Lei Xing
Consistency-guided Meta-Learning for Bootstrapping Semi-Supervised Medical Image Segmentation
Jul 21, 2023



Abstract:Medical imaging has witnessed remarkable progress but usually requires a large amount of high-quality annotated data which is time-consuming and costly to obtain. To alleviate this burden, semi-supervised learning has garnered attention as a potential solution. In this paper, we present Meta-Learning for Bootstrapping Medical Image Segmentation (MLB-Seg), a novel method for tackling the challenge of semi-supervised medical image segmentation. Specifically, our approach first involves training a segmentation model on a small set of clean labeled images to generate initial labels for unlabeled data. To further optimize this bootstrapping process, we introduce a per-pixel weight mapping system that dynamically assigns weights to both the initialized labels and the model's own predictions. These weights are determined using a meta-process that prioritizes pixels with loss gradient directions closer to those of clean data, which is based on a small set of precisely annotated images. To facilitate the meta-learning process, we additionally introduce a consistency-based Pseudo Label Enhancement (PLE) scheme that improves the quality of the model's own predictions by ensembling predictions from various augmented versions of the same input. In order to improve the quality of the weight maps obtained through multiple augmentations of a single input, we introduce a mean teacher into the PLE scheme. This method helps to reduce noise in the weight maps and stabilize its generation process. Our extensive experimental results on public atrial and prostate segmentation datasets demonstrate that our proposed method achieves state-of-the-art results under semi-supervision. Our code is available at https://github.com/aijinrjinr/MLB-Seg.
Towards Medical Artificial General Intelligence via Knowledge-Enhanced Multimodal Pretraining
Apr 26, 2023



Abstract:Medical artificial general intelligence (MAGI) enables one foundation model to solve different medical tasks, which is very practical in the medical domain. It can significantly reduce the requirement of large amounts of task-specific data by sufficiently sharing medical knowledge among different tasks. However, due to the challenges of designing strongly generalizable models with limited and complex medical data, most existing approaches tend to develop task-specific models. To take a step towards MAGI, we propose a new paradigm called Medical-knOwledge-enhanced mulTimOdal pretRaining (MOTOR). In MOTOR, we combine two kinds of basic medical knowledge, i.e., general and specific knowledge, in a complementary manner to boost the general pretraining process. As a result, the foundation model with comprehensive basic knowledge can learn compact representations from pretraining radiographic data for better cross-modal alignment. MOTOR unifies the understanding and generation, which are two kinds of core intelligence of an AI system, into a single medical foundation model, to flexibly handle more diverse medical tasks. To enable a comprehensive evaluation and facilitate further research, we construct a medical multimodal benchmark including a wide range of downstream tasks, such as chest x-ray report generation and medical visual question answering. Extensive experiments on our benchmark show that MOTOR obtains promising results through simple task-oriented adaptation. The visualization shows that the injected knowledge successfully highlights key information in the medical data, demonstrating the excellent interpretability of MOTOR. Our MOTOR successfully mimics the human practice of fulfilling a "medical student" to accelerate the process of becoming a "specialist". We believe that our work makes a significant stride in realizing MAGI.
Learning Treatment Plan Representations for Content Based Image Retrieval
Jun 06, 2022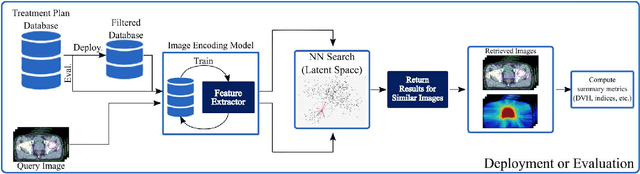
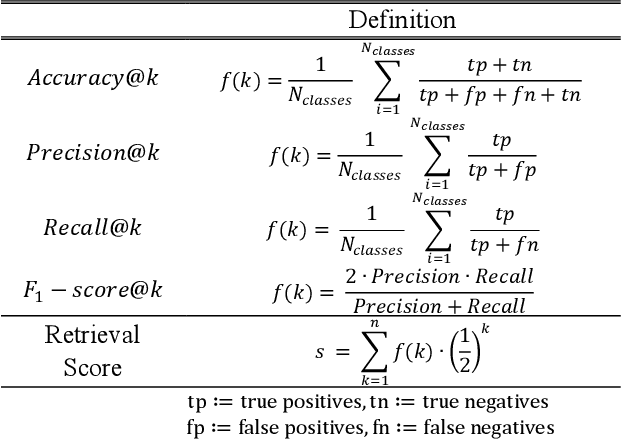

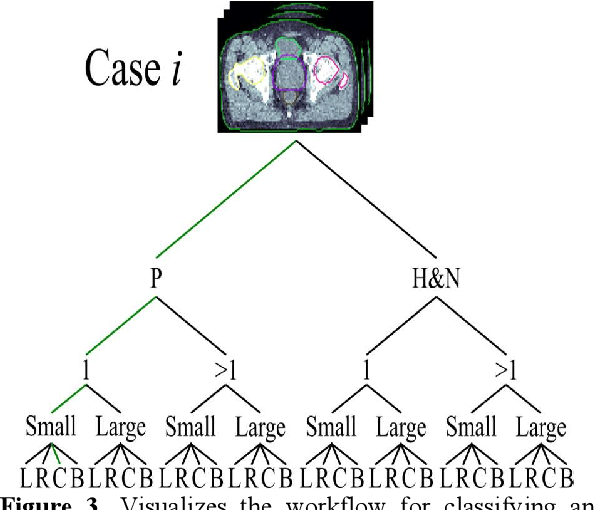
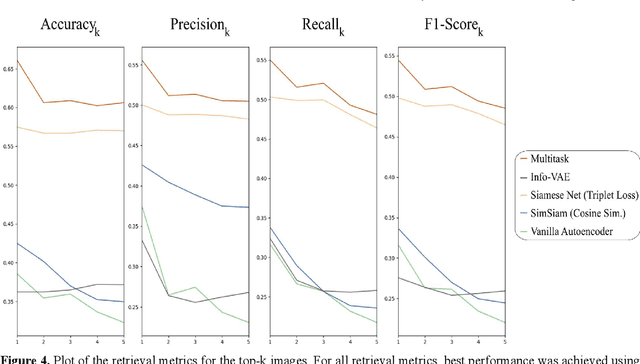
Abstract:Objective: Knowledge based planning (KBP) typically involves training an end-to-end deep learning model to predict dose distributions. However, training end-to-end KBP methods may be associated with practical limitations due to the limited size of medical datasets that are often used. To address these limitations, we propose a content based image retrieval (CBIR) method for retrieving dose distributions of previously planned patients based on anatomical similarity. Approach: Our proposed CBIR method trains a representation model that produces latent space embeddings of a patient's anatomical information. The latent space embeddings of new patients are then compared against those of previous patients in a database for image retrieval of dose distributions. Summary metrics (e.g. dose-volume histogram, conformity index, homogeneity index, etc.) are computed and can then be utilized in subsequent automated planning. All source code for this project is available on github. Main Results: The retrieval performance of various CBIR methods is evaluated on a dataset consisting of both publicly available plans and clinical plans from our institution. This study compares various encoding methods, ranging from simple autoencoders to more recent Siamese networks like SimSiam, and the best performance was observed for the multitask Siamese network. Significance: Applying CBIR to inform subsequent treatment planning potentially addresses many limitations associated with end-to-end KBP. Our current results demonstrate that excellent image retrieval performance can be obtained through slight changes to previously developed Siamese networks. We hope to integrate CBIR into automated planning workflow in future works, potentially through methods like the MetaPlanner framework.
Label-Efficient Self-Supervised Federated Learning for Tackling Data Heterogeneity in Medical Imaging
May 17, 2022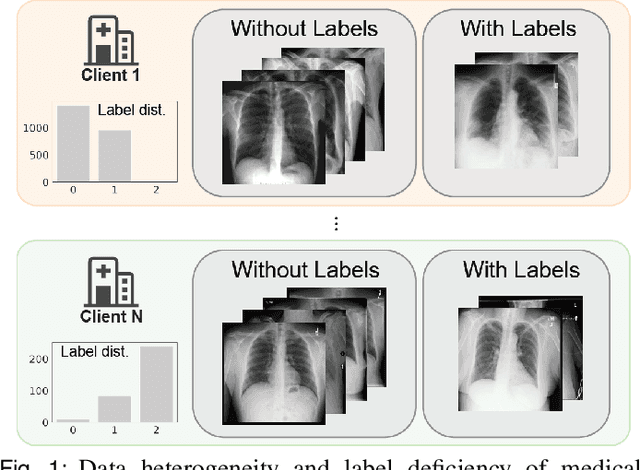
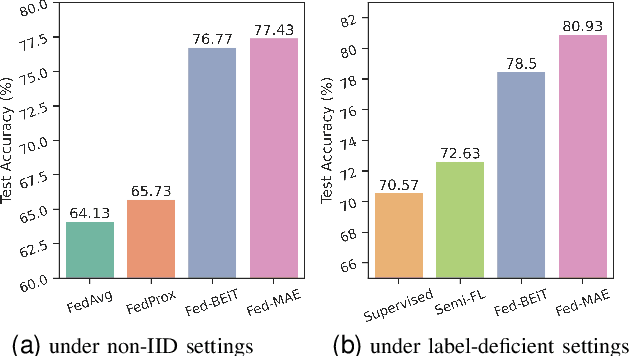
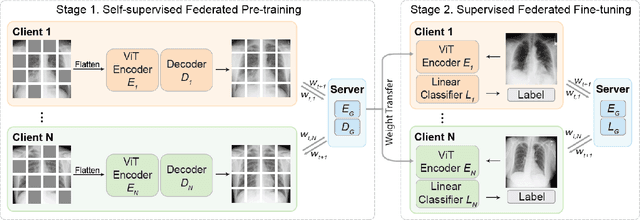
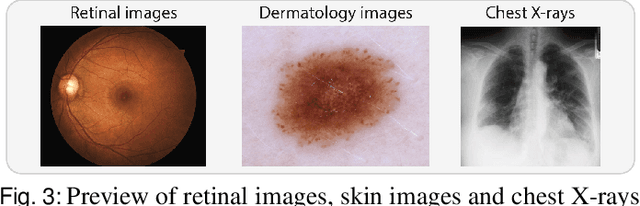
Abstract:The curation of large-scale medical datasets from multiple institutions necessary for training deep learning models is challenged by the difficulty in sharing patient data with privacy-preserving. Federated learning (FL), a paradigm that enables privacy-protected collaborative learning among different institutions, is a promising solution to this challenge. However, FL generally suffers from performance deterioration due to heterogeneous data distributions across institutions and the lack of quality labeled data. In this paper, we present a robust and label-efficient self-supervised FL framework for medical image analysis. Specifically, we introduce a novel distributed self-supervised pre-training paradigm into the existing FL pipeline (i.e., pre-training the models directly on the decentralized target task datasets). Built upon the recent success of Vision Transformers, we employ masked image encoding tasks for self-supervised pre-training, to facilitate more effective knowledge transfer to downstream federated models. Extensive empirical results on simulated and real-world medical imaging federated datasets show that self-supervised pre-training largely benefits the robustness of federated models against various degrees of data heterogeneity. Notably, under severe data heterogeneity, our method, without relying on any additional pre-training data, achieves an improvement of 5.06%, 1.53% and 4.58% in test accuracy on retinal, dermatology and chest X-ray classification compared with the supervised baseline with ImageNet pre-training. Moreover, we show that our self-supervised FL algorithm generalizes well to out-of-distribution data and learns federated models more effectively in limited label scenarios, surpassing the supervised baseline by 10.36% and the semi-supervised FL method by 8.3% in test accuracy.
CSN: Component-Supervised Network for Few-Shot Classification
Mar 15, 2022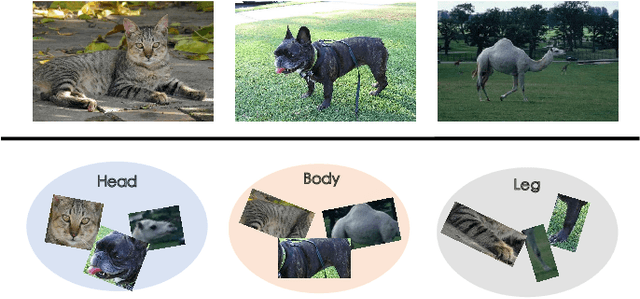
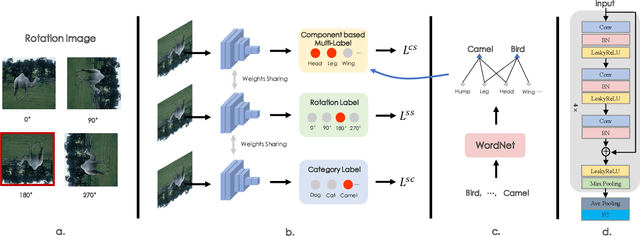

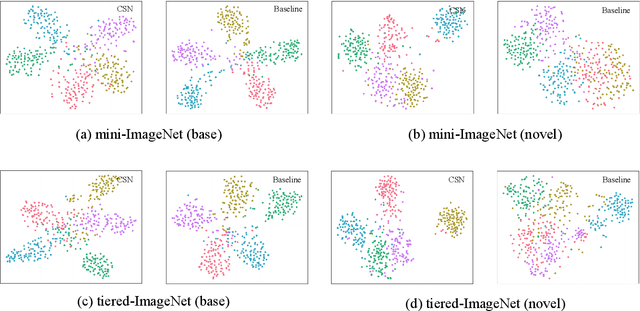
Abstract:The few-shot classification (FSC) task has been a hot research topic in recent years. It aims to address the classification problem with insufficient labeled data on a cross-category basis. Typically, researchers pre-train a feature extractor with base data, then use it to extract the features of novel data and recognize them. Notably, the novel set only has a few annotated samples and has entirely different categories from the base set, which leads to that the pre-trained feature extractor can not adapt to the novel data flawlessly. We dub this problem as Feature-Extractor-Maladaptive (FEM) problem. Starting from the root cause of this problem, this paper presents a new scheme, Component-Supervised Network (CSN), to improve the performance of FSC. We believe that although the categories of base and novel sets are different, the composition of the sample's components is similar. For example, both cat and dog contain leg and head components. Actually, such entity components are intra-class stable. They have fine cross-category versatility and new category generalization. Therefore, we refer to WordNet, a dictionary commonly used in natural language processing, to collect component information of samples and construct a component-based auxiliary task to improve the adaptability of the feature extractor. We conduct experiments on two benchmark datasets (mini-ImageNet and tiered-ImageNet), the improvements of $0.9\%$-$5.8\%$ compared with state-of-the-arts have evaluated the efficiency of our CSN.
Learning to Bootstrap for Combating Label Noise
Feb 09, 2022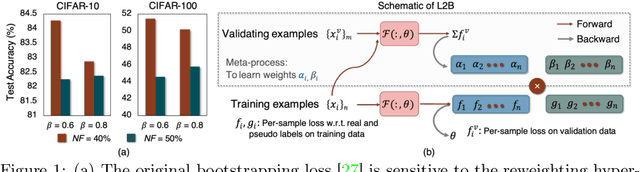
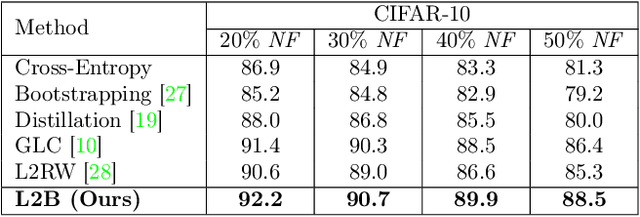
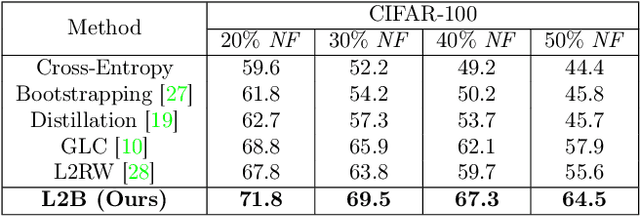
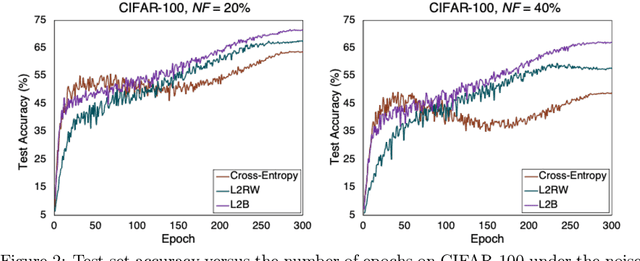
Abstract:Deep neural networks are powerful tools for representation learning, but can easily overfit to noisy labels which are prevalent in many real-world scenarios. Generally, noisy supervision could stem from variation among labelers, label corruption by adversaries, etc. To combat such label noises, one popular line of approach is to apply customized weights to the training instances, so that the corrupted examples contribute less to the model learning. However, such learning mechanisms potentially erase important information about the data distribution and therefore yield suboptimal results. To leverage useful information from the corrupted instances, an alternative is the bootstrapping loss, which reconstructs new training targets on-the-fly by incorporating the network's own predictions (i.e., pseudo-labels). In this paper, we propose a more generic learnable loss objective which enables a joint reweighting of instances and labels at once. Specifically, our method dynamically adjusts the per-sample importance weight between the real observed labels and pseudo-labels, where the weights are efficiently determined in a meta process. Compared to the previous instance reweighting methods, our approach concurrently conducts implicit relabeling, and thereby yield substantial improvements with almost no extra cost. Extensive experimental results demonstrated the strengths of our approach over existing methods on multiple natural and medical image benchmark datasets, including CIFAR-10, CIFAR-100, ISIC2019 and Clothing 1M. The code is publicly available at https://github.com/yuyinzhou/L2B.
Image Classification using Graph Neural Network and Multiscale Wavelet Superpixels
Jan 29, 2022



Abstract:Prior studies using graph neural networks (GNNs) for image classification have focused on graphs generated from a regular grid of pixels or similar-sized superpixels. In the latter, a single target number of superpixels is defined for an entire dataset irrespective of differences across images and their intrinsic multiscale structure. On the contrary, this study investigates image classification using graphs generated from an image-specific number of multiscale superpixels. We propose WaveMesh, a new wavelet-based superpixeling algorithm, where the number and sizes of superpixels in an image are systematically computed based on its content. WaveMesh superpixel graphs are structurally different from similar-sized superpixel graphs. We use SplineCNN, a state-of-the-art network for image graph classification, to compare WaveMesh and similar-sized superpixels. Using SplineCNN, we perform extensive experiments on three benchmark datasets under three local-pooling settings: 1) no pooling, 2) GraclusPool, and 3) WavePool, a novel spatially heterogeneous pooling scheme tailored to WaveMesh superpixels. Our experiments demonstrate that SplineCNN learns from multiscale WaveMesh superpixels on-par with similar-sized superpixels. In all WaveMesh experiments, GraclusPool performs poorer than no pooling / WavePool, indicating that poor choice of pooling can result in inferior performance while learning from multiscale superpixels.
SSDL: Self-Supervised Dictionary Learning
Dec 03, 2021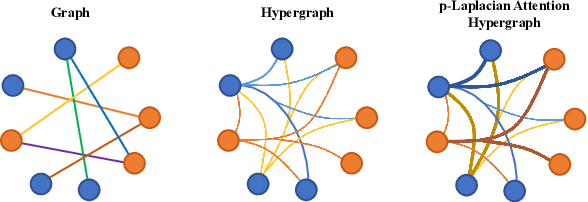
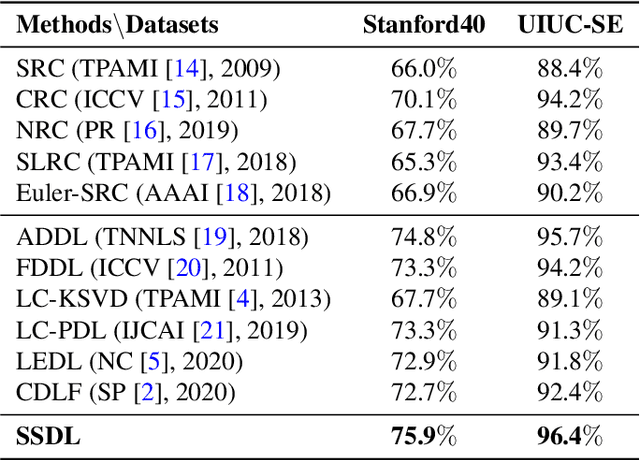
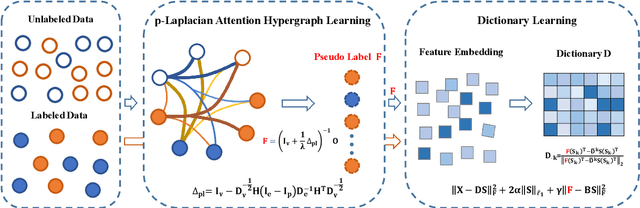
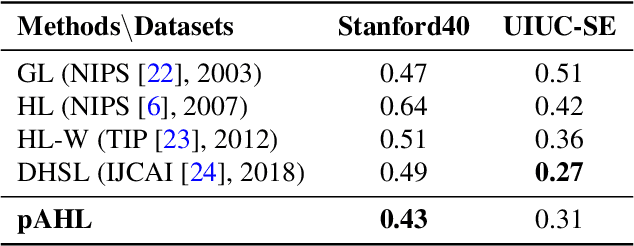
Abstract:The label-embedded dictionary learning (DL) algorithms generate influential dictionaries by introducing discriminative information. However, there exists a limitation: All the label-embedded DL methods rely on the labels due that this way merely achieves ideal performances in supervised learning. While in semi-supervised and unsupervised learning, it is no longer sufficient to be effective. Inspired by the concept of self-supervised learning (e.g., setting the pretext task to generate a universal model for the downstream task), we propose a Self-Supervised Dictionary Learning (SSDL) framework to address this challenge. Specifically, we first design a $p$-Laplacian Attention Hypergraph Learning (pAHL) block as the pretext task to generate pseudo soft labels for DL. Then, we adopt the pseudo labels to train a dictionary from a primary label-embedded DL method. We evaluate our SSDL on two human activity recognition datasets. The comparison results with other state-of-the-art methods have demonstrated the efficiency of SSDL.
MDFM: Multi-Decision Fusing Model for Few-Shot Learning
Dec 03, 2021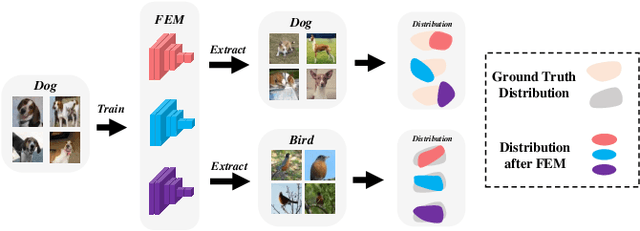
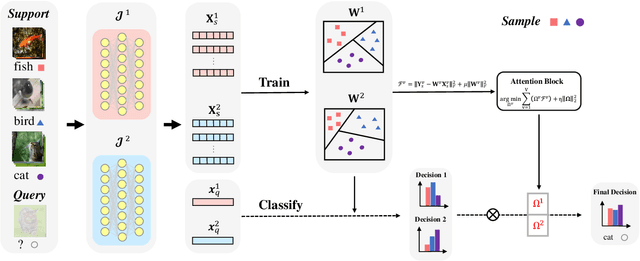
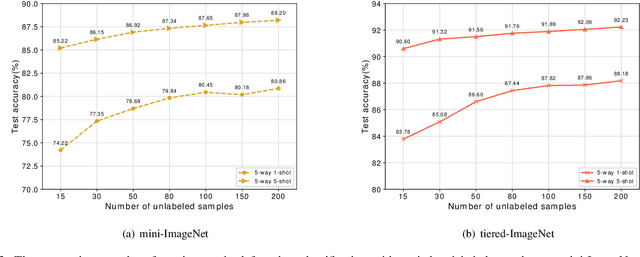
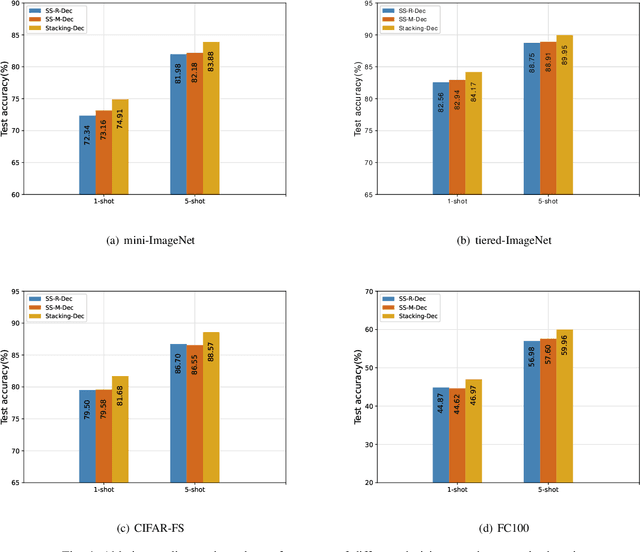
Abstract:In recent years, researchers pay growing attention to the few-shot learning (FSL) task to address the data-scarce problem. A standard FSL framework is composed of two components: i) Pre-train. Employ the base data to generate a CNN-based feature extraction model (FEM). ii) Meta-test. Apply the trained FEM to the novel data (category is different from base data) to acquire the feature embeddings and recognize them. Although researchers have made remarkable breakthroughs in FSL, there still exists a fundamental problem. Since the trained FEM with base data usually cannot adapt to the novel class flawlessly, the novel data's feature may lead to the distribution shift problem. To address this challenge, we hypothesize that even if most of the decisions based on different FEMs are viewed as weak decisions, which are not available for all classes, they still perform decently in some specific categories. Inspired by this assumption, we propose a novel method Multi-Decision Fusing Model (MDFM), which comprehensively considers the decisions based on multiple FEMs to enhance the efficacy and robustness of the model. MDFM is a simple, flexible, non-parametric method that can directly apply to the existing FEMs. Besides, we extend the proposed MDFM to two FSL settings (i.e., supervised and semi-supervised settings). We evaluate the proposed method on five benchmark datasets and achieve significant improvements of 3.4%-7.3% compared with state-of-the-arts.
RadFusion: Benchmarking Performance and Fairness for Multimodal Pulmonary Embolism Detection from CT and EHR
Nov 27, 2021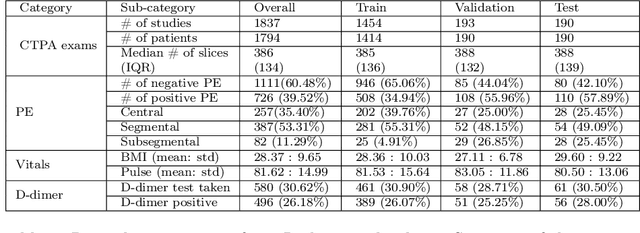
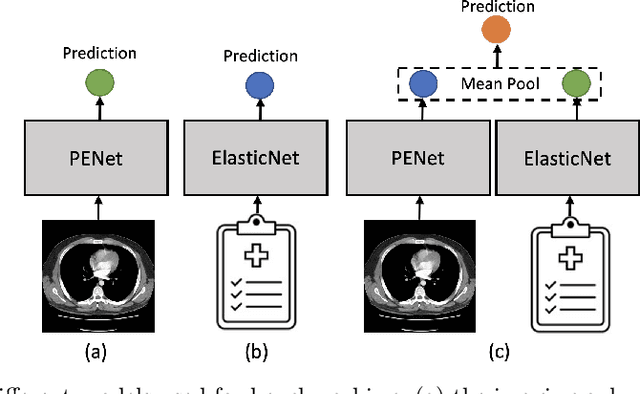
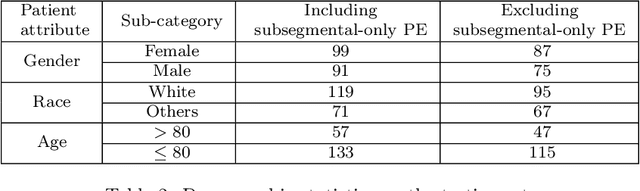
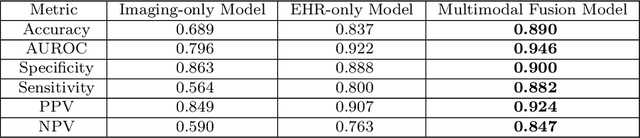
Abstract:Despite the routine use of electronic health record (EHR) data by radiologists to contextualize clinical history and inform image interpretation, the majority of deep learning architectures for medical imaging are unimodal, i.e., they only learn features from pixel-level information. Recent research revealing how race can be recovered from pixel data alone highlights the potential for serious biases in models which fail to account for demographics and other key patient attributes. Yet the lack of imaging datasets which capture clinical context, inclusive of demographics and longitudinal medical history, has left multimodal medical imaging underexplored. To better assess these challenges, we present RadFusion, a multimodal, benchmark dataset of 1794 patients with corresponding EHR data and high-resolution computed tomography (CT) scans labeled for pulmonary embolism. We evaluate several representative multimodal fusion models and benchmark their fairness properties across protected subgroups, e.g., gender, race/ethnicity, age. Our results suggest that integrating imaging and EHR data can improve classification performance and robustness without introducing large disparities in the true positive rate between population groups.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge