been negatively associated with clinician wellness, in part due to poor user experience, increased burden of documentation, and alert fatigue. In this study, we present Almanac Copilot, an autonomous agent capable of assisting clinicians with EMR-specific tasks such as information retrieval and order placement. On EHR-QA, a synthetic evaluation dataset of 300 common EHR queries based on real patient data, Almanac Copilot obtains a successful task completion rate of 74% (n = 221 tasks) with a mean score of 2.45 over 3 (95% CI:2.34-2.56). By automating routine tasks and streamlining the documentation process, our findings highlight the significant potential of autonomous agents to mitigate the cognitive load imposed on clinicians by current EMR systems.
A Generalizable Deep Learning System for Cardiac MRI
Dec 01, 2023



Abstract:Cardiac MRI allows for a comprehensive assessment of myocardial structure, function, and tissue characteristics. Here we describe a foundational vision system for cardiac MRI, capable of representing the breadth of human cardiovascular disease and health. Our deep learning model is trained via self-supervised contrastive learning, by which visual concepts in cine-sequence cardiac MRI scans are learned from the raw text of the accompanying radiology reports. We train and evaluate our model on data from four large academic clinical institutions in the United States. We additionally showcase the performance of our models on the UK BioBank, and two additional publicly available external datasets. We explore emergent zero-shot capabilities of our system, and demonstrate remarkable performance across a range of tasks; including the problem of left ventricular ejection fraction regression, and the diagnosis of 35 different conditions such as cardiac amyloidosis and hypertrophic cardiomyopathy. We show that our deep learning system is capable of not only understanding the staggering complexity of human cardiovascular disease, but can be directed towards clinical problems of interest yielding impressive, clinical grade diagnostic accuracy with a fraction of the training data typically required for such tasks.
Echocardiogram Foundation Model -- Application 1: Estimating Ejection Fraction
Nov 21, 2023



Abstract:Cardiovascular diseases stand as the primary global cause of mortality. Among the various imaging techniques available for visualising the heart and evaluating its function, echocardiograms emerge as the preferred choice due to their safety and low cost. Quantifying cardiac function based on echocardiograms is very laborious, time-consuming and subject to high interoperator variability. In this work, we introduce EchoAI, an echocardiogram foundation model, that is trained using self-supervised learning (SSL) on 1.5 million echocardiograms. We evaluate our approach by fine-tuning EchoAI to estimate the ejection fraction achieving a mean absolute percentage error of 9.40%. This level of accuracy aligns with the performance of expert sonographers.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge