Jessica Schrouff
Evaluating Model Bias Requires Characterizing its Mistakes
Jul 15, 2024



Abstract:The ability to properly benchmark model performance in the face of spurious correlations is important to both build better predictors and increase confidence that models are operating as intended. We demonstrate that characterizing (as opposed to simply quantifying) model mistakes across subgroups is pivotal to properly reflect model biases, which are ignored by standard metrics such as worst-group accuracy or accuracy gap. Inspired by the hypothesis testing framework, we introduce SkewSize, a principled and flexible metric that captures bias from mistakes in a model's predictions. It can be used in multi-class settings or generalised to the open vocabulary setting of generative models. SkewSize is an aggregation of the effect size of the interaction between two categorical variables: the spurious variable representing the bias attribute and the model's prediction. We demonstrate the utility of SkewSize in multiple settings including: standard vision models trained on synthetic data, vision models trained on ImageNet, and large scale vision-and-language models from the BLIP-2 family. In each case, the proposed SkewSize is able to highlight biases not captured by other metrics, while also providing insights on the impact of recently proposed techniques, such as instruction tuning.
Mind the Graph When Balancing Data for Fairness or Robustness
Jun 25, 2024



Abstract:Failures of fairness or robustness in machine learning predictive settings can be due to undesired dependencies between covariates, outcomes and auxiliary factors of variation. A common strategy to mitigate these failures is data balancing, which attempts to remove those undesired dependencies. In this work, we define conditions on the training distribution for data balancing to lead to fair or robust models. Our results display that, in many cases, the balanced distribution does not correspond to selectively removing the undesired dependencies in a causal graph of the task, leading to multiple failure modes and even interference with other mitigation techniques such as regularization. Overall, our results highlight the importance of taking the causal graph into account before performing data balancing.
FunBO: Discovering Acquisition Functions for Bayesian Optimization with FunSearch
Jun 07, 2024



Abstract:The sample efficiency of Bayesian optimization algorithms depends on carefully crafted acquisition functions (AFs) guiding the sequential collection of function evaluations. The best-performing AF can vary significantly across optimization problems, often requiring ad-hoc and problem-specific choices. This work tackles the challenge of designing novel AFs that perform well across a variety of experimental settings. Based on FunSearch, a recent work using Large Language Models (LLMs) for discovery in mathematical sciences, we propose FunBO, an LLM-based method that can be used to learn new AFs written in computer code by leveraging access to a limited number of evaluations for a set of objective functions. We provide the analytic expression of all discovered AFs and evaluate them on various global optimization benchmarks and hyperparameter optimization tasks. We show how FunBO identifies AFs that generalize well in and out of the training distribution of functions, thus outperforming established general-purpose AFs and achieving competitive performance against AFs that are customized to specific function types and are learned via transfer-learning algorithms.
Adapting to Latent Subgroup Shifts via Concepts and Proxies
Dec 21, 2022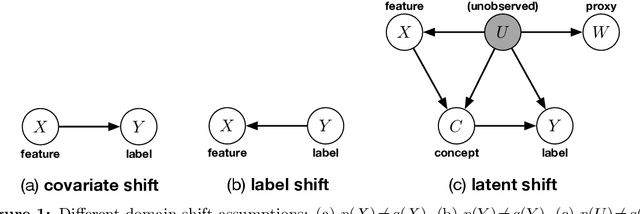

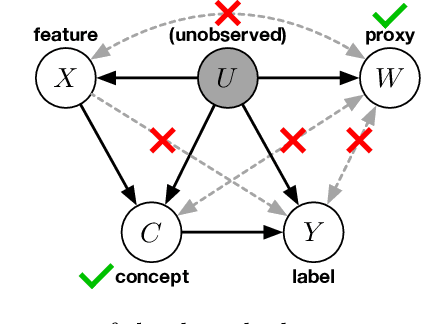
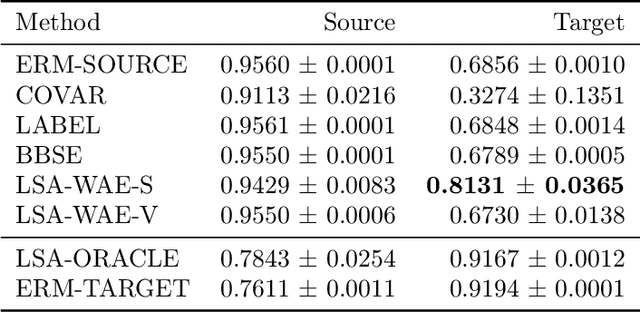
Abstract:We address the problem of unsupervised domain adaptation when the source domain differs from the target domain because of a shift in the distribution of a latent subgroup. When this subgroup confounds all observed data, neither covariate shift nor label shift assumptions apply. We show that the optimal target predictor can be non-parametrically identified with the help of concept and proxy variables available only in the source domain, and unlabeled data from the target. The identification results are constructive, immediately suggesting an algorithm for estimating the optimal predictor in the target. For continuous observations, when this algorithm becomes impractical, we propose a latent variable model specific to the data generation process at hand. We show how the approach degrades as the size of the shift changes, and verify that it outperforms both covariate and label shift adjustment.
Detecting and Preventing Shortcut Learning for Fair Medical AI using Shortcut Testing (ShorT)
Jul 21, 2022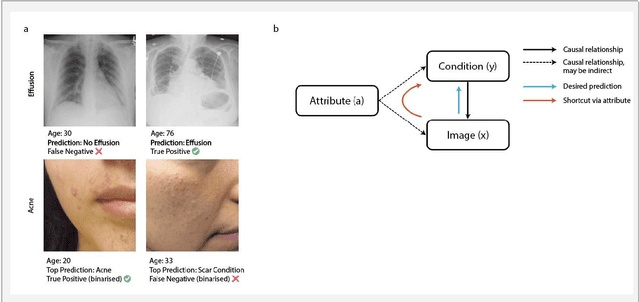
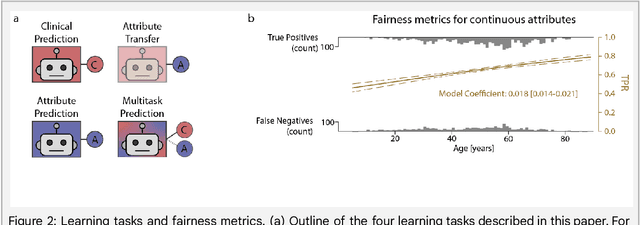
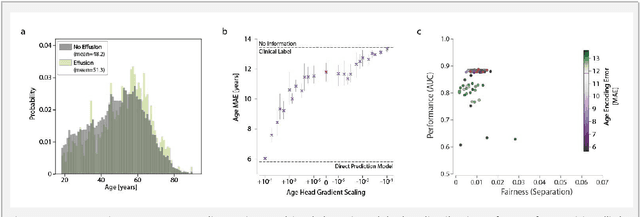
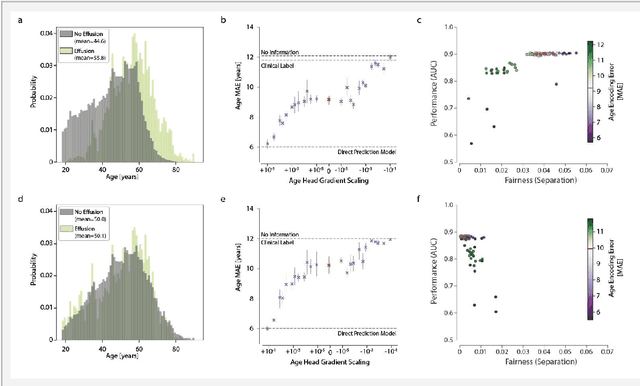
Abstract:Machine learning (ML) holds great promise for improving healthcare, but it is critical to ensure that its use will not propagate or amplify health disparities. An important step is to characterize the (un)fairness of ML models - their tendency to perform differently across subgroups of the population - and to understand its underlying mechanisms. One potential driver of algorithmic unfairness, shortcut learning, arises when ML models base predictions on improper correlations in the training data. However, diagnosing this phenomenon is difficult, especially when sensitive attributes are causally linked with disease. Using multi-task learning, we propose the first method to assess and mitigate shortcut learning as a part of the fairness assessment of clinical ML systems, and demonstrate its application to clinical tasks in radiology and dermatology. Finally, our approach reveals instances when shortcutting is not responsible for unfairness, highlighting the need for a holistic approach to fairness mitigation in medical AI.
A Reduction to Binary Approach for Debiasing Multiclass Datasets
May 31, 2022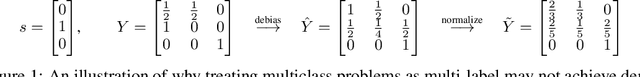
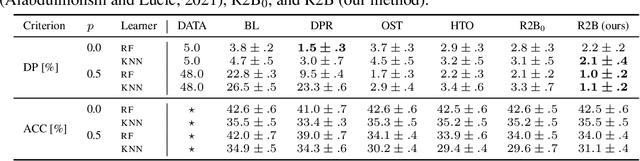
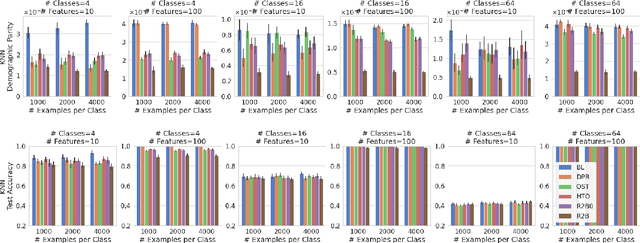
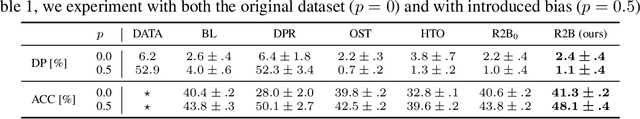
Abstract:We propose a novel reduction-to-binary (R2B) approach that enforces demographic parity for multiclass classification with non-binary sensitive attributes via a reduction to a sequence of binary debiasing tasks. We prove that R2B satisfies optimality and bias guarantees and demonstrate empirically that it can lead to an improvement over two baselines: (1) treating multiclass problems as multi-label by debiasing labels independently and (2) transforming the features instead of the labels. Surprisingly, we also demonstrate that independent label debiasing yields competitive results in most (but not all) settings. We validate these conclusions on synthetic and real-world datasets from social science, computer vision, and healthcare.
Disability prediction in multiple sclerosis using performance outcome measures and demographic data
Apr 08, 2022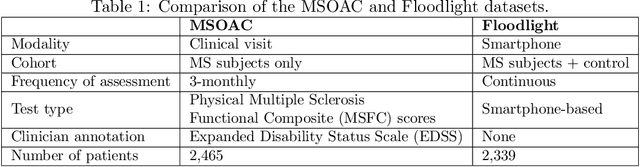
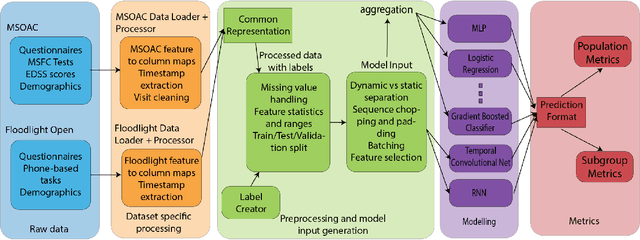
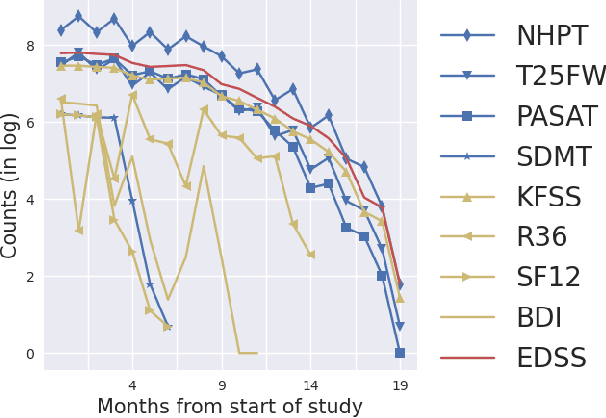
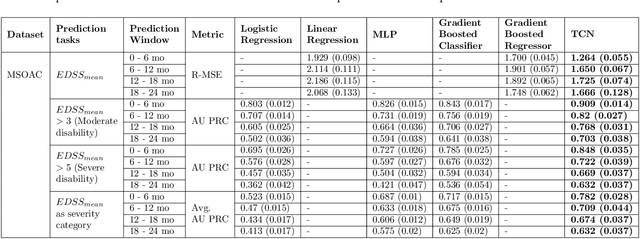
Abstract:Literature on machine learning for multiple sclerosis has primarily focused on the use of neuroimaging data such as magnetic resonance imaging and clinical laboratory tests for disease identification. However, studies have shown that these modalities are not consistent with disease activity such as symptoms or disease progression. Furthermore, the cost of collecting data from these modalities is high, leading to scarce evaluations. In this work, we used multi-dimensional, affordable, physical and smartphone-based performance outcome measures (POM) in conjunction with demographic data to predict multiple sclerosis disease progression. We performed a rigorous benchmarking exercise on two datasets and present results across 13 clinically actionable prediction endpoints and 6 machine learning models. To the best of our knowledge, our results are the first to show that it is possible to predict disease progression using POMs and demographic data in the context of both clinical trials and smartphone-base studies by using two datasets. Moreover, we investigate our models to understand the impact of different POMs and demographics on model performance through feature ablation studies. We also show that model performance is similar across different demographic subgroups (based on age and sex). To enable this work, we developed an end-to-end reusable pre-processing and machine learning framework which allows quicker experimentation over disparate MS datasets.
Healthsheet: Development of a Transparency Artifact for Health Datasets
Feb 26, 2022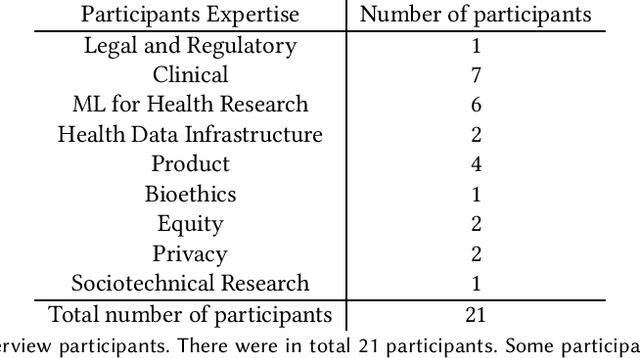
Abstract:Machine learning (ML) approaches have demonstrated promising results in a wide range of healthcare applications. Data plays a crucial role in developing ML-based healthcare systems that directly affect people's lives. Many of the ethical issues surrounding the use of ML in healthcare stem from structural inequalities underlying the way we collect, use, and handle data. Developing guidelines to improve documentation practices regarding the creation, use, and maintenance of ML healthcare datasets is therefore of critical importance. In this work, we introduce Healthsheet, a contextualized adaptation of the original datasheet questionnaire ~\cite{gebru2018datasheets} for health-specific applications. Through a series of semi-structured interviews, we adapt the datasheets for healthcare data documentation. As part of the Healthsheet development process and to understand the obstacles researchers face in creating datasheets, we worked with three publicly-available healthcare datasets as our case studies, each with different types of structured data: Electronic health Records (EHR), clinical trial study data, and smartphone-based performance outcome measures. Our findings from the interviewee study and case studies show 1) that datasheets should be contextualized for healthcare, 2) that despite incentives to adopt accountability practices such as datasheets, there is a lack of consistency in the broader use of these practices 3) how the ML for health community views datasheets and particularly \textit{Healthsheets} as diagnostic tool to surface the limitations and strength of datasets and 4) the relative importance of different fields in the datasheet to healthcare concerns.
Maintaining fairness across distribution shift: do we have viable solutions for real-world applications?
Feb 02, 2022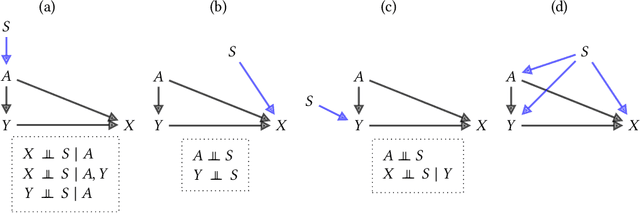
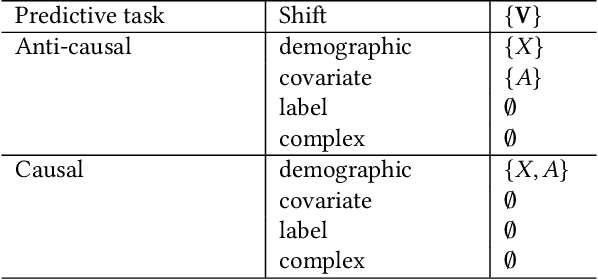
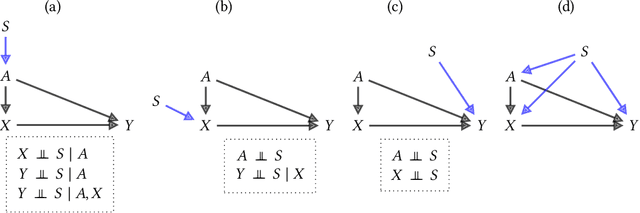
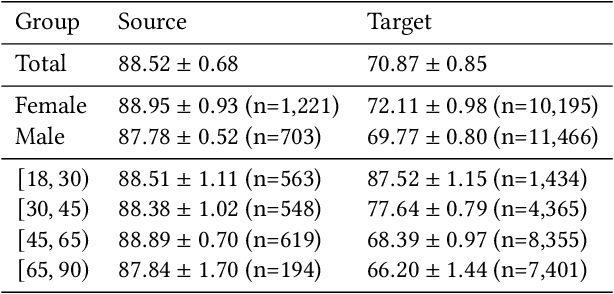
Abstract:Fairness and robustness are often considered as orthogonal dimensions when evaluating machine learning models. However, recent work has revealed interactions between fairness and robustness, showing that fairness properties are not necessarily maintained under distribution shift. In healthcare settings, this can result in e.g. a model that performs fairly according to a selected metric in "hospital A" showing unfairness when deployed in "hospital B". While a nascent field has emerged to develop provable fair and robust models, it typically relies on strong assumptions about the shift, limiting its impact for real-world applications. In this work, we explore the settings in which recently proposed mitigation strategies are applicable by referring to a causal framing. Using examples of predictive models in dermatology and electronic health records, we show that real-world applications are complex and often invalidate the assumptions of such methods. Our work hence highlights technical, practical, and engineering gaps that prevent the development of robustly fair machine learning models for real-world applications. Finally, we discuss potential remedies at each step of the machine learning pipeline.
Best of both worlds: local and global explanations with human-understandable concepts
Jun 16, 2021



Abstract:Interpretability techniques aim to provide the rationale behind a model's decision, typically by explaining either an individual prediction (local explanation, e.g. `why is this patient diagnosed with this condition') or a class of predictions (global explanation, e.g. `why are patients diagnosed with this condition in general'). While there are many methods focused on either one, few frameworks can provide both local and global explanations in a consistent manner. In this work, we combine two powerful existing techniques, one local (Integrated Gradients, IG) and one global (Testing with Concept Activation Vectors), to provide local, and global concept-based explanations. We first validate our idea using two synthetic datasets with a known ground truth, and further demonstrate with a benchmark natural image dataset. We test our method with various concepts, target classes, model architectures and IG baselines. We show that our method improves global explanations over TCAV when compared to ground truth, and provides useful insights. We hope our work provides a step towards building bridges between many existing local and global methods to get the best of both worlds.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge