Agustina Saenz
ReXamine-Global: A Framework for Uncovering Inconsistencies in Radiology Report Generation Metrics
Aug 29, 2024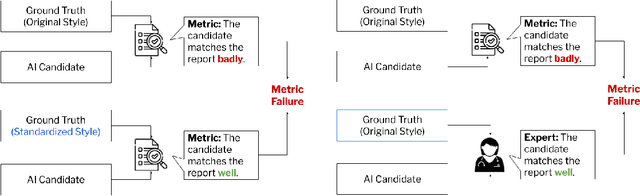



Abstract:Given the rapidly expanding capabilities of generative AI models for radiology, there is a need for robust metrics that can accurately measure the quality of AI-generated radiology reports across diverse hospitals. We develop ReXamine-Global, a LLM-powered, multi-site framework that tests metrics across different writing styles and patient populations, exposing gaps in their generalization. First, our method tests whether a metric is undesirably sensitive to reporting style, providing different scores depending on whether AI-generated reports are stylistically similar to ground-truth reports or not. Second, our method measures whether a metric reliably agrees with experts, or whether metric and expert scores of AI-generated report quality diverge for some sites. Using 240 reports from 6 hospitals around the world, we apply ReXamine-Global to 7 established report evaluation metrics and uncover serious gaps in their generalizability. Developers can apply ReXamine-Global when designing new report evaluation metrics, ensuring their robustness across sites. Additionally, our analysis of existing metrics can guide users of those metrics towards evaluation procedures that work reliably at their sites of interest.
Style-Aware Radiology Report Generation with RadGraph and Few-Shot Prompting
Oct 31, 2023



Abstract:Automatically generated reports from medical images promise to improve the workflow of radiologists. Existing methods consider an image-to-report modeling task by directly generating a fully-fledged report from an image. However, this conflates the content of the report (e.g., findings and their attributes) with its style (e.g., format and choice of words), which can lead to clinically inaccurate reports. To address this, we propose a two-step approach for radiology report generation. First, we extract the content from an image; then, we verbalize the extracted content into a report that matches the style of a specific radiologist. For this, we leverage RadGraph -- a graph representation of reports -- together with large language models (LLMs). In our quantitative evaluations, we find that our approach leads to beneficial performance. Our human evaluation with clinical raters highlights that the AI-generated reports are indistinguishably tailored to the style of individual radiologist despite leveraging only a few examples as context.
RadGraph2: Modeling Disease Progression in Radiology Reports via Hierarchical Information Extraction
Aug 09, 2023



Abstract:We present RadGraph2, a novel dataset for extracting information from radiology reports that focuses on capturing changes in disease state and device placement over time. We introduce a hierarchical schema that organizes entities based on their relationships and show that using this hierarchy during training improves the performance of an information extraction model. Specifically, we propose a modification to the DyGIE++ framework, resulting in our model HGIE, which outperforms previous models in entity and relation extraction tasks. We demonstrate that RadGraph2 enables models to capture a wider variety of findings and perform better at relation extraction compared to those trained on the original RadGraph dataset. Our work provides the foundation for developing automated systems that can track disease progression over time and develop information extraction models that leverage the natural hierarchy of labels in the medical domain.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge