Eric P. Xing
Personalized Survival Prediction with Contextual Explanation Networks
Jan 30, 2018
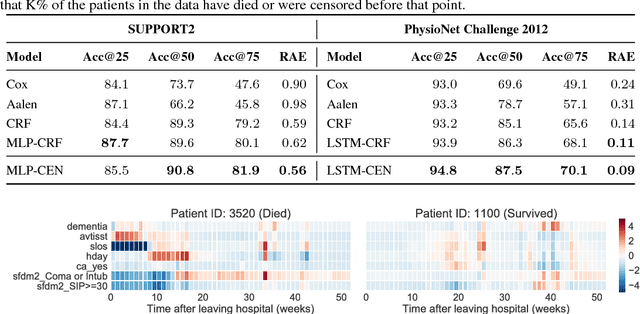
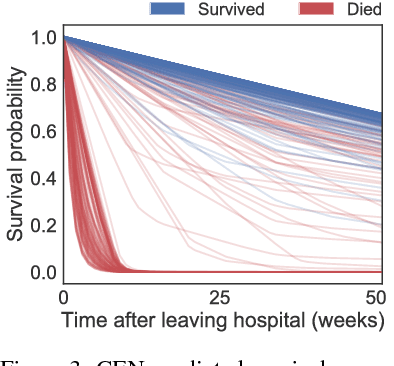
Abstract:Accurate and transparent prediction of cancer survival times on the level of individual patients can inform and improve patient care and treatment practices. In this paper, we design a model that concurrently learns to accurately predict patient-specific survival distributions and to explain its predictions in terms of patient attributes such as clinical tests or assessments. Our model is flexible and based on a recurrent network, can handle various modalities of data including temporal measurements, and yet constructs and uses simple explanations in the form of patient- and time-specific linear regression. For analysis, we use two publicly available datasets and show that our networks outperform a number of baselines in prediction while providing a way to inspect the reasons behind each prediction.
The Intriguing Properties of Model Explanations
Jan 30, 2018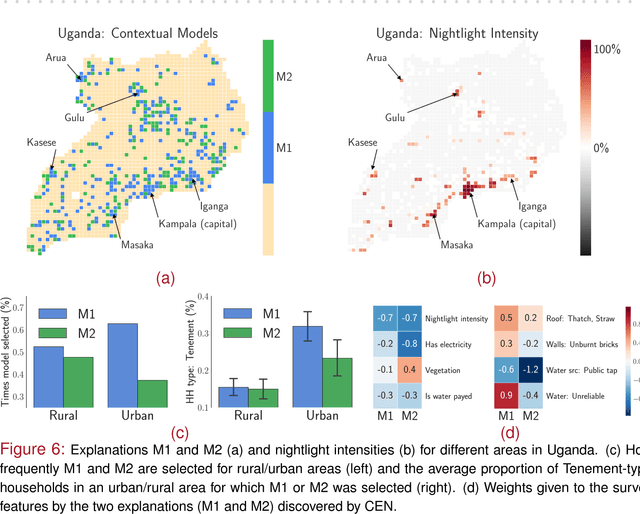
Abstract:Linear approximations to the decision boundary of a complex model have become one of the most popular tools for interpreting predictions. In this paper, we study such linear explanations produced either post-hoc by a few recent methods or generated along with predictions with contextual explanation networks (CENs). We focus on two questions: (i) whether linear explanations are always consistent or can be misleading, and (ii) when integrated into the prediction process, whether and how explanations affect the performance of the model. Our analysis sheds more light on certain properties of explanations produced by different methods and suggests that learning models that explain and predict jointly is often advantageous.
Contextual Explanation Networks
Jan 30, 2018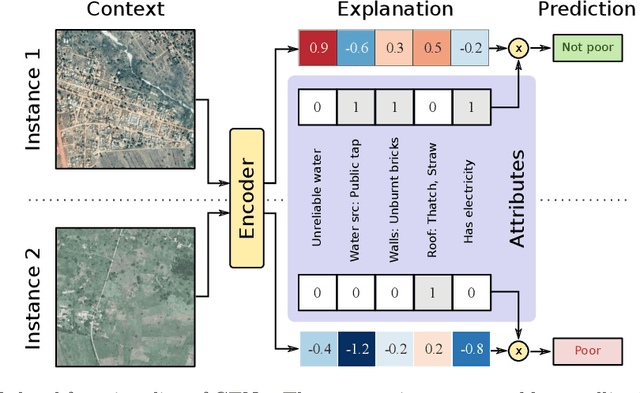

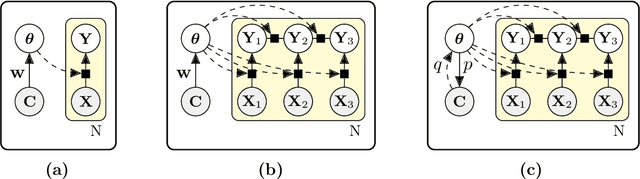
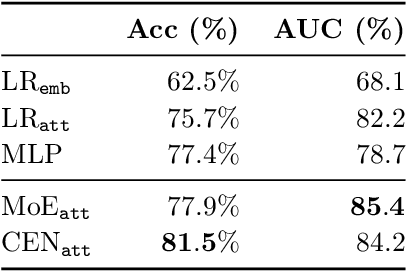
Abstract:We introduce contextual explanation networks (CENs)---a class of models that learn to predict by generating and leveraging intermediate explanations. CENs are deep networks that generate parameters for context-specific probabilistic graphical models which are further used for prediction and play the role of explanations. Contrary to the existing post-hoc model-explanation tools, CENs learn to predict and to explain jointly. Our approach offers two major advantages: (i) for each prediction, valid instance-specific explanations are generated with no computational overhead and (ii) prediction via explanation acts as a regularization and boosts performance in low-resource settings. We prove that local approximations to the decision boundary of our networks are consistent with the generated explanations. Our results on image and text classification and survival analysis tasks demonstrate that CENs are competitive with the state-of-the-art while offering additional insights behind each prediction, valuable for decision support.
Stability Selection for Structured Variable Selection
Dec 13, 2017
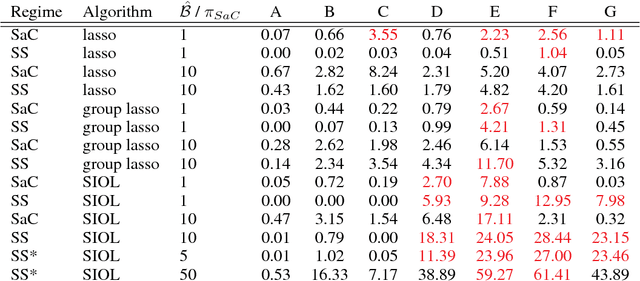

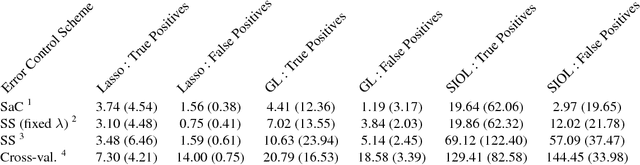
Abstract:In variable or graph selection problems, finding a right-sized model or controlling the number of false positives is notoriously difficult. Recently, a meta-algorithm called Stability Selection was proposed that can provide reliable finite-sample control of the number of false positives. Its benefits were demonstrated when used in conjunction with the lasso and orthogonal matching pursuit algorithms. In this paper, we investigate the applicability of stability selection to structured selection algorithms: the group lasso and the structured input-output lasso. We find that using stability selection often increases the power of both algorithms, but that the presence of complex structure reduces the reliability of error control under stability selection. We give strategies for setting tuning parameters to obtain a good model size under stability selection, and highlight its strengths and weaknesses compared to competing methods screen and clean and cross-validation. We give guidelines about when to use which error control method.
Cavs: A Vertex-centric Programming Interface for Dynamic Neural Networks
Dec 11, 2017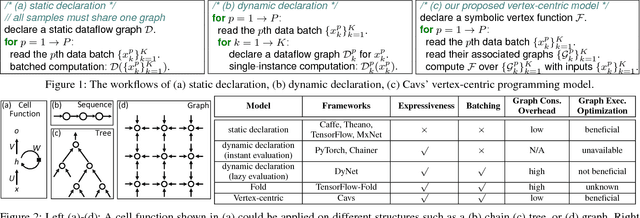
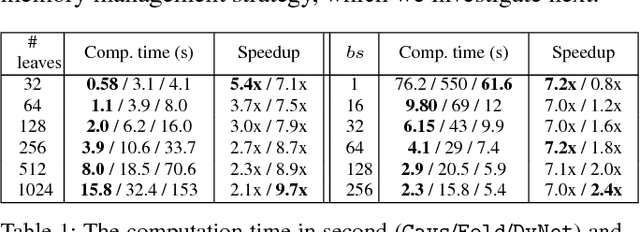
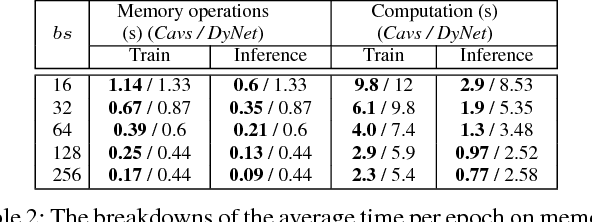
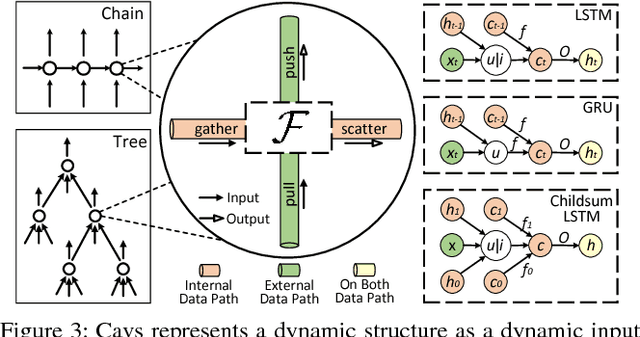
Abstract:Recent deep learning (DL) models have moved beyond static network architectures to dynamic ones, handling data where the network structure changes every example, such as sequences of variable lengths, trees, and graphs. Existing dataflow-based programming models for DL---both static and dynamic declaration---either cannot readily express these dynamic models, or are inefficient due to repeated dataflow graph construction and processing, and difficulties in batched execution. We present Cavs, a vertex-centric programming interface and optimized system implementation for dynamic DL models. Cavs represents dynamic network structure as a static vertex function $\mathcal{F}$ and a dynamic instance-specific graph $\mathcal{G}$, and performs backpropagation by scheduling the execution of $\mathcal{F}$ following the dependencies in $\mathcal{G}$. Cavs bypasses expensive graph construction and preprocessing overhead, allows for the use of static graph optimization techniques on pre-defined operations in $\mathcal{F}$, and naturally exposes batched execution opportunities over different graphs. Experiments comparing Cavs to two state-of-the-art frameworks for dynamic NNs (TensorFlow Fold and DyNet) demonstrate the efficacy of this approach: Cavs achieves a near one order of magnitude speedup on training of various dynamic NN architectures, and ablations demonstrate the contribution of our proposed batching and memory management strategies.
Towards Automated ICD Coding Using Deep Learning
Nov 30, 2017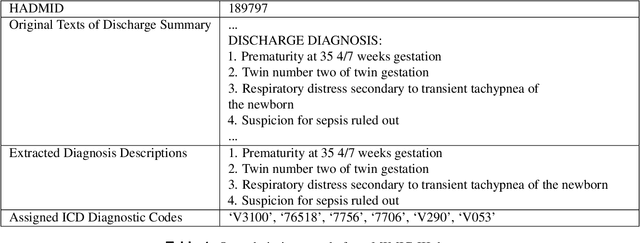
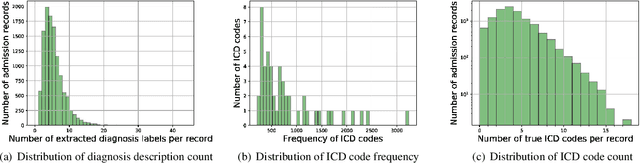
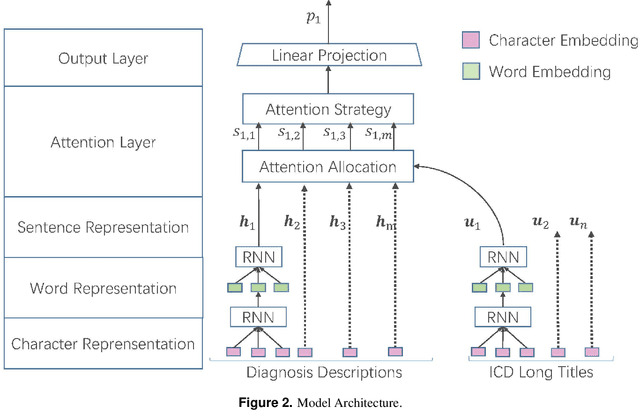
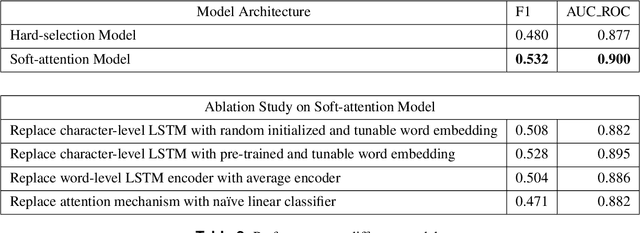
Abstract:International Classification of Diseases(ICD) is an authoritative health care classification system of different diseases and conditions for clinical and management purposes. Considering the complicated and dedicated process to assign correct codes to each patient admission based on overall diagnosis, we propose a hierarchical deep learning model with attention mechanism which can automatically assign ICD diagnostic codes given written diagnosis. We utilize character-aware neural language models to generate hidden representations of written diagnosis descriptions and ICD codes, and design an attention mechanism to address the mismatch between the numbers of descriptions and corresponding codes. Our experimental results show the strong potential of automated ICD coding from diagnosis descriptions. Our best model achieves 0.53 and 0.90 of F1 score and area under curve of receiver operating characteristic respectively. The result outperforms those achieved using character-unaware encoding method or without attention mechanism. It indicates that our proposed deep learning model can code automatically in a reasonable way and provide a framework for computer-auxiliary ICD coding.
Learning Less-Overlapping Representations
Nov 25, 2017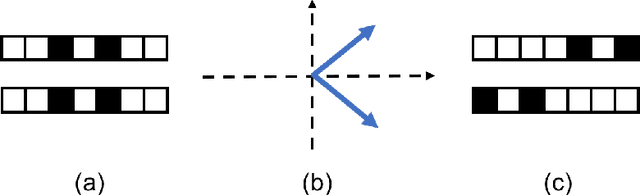
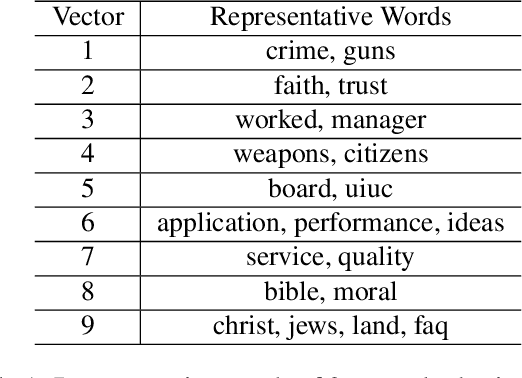
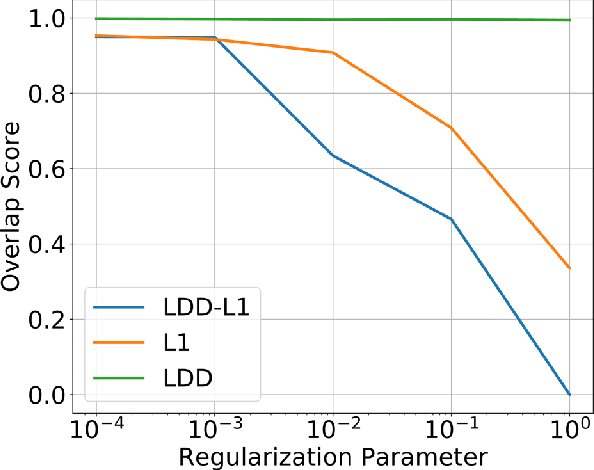
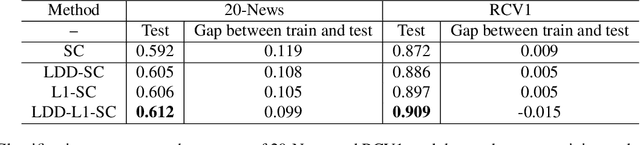
Abstract:In representation learning (RL), how to make the learned representations easy to interpret and less overfitted to training data are two important but challenging issues. To address these problems, we study a new type of regulariza- tion approach that encourages the supports of weight vectors in RL models to have small overlap, by simultaneously promoting near-orthogonality among vectors and sparsity of each vector. We apply the proposed regularizer to two models: neural networks (NNs) and sparse coding (SC), and develop an efficient ADMM-based algorithm for regu- larized SC. Experiments on various datasets demonstrate that weight vectors learned under our regularizer are more interpretable and have better generalization performance.
Diversity-Promoting Bayesian Learning of Latent Variable Models
Nov 23, 2017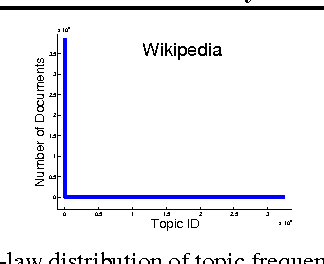
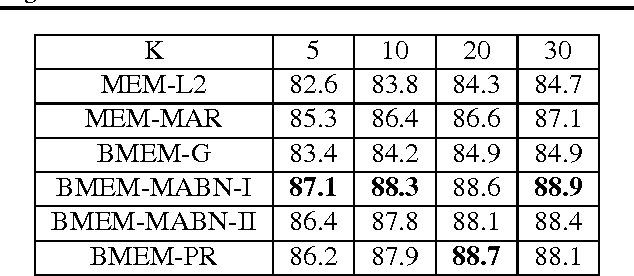
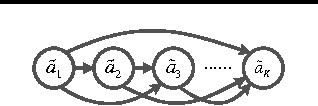
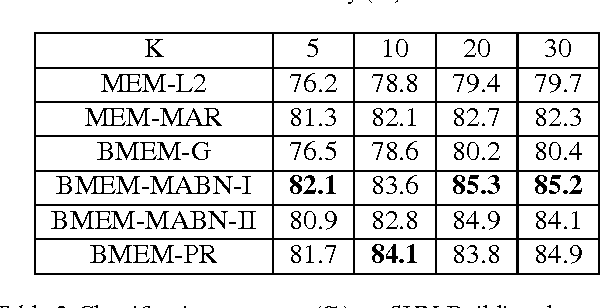
Abstract:To address three important issues involved in latent variable models (LVMs), including capturing infrequent patterns, achieving small-sized but expressive models and alleviating overfitting, several studies have been devoted to "diversifying" LVMs, which aim at encouraging the components in LVMs to be diverse. Most existing studies fall into a frequentist-style regularization framework, where the components are learned via point estimation. In this paper, we investigate how to "diversify" LVMs in the paradigm of Bayesian learning. We propose two approaches that have complementary advantages. One is to define a diversity-promoting mutual angular prior which assigns larger density to components with larger mutual angles and use this prior to affect the posterior via Bayes' rule. We develop two efficient approximate posterior inference algorithms based on variational inference and MCMC sampling. The other approach is to impose diversity-promoting regularization directly over the post-data distribution of components. We also extend our approach to "diversify" Bayesian nonparametric models where the number of components is infinite. A sampling algorithm based on slice sampling and Hamiltonian Monte Carlo is developed. We apply these methods to "diversify" Bayesian mixture of experts model and infinite latent feature model. Experiments on various datasets demonstrate the effectiveness and efficiency of our methods.
Medical Diagnosis From Laboratory Tests by Combining Generative and Discriminative Learning
Nov 16, 2017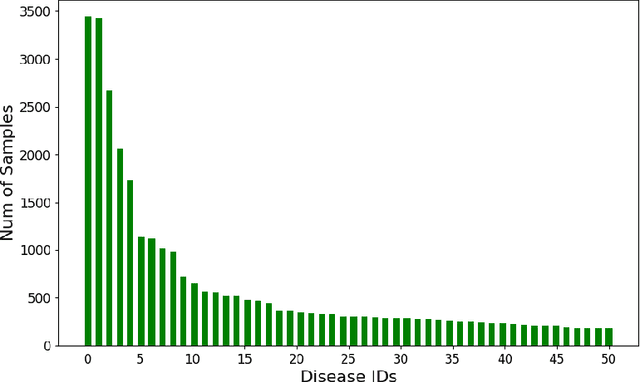
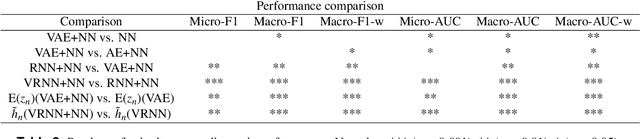
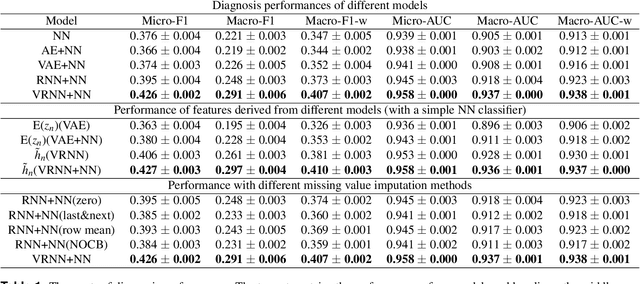
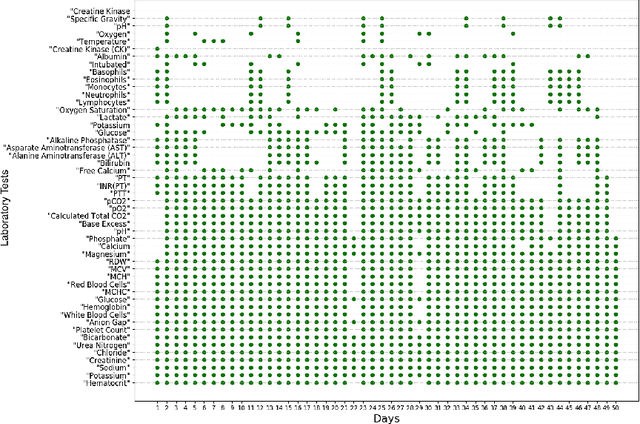

Abstract:A primary goal of computational phenotype research is to conduct medical diagnosis. In hospital, physicians rely on massive clinical data to make diagnosis decisions, among which laboratory tests are one of the most important resources. However, the longitudinal and incomplete nature of laboratory test data casts a significant challenge on its interpretation and usage, which may result in harmful decisions by both human physicians and automatic diagnosis systems. In this work, we take advantage of deep generative models to deal with the complex laboratory tests. Specifically, we propose an end-to-end architecture that involves a deep generative variational recurrent neural networks (VRNN) to learn robust and generalizable features, and a discriminative neural network (NN) model to learn diagnosis decision making, and the two models are trained jointly. Our experiments are conducted on a dataset involving 46,252 patients, and the 50 most frequent tests are used to predict the 50 most common diagnoses. The results show that our model, VRNN+NN, significantly (p<0.001) outperforms other baseline models. Moreover, we demonstrate that the representations learned by the joint training are more informative than those learned by pure generative models. Finally, we find that our model offers a surprisingly good imputation for missing values.
A Sparse Graph-Structured Lasso Mixed Model for Genetic Association with Confounding Correction
Nov 11, 2017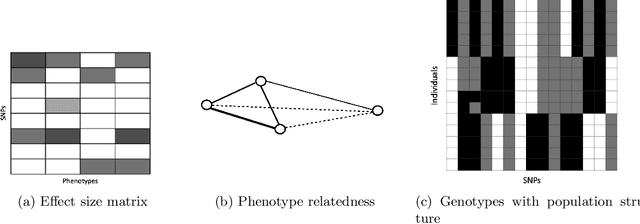
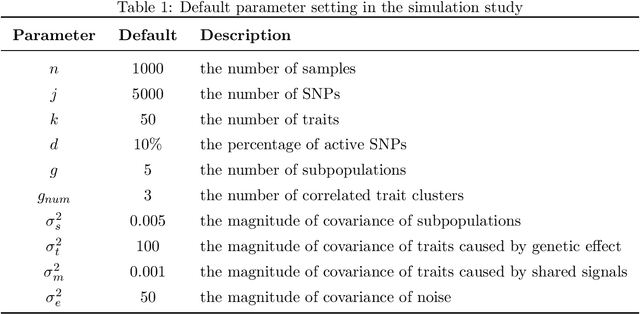

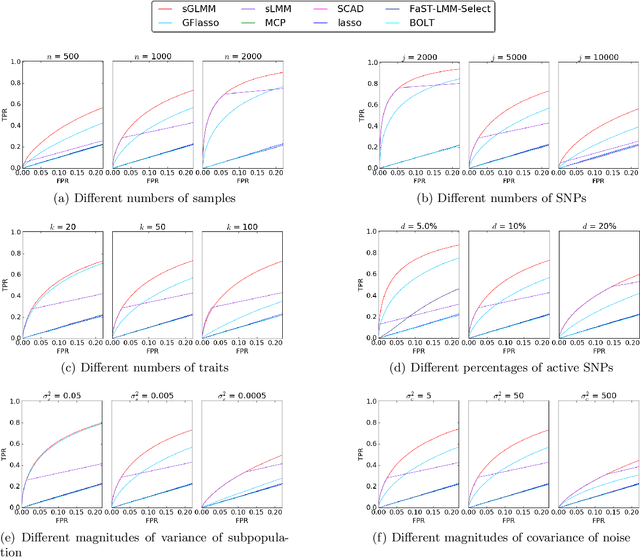
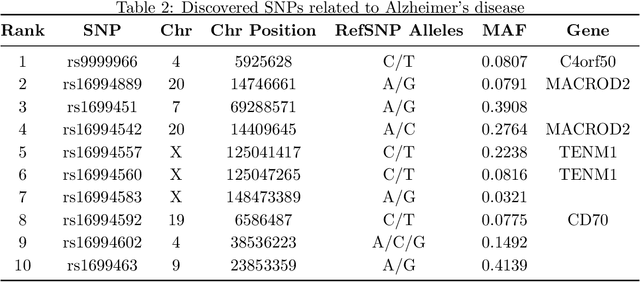
Abstract:While linear mixed model (LMM) has shown a competitive performance in correcting spurious associations raised by population stratification, family structures, and cryptic relatedness, more challenges are still to be addressed regarding the complex structure of genotypic and phenotypic data. For example, geneticists have discovered that some clusters of phenotypes are more co-expressed than others. Hence, a joint analysis that can utilize such relatedness information in a heterogeneous data set is crucial for genetic modeling. We proposed the sparse graph-structured linear mixed model (sGLMM) that can incorporate the relatedness information from traits in a dataset with confounding correction. Our method is capable of uncovering the genetic associations of a large number of phenotypes together while considering the relatedness of these phenotypes. Through extensive simulation experiments, we show that the proposed model outperforms other existing approaches and can model correlation from both population structure and shared signals. Further, we validate the effectiveness of sGLMM in the real-world genomic dataset on two different species from plants and humans. In Arabidopsis thaliana data, sGLMM behaves better than all other baseline models for 63.4% traits. We also discuss the potential causal genetic variation of Human Alzheimer's disease discovered by our model and justify some of the most important genetic loci.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge