Marc Modat
Weakly-Supervised Convolutional Neural Networks for Multimodal Image Registration
Jul 09, 2018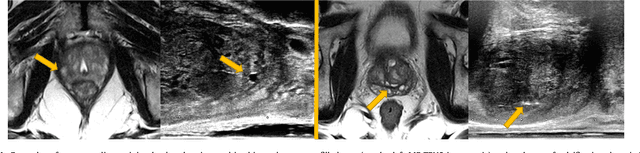
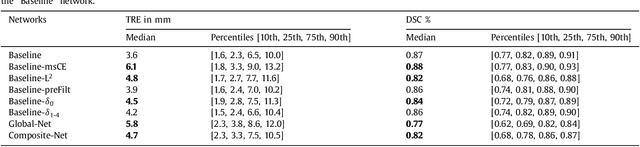
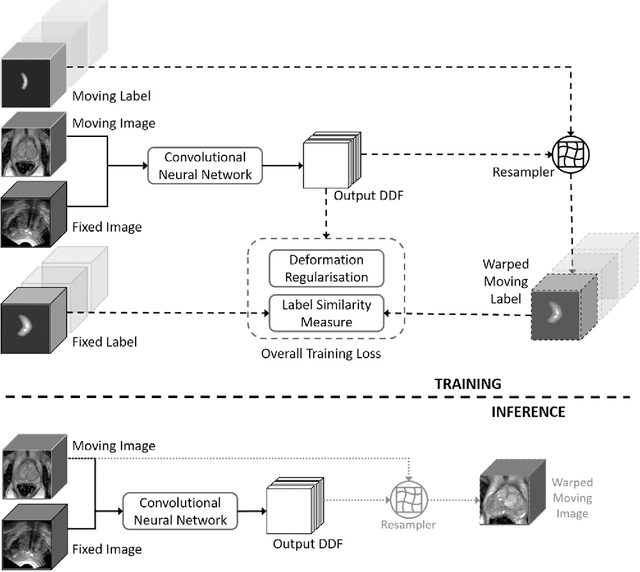
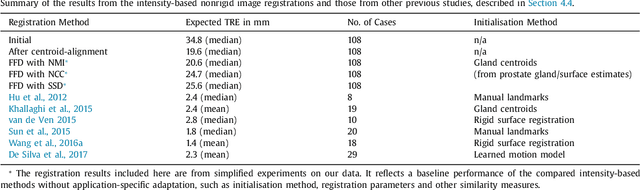
Abstract:One of the fundamental challenges in supervised learning for multimodal image registration is the lack of ground-truth for voxel-level spatial correspondence. This work describes a method to infer voxel-level transformation from higher-level correspondence information contained in anatomical labels. We argue that such labels are more reliable and practical to obtain for reference sets of image pairs than voxel-level correspondence. Typical anatomical labels of interest may include solid organs, vessels, ducts, structure boundaries and other subject-specific ad hoc landmarks. The proposed end-to-end convolutional neural network approach aims to predict displacement fields to align multiple labelled corresponding structures for individual image pairs during the training, while only unlabelled image pairs are used as the network input for inference. We highlight the versatility of the proposed strategy, for training, utilising diverse types of anatomical labels, which need not to be identifiable over all training image pairs. At inference, the resulting 3D deformable image registration algorithm runs in real-time and is fully-automated without requiring any anatomical labels or initialisation. Several network architecture variants are compared for registering T2-weighted magnetic resonance images and 3D transrectal ultrasound images from prostate cancer patients. A median target registration error of 3.6 mm on landmark centroids and a median Dice of 0.87 on prostate glands are achieved from cross-validation experiments, in which 108 pairs of multimodal images from 76 patients were tested with high-quality anatomical labels.
Joint registration and synthesis using a probabilistic model for alignment of MRI and histological sections
Jan 16, 2018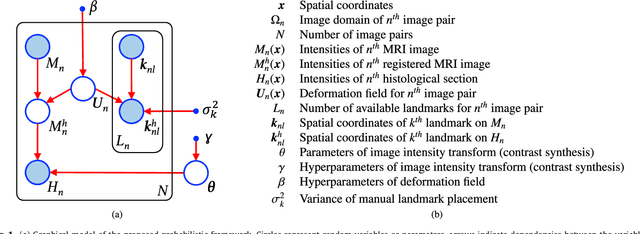
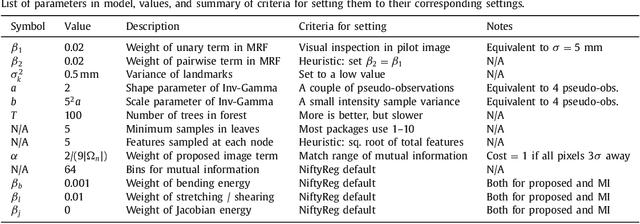
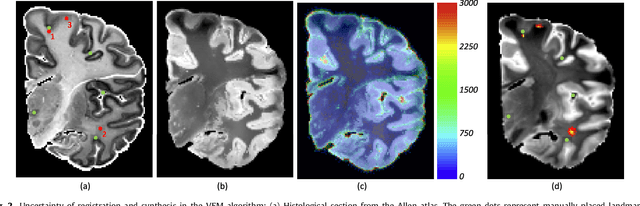
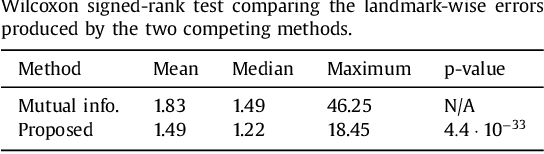
Abstract:Nonlinear registration of 2D histological sections with corresponding slices of MRI data is a critical step of 3D histology reconstruction. This task is difficult due to the large differences in image contrast and resolution, as well as the complex nonrigid distortions produced when sectioning the sample and mounting it on the glass slide. It has been shown in brain MRI registration that better spatial alignment across modalities can be obtained by synthesizing one modality from the other and then using intra-modality registration metrics, rather than by using mutual information (MI) as metric. However, such an approach typically requires a database of aligned images from the two modalities, which is very difficult to obtain for histology/MRI. Here, we overcome this limitation with a probabilistic method that simultaneously solves for registration and synthesis directly on the target images, without any training data. In our model, the MRI slice is assumed to be a contrast-warped, spatially deformed version of the histological section. We use approximate Bayesian inference to iteratively refine the probabilistic estimate of the synthesis and the registration, while accounting for each other's uncertainty. Moreover, manually placed landmarks can be seamlessly integrated in the framework for increased performance. Experiments on a synthetic dataset show that, compared with MI, the proposed method makes it possible to use a much more flexible deformation model in the registration to improve its accuracy, without compromising robustness. Moreover, our framework also exploits information in manually placed landmarks more efficiently than MI, since landmarks inform both synthesis and registration - as opposed to registration alone. Finally, we show qualitative results on the public Allen atlas, in which the proposed method provides a clear improvement over MI based registration.
Label-driven weakly-supervised learning for multimodal deformable image registration
Dec 24, 2017Abstract:Spatially aligning medical images from different modalities remains a challenging task, especially for intraoperative applications that require fast and robust algorithms. We propose a weakly-supervised, label-driven formulation for learning 3D voxel correspondence from higher-level label correspondence, thereby bypassing classical intensity-based image similarity measures. During training, a convolutional neural network is optimised by outputting a dense displacement field (DDF) that warps a set of available anatomical labels from the moving image to match their corresponding counterparts in the fixed image. These label pairs, including solid organs, ducts, vessels, point landmarks and other ad hoc structures, are only required at training time and can be spatially aligned by minimising a cross-entropy function of the warped moving label and the fixed label. During inference, the trained network takes a new image pair to predict an optimal DDF, resulting in a fully-automatic, label-free, real-time and deformable registration. For interventional applications where large global transformation prevails, we also propose a neural network architecture to jointly optimise the global- and local displacements. Experiment results are presented based on cross-validating registrations of 111 pairs of T2-weighted magnetic resonance images and 3D transrectal ultrasound images from prostate cancer patients with a total of over 4000 anatomical labels, yielding a median target registration error of 4.2 mm on landmark centroids and a median Dice of 0.88 on prostate glands.
NiftyNet: a deep-learning platform for medical imaging
Oct 16, 2017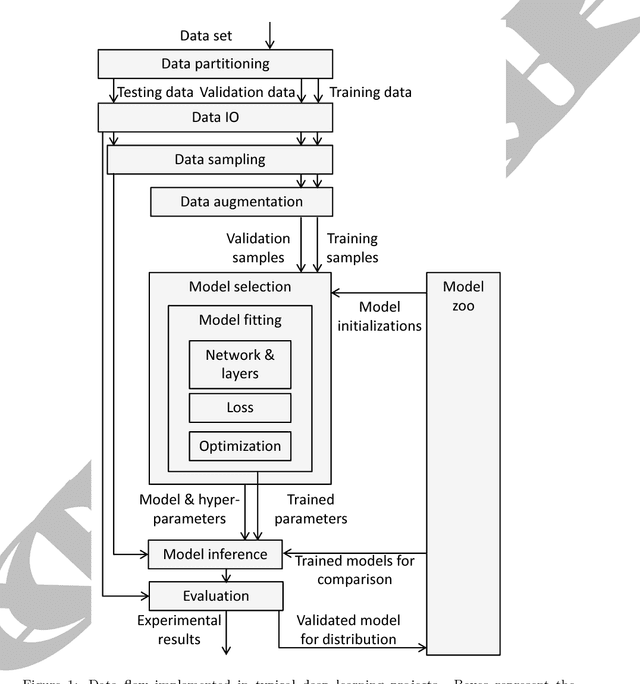
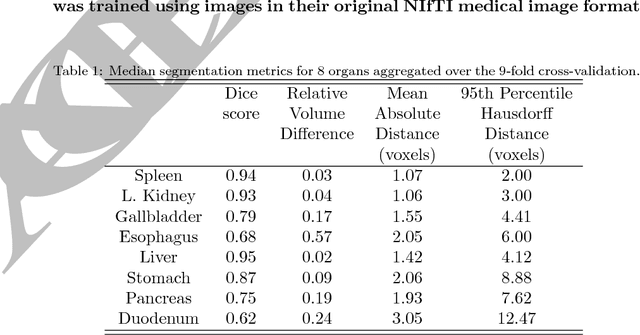
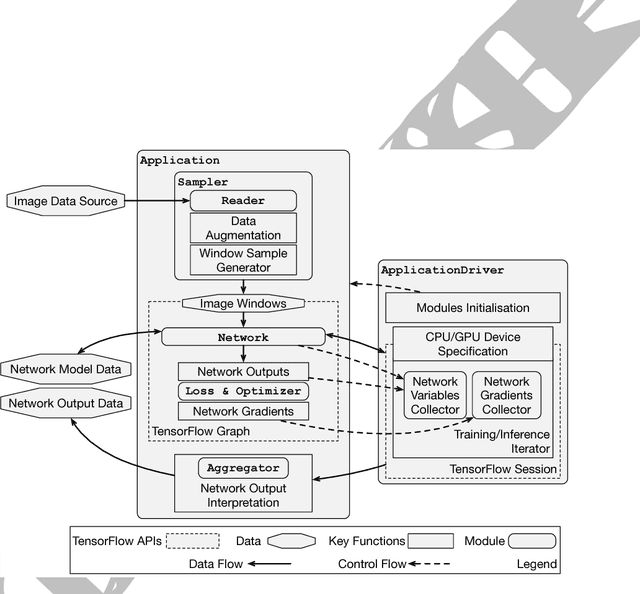
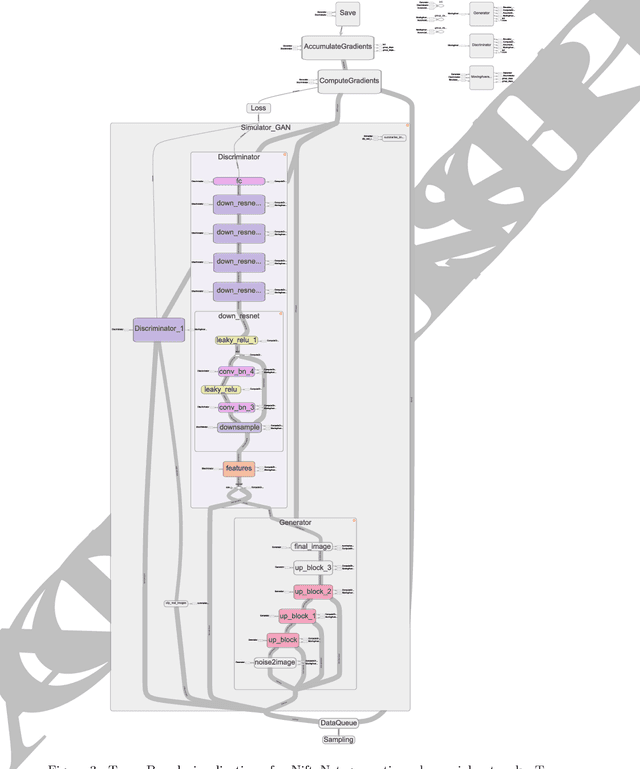
Abstract:Medical image analysis and computer-assisted intervention problems are increasingly being addressed with deep-learning-based solutions. Established deep-learning platforms are flexible but do not provide specific functionality for medical image analysis and adapting them for this application requires substantial implementation effort. Thus, there has been substantial duplication of effort and incompatible infrastructure developed across many research groups. This work presents the open-source NiftyNet platform for deep learning in medical imaging. The ambition of NiftyNet is to accelerate and simplify the development of these solutions, and to provide a common mechanism for disseminating research outputs for the community to use, adapt and build upon. NiftyNet provides a modular deep-learning pipeline for a range of medical imaging applications including segmentation, regression, image generation and representation learning applications. Components of the NiftyNet pipeline including data loading, data augmentation, network architectures, loss functions and evaluation metrics are tailored to, and take advantage of, the idiosyncracies of medical image analysis and computer-assisted intervention. NiftyNet is built on TensorFlow and supports TensorBoard visualization of 2D and 3D images and computational graphs by default. We present 3 illustrative medical image analysis applications built using NiftyNet: (1) segmentation of multiple abdominal organs from computed tomography; (2) image regression to predict computed tomography attenuation maps from brain magnetic resonance images; and (3) generation of simulated ultrasound images for specified anatomical poses. NiftyNet enables researchers to rapidly develop and distribute deep learning solutions for segmentation, regression, image generation and representation learning applications, or extend the platform to new applications.
Part-to-whole Registration of Histology and MRI using Shape Elements
Aug 27, 2017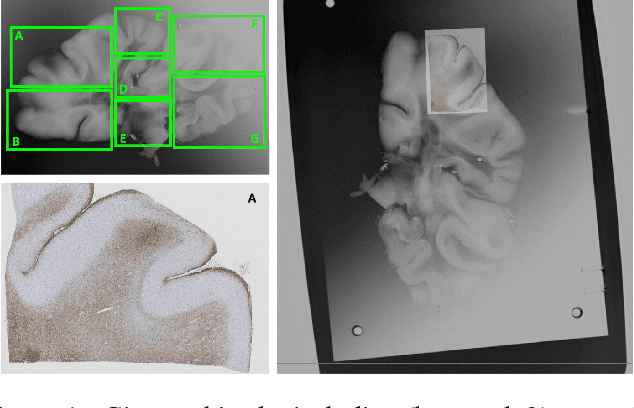
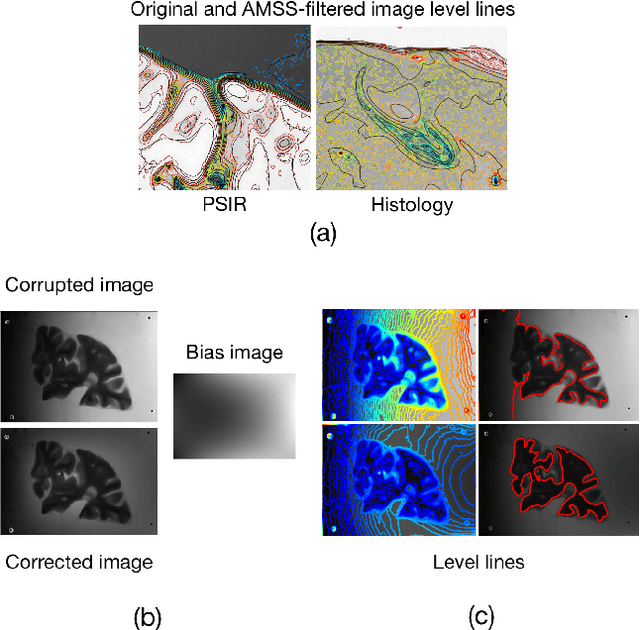

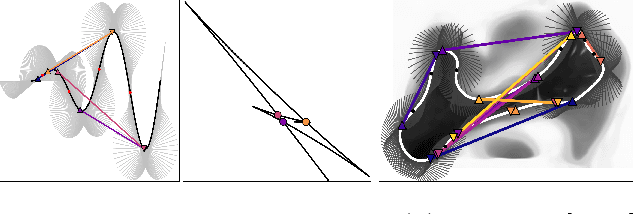
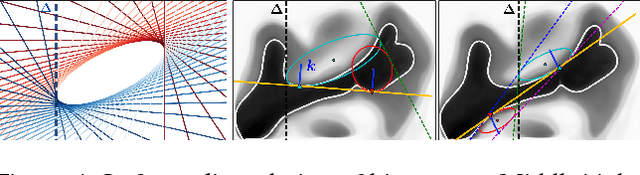
Abstract:Image registration between histology and magnetic resonance imaging (MRI) is a challenging task due to differences in structural content and contrast. Too thick and wide specimens cannot be processed all at once and must be cut into smaller pieces. This dramatically increases the complexity of the problem, since each piece should be individually and manually pre-aligned. To the best of our knowledge, no automatic method can reliably locate such piece of tissue within its respective whole in the MRI slice, and align it without any prior information. We propose here a novel automatic approach to the joint problem of multimodal registration between histology and MRI, when only a fraction of tissue is available from histology. The approach relies on the representation of images using their level lines so as to reach contrast invariance. Shape elements obtained via the extraction of bitangents are encoded in a projective-invariant manner, which permits the identification of common pieces of curves between two images. We evaluated the approach on human brain histology and compared resulting alignments against manually annotated ground truths. Considering the complexity of the brain folding patterns, preliminary results are promising and suggest the use of characteristic and meaningful shape elements for improved robustness and efficiency.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge