Qian Huang
Toward Real-World Adoption of Portrait Relighting via Hybrid Domain Knowledge Fusion
Apr 25, 2026Abstract:The real-world adoption of portrait relighting is hindered by dataset domain gaps, camera sensitivity, and computational costs. We address these challenges with Hybrid Domain Knowledge Fusion, a paradigm that fuses the specialized strengths of synthetic, One-Light-at-A-Time (OLAT), and real-world datasets into a compact model. Our approach features specialized prior models hardened by domain-aware adaptation, followed by augmented knowledge distillation into a lightweight student model with multi-domain expertise. Our method demonstrates a 6x to 240x inference speedup while maintaining state-of-the-art (SOTA) visual quality in the experiments. Additionally, we construct a massive, high-fidelity synthetic dataset with diverse ground-truth intrinsics to support our training pipeline.
The Trilingual Triad Framework: Integrating Design, AI, and Domain Knowledge in No-code AI Smart City Course
Mar 05, 2026Abstract:This paper introduces the "Trilingual Triad" framework, a model that explains how students learn to design with generative artificial intelligence (AI) through the integration of Design, AI, and Domain Knowledge. As generative AI rapidly enters higher education, students often engage with these systems as passive users of generated outputs rather than active creators of AI-enabled knowledge tools. This study investigates how students can transition from using AI as a tool to designing AI as a collaborative teammate. The research examines a graduate course, Creating the Frontier of No-code Smart Cities at the Singapore University of Technology and Design (SUTD), in which students developed domain-specific custom GPT systems without coding. Using a qualitative multi-case study approach, three projects - the Interview Companion GPT, the Urban Observer GPT, and Buddy Buddy - were analyzed across three dimensions: design, AI architecture, and domain expertise. The findings show that effective human-AI collaboration emerges when these three "languages" are orchestrated together: domain knowledge structures the AI's logic, design mediates human-AI interaction, and AI extends learners' cognitive capacity. The Trilingual Triad framework highlights how building AI systems can serve as a constructionist learning process that strengthens AI literacy, metacognition, and learner agency.
Making medical vision-language models think causally across modalities with retrieval-augmented cross-modal reasoning
Jan 26, 2026Abstract:Medical vision-language models (VLMs) achieve strong performance in diagnostic reporting and image-text alignment, yet their underlying reasoning mechanisms remain fundamentally correlational, exhibiting reliance on superficial statistical associations that fail to capture the causal pathophysiological mechanisms central to clinical decision-making. This limitation makes them fragile, prone to hallucinations, and sensitive to dataset biases. Retrieval-augmented generation (RAG) offers a partial remedy by grounding predictions in external knowledge. However, conventional RAG depends on semantic similarity, introducing new spurious correlations. We propose Multimodal Causal Retrieval-Augmented Generation, a framework that integrates causal inference principles with multimodal retrieval. It retrieves clinically relevant exemplars and causal graphs from external sources, conditioning model reasoning on counterfactual and interventional evidence rather than correlations alone. Applied to radiology report generation, diagnosis prediction, and visual question answering, it improves factual accuracy, robustness to distribution shifts, and interpretability. Our results highlight causal retrieval as a scalable path toward medical VLMs that think beyond pattern matching, enabling trustworthy multimodal reasoning in high-stakes clinical settings.
FinMMDocR: Benchmarking Financial Multimodal Reasoning with Scenario Awareness, Document Understanding, and Multi-Step Computation
Dec 31, 2025Abstract:We introduce FinMMDocR, a novel bilingual multimodal benchmark for evaluating multimodal large language models (MLLMs) on real-world financial numerical reasoning. Compared to existing benchmarks, our work delivers three major advancements. (1) Scenario Awareness: 57.9% of 1,200 expert-annotated problems incorporate 12 types of implicit financial scenarios (e.g., Portfolio Management), challenging models to perform expert-level reasoning based on assumptions; (2) Document Understanding: 837 Chinese/English documents spanning 9 types (e.g., Company Research) average 50.8 pages with rich visual elements, significantly surpassing existing benchmarks in both breadth and depth of financial documents; (3) Multi-Step Computation: Problems demand 11-step reasoning on average (5.3 extraction + 5.7 calculation steps), with 65.0% requiring cross-page evidence (2.4 pages average). The best-performing MLLM achieves only 58.0% accuracy, and different retrieval-augmented generation (RAG) methods show significant performance variations on this task. We expect FinMMDocR to drive improvements in MLLMs and reasoning-enhanced methods on complex multimodal reasoning tasks in real-world scenarios.
Language-Enhanced Mobile Manipulation for Efficient Object Search in Indoor Environments
Aug 28, 2025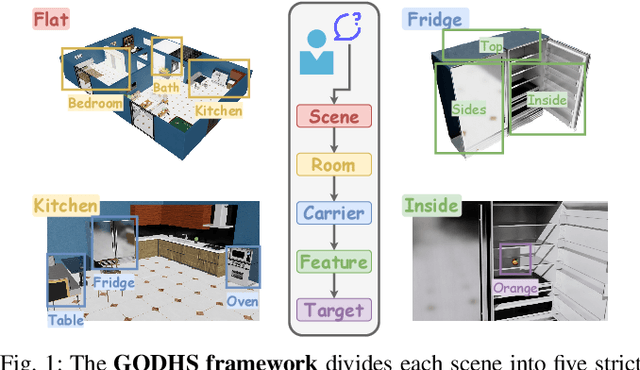
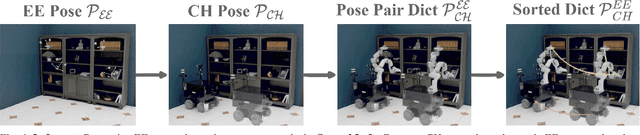
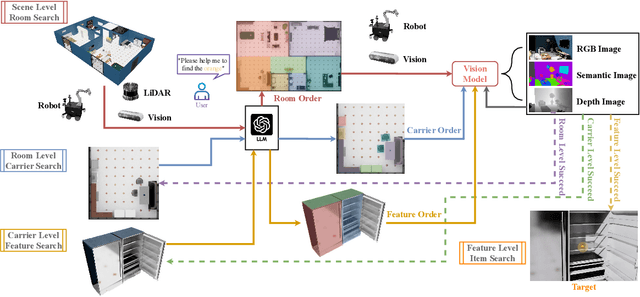
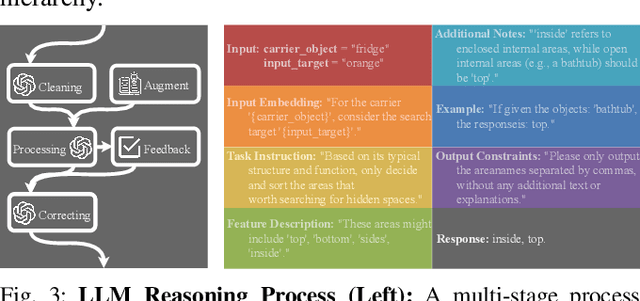

Abstract:Enabling robots to efficiently search for and identify objects in complex, unstructured environments is critical for diverse applications ranging from household assistance to industrial automation. However, traditional scene representations typically capture only static semantics and lack interpretable contextual reasoning, limiting their ability to guide object search in completely unfamiliar settings. To address this challenge, we propose a language-enhanced hierarchical navigation framework that tightly integrates semantic perception and spatial reasoning. Our method, Goal-Oriented Dynamically Heuristic-Guided Hierarchical Search (GODHS), leverages large language models (LLMs) to infer scene semantics and guide the search process through a multi-level decision hierarchy. Reliability in reasoning is achieved through the use of structured prompts and logical constraints applied at each stage of the hierarchy. For the specific challenges of mobile manipulation, we introduce a heuristic-based motion planner that combines polar angle sorting with distance prioritization to efficiently generate exploration paths. Comprehensive evaluations in Isaac Sim demonstrate the feasibility of our framework, showing that GODHS can locate target objects with higher search efficiency compared to conventional, non-semantic search strategies. Website and Video are available at: https://drapandiger.github.io/GODHS
Human and AI collaboration in Fitness Education:A Longitudinal Study with a Pilates Instructor
Jun 05, 2025Abstract:Artificial intelligence is poised to transform teaching and coaching practices,yet its optimal role alongside human expertise remains unclear.This study investigates human and AI collaboration in fitness education through a one year qualitative case study with a Pilates instructor.The researcher participated in the instructor classes and conducted biweekly semi structured interviews to explore how generative AI could be integrated into class planning and instruction.
Detecting Sybil Addresses in Blockchain Airdrops: A Subgraph-based Feature Propagation and Fusion Approach
May 14, 2025Abstract:Sybil attacks pose a significant security threat to blockchain ecosystems, particularly in token airdrop events. This paper proposes a novel sybil address identification method based on subgraph feature extraction lightGBM. The method first constructs a two-layer deep transaction subgraph for each address, then extracts key event operation features according to the lifecycle of sybil addresses, including the time of first transaction, first gas acquisition, participation in airdrop activities, and last transaction. These temporal features effectively capture the consistency of sybil address behavior operations. Additionally, the method extracts amount and network structure features, comprehensively describing address behavior patterns and network topology through feature propagation and fusion. Experiments conducted on a dataset containing 193,701 addresses (including 23,240 sybil addresses) show that this method outperforms existing approaches in terms of precision, recall, F1 score, and AUC, with all metrics exceeding 0.9. The methods and results of this study can be further applied to broader blockchain security areas such as transaction manipulation identification and token liquidity risk assessment, contributing to the construction of a more secure and fair blockchain ecosystem.
BIGCity: A Universal Spatiotemporal Model for Unified Trajectory and Traffic State Data Analysis
Dec 01, 2024



Abstract:Typical dynamic ST data includes trajectory data (representing individual-level mobility) and traffic state data (representing population-level mobility). Traditional studies often treat trajectory and traffic state data as distinct, independent modalities, each tailored to specific tasks within a single modality. However, real-world applications, such as navigation apps, require joint analysis of trajectory and traffic state data. Treating these data types as two separate domains can lead to suboptimal model performance. Although recent advances in ST data pre-training and ST foundation models aim to develop universal models for ST data analysis, most existing models are "multi-task, solo-data modality" (MTSM), meaning they can handle multiple tasks within either trajectory data or traffic state data, but not both simultaneously. To address this gap, this paper introduces BIGCity, the first multi-task, multi-data modality (MTMD) model for ST data analysis. The model targets two key challenges in designing an MTMD ST model: (1) unifying the representations of different ST data modalities, and (2) unifying heterogeneous ST analysis tasks. To overcome the first challenge, BIGCity introduces a novel ST-unit that represents both trajectories and traffic states in a unified format. Additionally, for the second challenge, BIGCity adopts a tunable large model with ST task-oriented prompt, enabling it to perform a range of heterogeneous tasks without the need for fine-tuning. Extensive experiments on real-world datasets demonstrate that BIGCity achieves state-of-the-art performance across 8 tasks, outperforming 18 baselines. To the best of our knowledge, BIGCity is the first model capable of handling both trajectories and traffic states for diverse heterogeneous tasks. Our code are available at https://github.com/bigscity/BIGCity
The application of GPT-4 in grading design university students' assignment and providing feedback: An exploratory study
Sep 26, 2024



Abstract:This study aims to investigate whether GPT-4 can effectively grade assignments for design university students and provide useful feedback. In design education, assignments do not have a single correct answer and often involve solving an open-ended design problem. This subjective nature of design projects often leads to grading problems,as grades can vary between different raters,for instance instructor from engineering background or architecture background. This study employs an iterative research approach in developing a Custom GPT with the aim of achieving more reliable results and testing whether it can provide design students with constructive feedback. The findings include: First,through several rounds of iterations the inter-reliability between GPT and human raters reached a level that is generally accepted by educators. This indicates that by providing accurate prompts to GPT,and continuously iterating to build a Custom GPT, it can be used to effectively grade students' design assignments, serving as a reliable complement to human raters. Second, the intra-reliability of GPT's scoring at different times is between 0.65 and 0.78. This indicates that, with adequate instructions, a Custom GPT gives consistent results which is a precondition for grading students. As consistency and comparability are the two main rules to ensure the reliability of educational assessment, this study has looked at whether a Custom GPT can be developed that adheres to these two rules. We finish the paper by testing whether Custom GPT can provide students with useful feedback and reflecting on how educators can develop and iterate a Custom GPT to serve as a complementary rater.
Exploring the Use of ChatGPT for a Systematic Literature Review: a Design-Based Research
Sep 25, 2024



Abstract:ChatGPT has been used in several educational contexts,including learning, teaching and research. It also has potential to conduct the systematic literature review (SLR). However, there are limited empirical studies on how to use ChatGPT in conducting a SLR. Based on a SLR published,this study used ChatGPT to conduct a SLR of the same 33 papers in a design-based approach, to see what the differences are by comparing the reviews' results,and to answer: To what extent can ChatGPT conduct SLR? What strategies can human researchers utilize to structure prompts for ChatGPT that enhance the reliability and validity of a SLR? This study found that ChatGPT could conduct a SLR. It needs detailed and accurate prompts to analyze the literature. It also has limitations. Guiding principles are summarized from this study for researchers to follow when they need to conduct SLRs using ChatGPT.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge