Jian K. Liu
University of Birmingham, Birmingham, United Kingdom
Copilot-Assisted Second-Thought Framework for Brain-to-Robot Hand Motion Decoding
Mar 29, 2026Abstract:Motor kinematics prediction (MKP) from electroencephalography (EEG) is an important research area for developing movement-related brain-computer interfaces (BCIs). While traditional methods often rely on convolutional neural networks (CNNs) or recurrent neural networks (RNNs), Transformer-based models have shown strong ability in modeling long sequential EEG data. In this study, we propose a CNN-attention hybrid model for decoding hand kinematics from EEG during grasp-and-lift tasks, achieving strong performance in within-subject experiments. We further extend this approach to EEG-EMG multimodal decoding, which yields substantially improved results. Within-subject tests achieve PCC values of 0.9854, 0.9946, and 0.9065 for the X, Y, and Z axes, respectively, computed on the midpoint trajectory between the thumb and index finger, while cross-subject tests result in 0.9643, 0.9795, and 0.5852. The decoded trajectories from both modalities are then used to control a Franka Panda robotic arm in a MuJoCo simulation. To enhance trajectory fidelity, we introduce a copilot framework that filters low-confidence decoded points using a motion-state-aware critic within a finite-state machine. This post-processing step improves the overall within-subject PCC of EEG-only decoding to 0.93 while excluding fewer than 20% of the data points.
SeqProFT: Applying LoRA Finetuning for Sequence-only Protein Property Predictions
Nov 18, 2024Abstract:Protein language models (PLMs) are capable of learning the relationships between protein sequences and functions by treating amino acid sequences as textual data in a self-supervised manner. However, fine-tuning these models typically demands substantial computational resources and time, with results that may not always be optimized for specific tasks. To overcome these challenges, this study employs the LoRA method to perform end-to-end fine-tuning of the ESM-2 model specifically for protein property prediction tasks, utilizing only sequence information. Additionally, a multi-head attention mechanism is integrated into the downstream network to combine sequence features with contact map information, thereby enhancing the model's comprehension of protein sequences. Experimental results of extensive classification and regression tasks demonstrate that the fine-tuned model achieves strong performance and faster convergence across multiple regression and classification tasks.
Aligning Neuronal Coding of Dynamic Visual Scenes with Foundation Vision Models
Jul 15, 2024



Abstract:Our brains represent the ever-changing environment with neurons in a highly dynamic fashion. The temporal features of visual pixels in dynamic natural scenes are entrapped in the neuronal responses of the retina. It is crucial to establish the intrinsic temporal relationship between visual pixels and neuronal responses. Recent foundation vision models have paved an advanced way of understanding image pixels. Yet, neuronal coding in the brain largely lacks a deep understanding of its alignment with pixels. Most previous studies employ static images or artificial videos derived from static images for emulating more real and complicated stimuli. Despite these simple scenarios effectively help to separate key factors influencing visual coding, complex temporal relationships receive no consideration. To decompose the temporal features of visual coding in natural scenes, here we propose Vi-ST, a spatiotemporal convolutional neural network fed with a self-supervised Vision Transformer (ViT) prior, aimed at unraveling the temporal-based encoding patterns of retinal neuronal populations. The model demonstrates robust predictive performance in generalization tests. Furthermore, through detailed ablation experiments, we demonstrate the significance of each temporal module. Furthermore, we introduce a visual coding evaluation metric designed to integrate temporal considerations and compare the impact of different numbers of neuronal populations on complementary coding. In conclusion, our proposed Vi-ST demonstrates a novel modeling framework for neuronal coding of dynamic visual scenes in the brain, effectively aligning our brain representation of video with neuronal activity. The code is available at https://github.com/wurining/Vi-ST.
Converting High-Performance and Low-Latency SNNs through Explicit Modelling of Residual Error in ANNs
Apr 26, 2024



Abstract:Spiking neural networks (SNNs) have garnered interest due to their energy efficiency and superior effectiveness on neuromorphic chips compared with traditional artificial neural networks (ANNs). One of the mainstream approaches to implementing deep SNNs is the ANN-SNN conversion, which integrates the efficient training strategy of ANNs with the energy-saving potential and fast inference capability of SNNs. However, under extreme low-latency conditions, the existing conversion theory suggests that the problem of misrepresentation of residual membrane potentials in SNNs, i.e., the inability of IF neurons with a reset-by-subtraction mechanism to respond to residual membrane potentials beyond the range from resting potential to threshold, leads to a performance gap in the converted SNNs compared to the original ANNs. This severely limits the possibility of practical application of SNNs on delay-sensitive edge devices. Existing conversion methods addressing this problem usually involve modifying the state of the conversion spiking neurons. However, these methods do not consider their adaptability and compatibility with neuromorphic chips. We propose a new approach based on explicit modeling of residual errors as additive noise. The noise is incorporated into the activation function of the source ANN, which effectively reduces the residual error. Our experiments on the CIFAR10/100 dataset verify that our approach exceeds the prevailing ANN-SNN conversion methods and directly trained SNNs concerning accuracy and the required time steps. Overall, our method provides new ideas for improving SNN performance under ultra-low-latency conditions and is expected to promote practical neuromorphic hardware applications for further development.
Deep Learning for Visual Neuroprosthesis
Jan 08, 2024Abstract:The visual pathway involves complex networks of cells and regions which contribute to the encoding and processing of visual information. While some aspects of visual perception are understood, there are still many unanswered questions regarding the exact mechanisms of visual encoding and the organization of visual information along the pathway. This chapter discusses the importance of visual perception and the challenges associated with understanding how visual information is encoded and represented in the brain. Furthermore, this chapter introduces the concept of neuroprostheses: devices designed to enhance or replace bodily functions, and highlights the importance of constructing computational models of the visual pathway in the implementation of such devices. A number of such models, employing the use of deep learning models, are outlined, and their value to understanding visual coding and natural vision is discussed.
Spike timing reshapes robustness against attacks in spiking neural networks
Jun 09, 2023



Abstract:The success of deep learning in the past decade is partially shrouded in the shadow of adversarial attacks. In contrast, the brain is far more robust at complex cognitive tasks. Utilizing the advantage that neurons in the brain communicate via spikes, spiking neural networks (SNNs) are emerging as a new type of neural network model, boosting the frontier of theoretical investigation and empirical application of artificial neural networks and deep learning. Neuroscience research proposes that the precise timing of neural spikes plays an important role in the information coding and sensory processing of the biological brain. However, the role of spike timing in SNNs is less considered and far from understood. Here we systematically explored the timing mechanism of spike coding in SNNs, focusing on the robustness of the system against various types of attacks. We found that SNNs can achieve higher robustness improvement using the coding principle of precise spike timing in neural encoding and decoding, facilitated by different learning rules. Our results suggest that the utility of spike timing coding in SNNs could improve the robustness against attacks, providing a new approach to reliable coding principles for developing next-generation brain-inspired deep learning.
ESL-SNNs: An Evolutionary Structure Learning Strategy for Spiking Neural Networks
Jun 06, 2023



Abstract:Spiking neural networks (SNNs) have manifested remarkable advantages in power consumption and event-driven property during the inference process. To take full advantage of low power consumption and improve the efficiency of these models further, the pruning methods have been explored to find sparse SNNs without redundancy connections after training. However, parameter redundancy still hinders the efficiency of SNNs during training. In the human brain, the rewiring process of neural networks is highly dynamic, while synaptic connections maintain relatively sparse during brain development. Inspired by this, here we propose an efficient evolutionary structure learning (ESL) framework for SNNs, named ESL-SNNs, to implement the sparse SNN training from scratch. The pruning and regeneration of synaptic connections in SNNs evolve dynamically during learning, yet keep the structural sparsity at a certain level. As a result, the ESL-SNNs can search for optimal sparse connectivity by exploring all possible parameters across time. Our experiments show that the proposed ESL-SNNs framework is able to learn SNNs with sparse structures effectively while reducing the limited accuracy. The ESL-SNNs achieve merely 0.28% accuracy loss with 10% connection density on the DVS-Cifar10 dataset. Our work presents a brand-new approach for sparse training of SNNs from scratch with biologically plausible evolutionary mechanisms, closing the gap in the expressibility between sparse training and dense training. Hence, it has great potential for SNN lightweight training and inference with low power consumption and small memory usage.
Biologically inspired structure learning with reverse knowledge distillation for spiking neural networks
Apr 19, 2023Abstract:Spiking neural networks (SNNs) have superb characteristics in sensory information recognition tasks due to their biological plausibility. However, the performance of some current spiking-based models is limited by their structures which means either fully connected or too-deep structures bring too much redundancy. This redundancy from both connection and neurons is one of the key factors hindering the practical application of SNNs. Although Some pruning methods were proposed to tackle this problem, they normally ignored the fact the neural topology in the human brain could be adjusted dynamically. Inspired by this, this paper proposed an evolutionary-based structure construction method for constructing more reasonable SNNs. By integrating the knowledge distillation and connection pruning method, the synaptic connections in SNNs can be optimized dynamically to reach an optimal state. As a result, the structure of SNNs could not only absorb knowledge from the teacher model but also search for deep but sparse network topology. Experimental results on CIFAR100 and DVS-Gesture show that the proposed structure learning method can get pretty well performance while reducing the connection redundancy. The proposed method explores a novel dynamical way for structure learning from scratch in SNNs which could build a bridge to close the gap between deep learning and bio-inspired neural dynamics.
Reconstruction of Natural Visual Scenes from Neural Spikes with Deep Neural Networks
Apr 30, 2019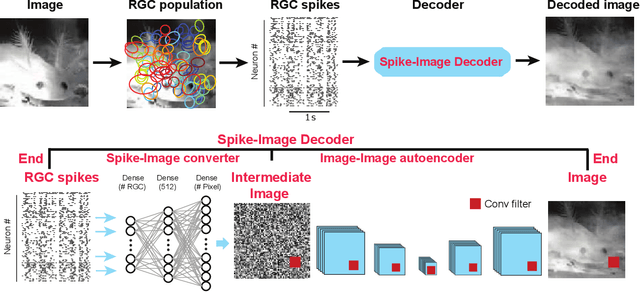
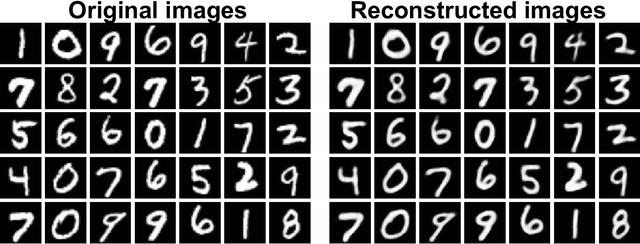
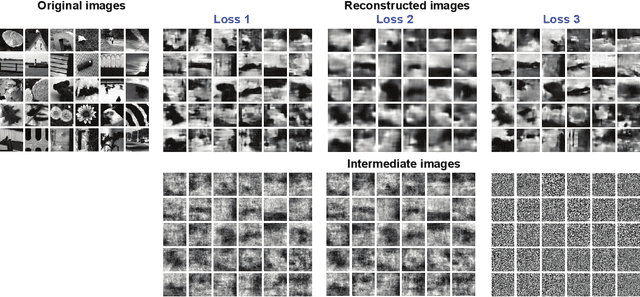


Abstract:Neural coding is one of the central questions in systems neuroscience for understanding how the brain processes stimulus from the environment, moreover, it is also a cornerstone for designing algorithms of brain-machine interface, where decoding incoming stimulus is needed for better performance of physical devices. Traditionally, the neural signal of interest for decoding visual scenes has been focused on fMRI data. However, our visual perception operates in a fast time scale of millisecond in terms of an event termed neural spike. So far there are few studies of decoding by using spikes. Here we fulfill this aim by developing a novel decoding framework based on deep neural networks, named spike-image decoder (SID), for reconstructing natural visual scenes, including static images and dynamic videos, from experimentally recorded spikes of a population of retinal ganglion cells. The SID is an end-to-end decoder with one end as neural spikes and the other end as images, which can be trained directly such that visual scenes are reconstructed from spikes in a highly accurate fashion. In addition, we show that SID can be generalized to arbitrary images by using image datasets of MNIST, CIFAR10, and CIFAR100. Furthermore, with a pre-trained SID, one can decode any dynamic videos, with the aid of an encoder, to achieve real-time encoding and decoding visual scenes by spikes. Altogether, our results shed new light on neuromorphic computing for artificial visual systems, such as event-based visual cameras and visual neuroprostheses.
Probabilistic Inference of Binary Markov Random Fields in Spiking Neural Networks through Mean-field Approximation
Feb 22, 2019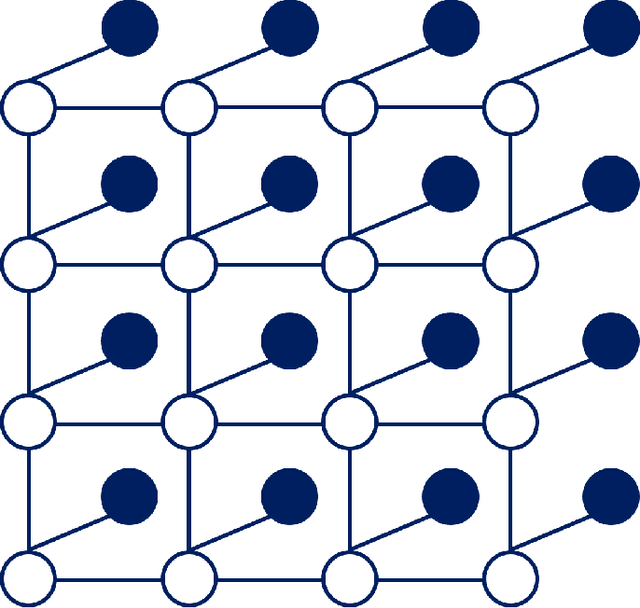
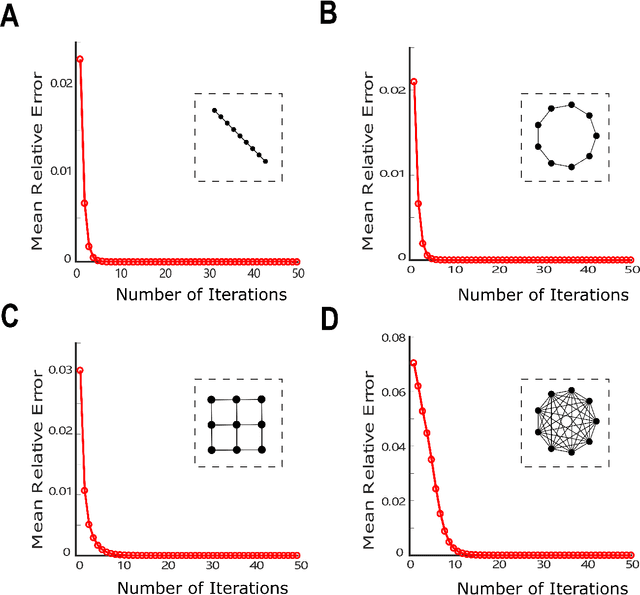
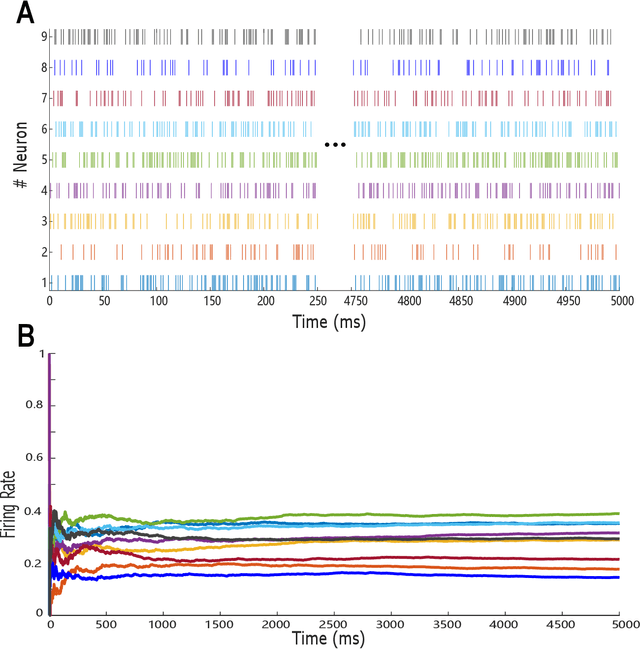
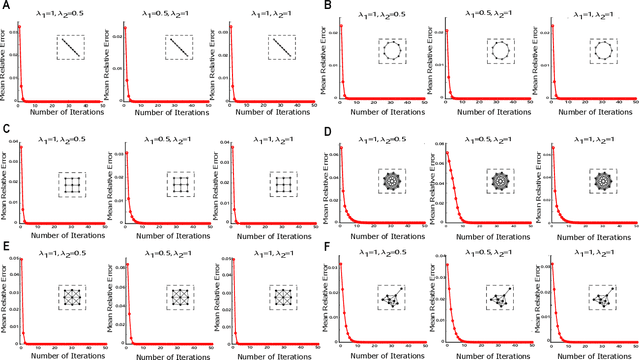
Abstract:Recent studies have suggested that the cognitive process of the human brain is realized as probabilistic inference and can be further modeled by probabilistic graphical models like Markov random fields. Nevertheless, it remains unclear how probabilistic inference can be implemented by a network of spiking neurons in the brain. Previous studies tried to relate the inference equation of binary Markov random fields to the dynamic equation of spiking neural networks through belief propagation algorithm and reparameterization, but they are valid only for Markov random fields with limited network structure. In this paper, we propose a spiking neural network model that can implement inference of arbitrary binary Markov random fields. Specifically, we design a spiking recurrent neural network and prove that its neuronal dynamics are mathematically equivalent to the inference process of Markov random fields by adopting mean-field theory. Theoretical analysis and experimental results, together with the application
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge