Di He
Rethinking Lipschitz Neural Networks for Certified L-infinity Robustness
Oct 04, 2022
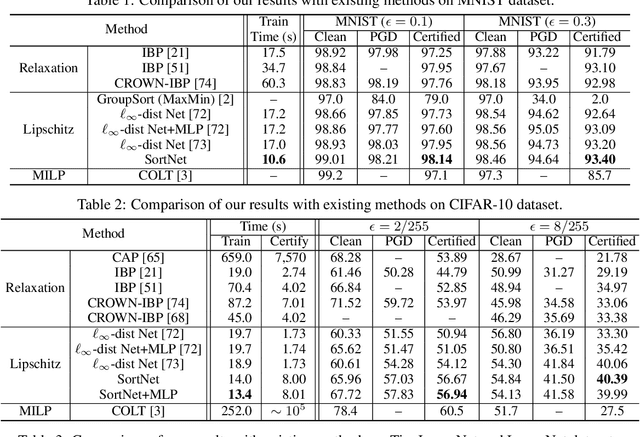
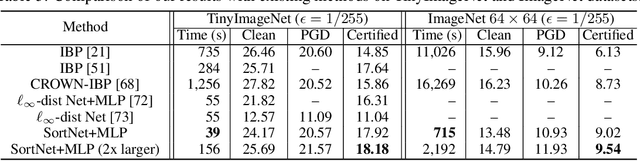
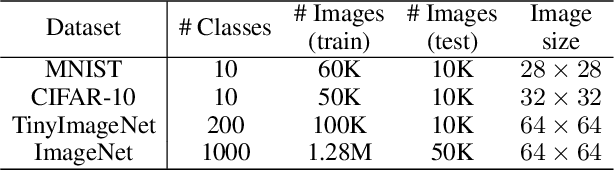
Abstract:Designing neural networks with bounded Lipschitz constant is a promising way to obtain certifiably robust classifiers against adversarial examples. However, the relevant progress for the important $\ell_\infty$ perturbation setting is rather limited, and a principled understanding of how to design expressive $\ell_\infty$ Lipschitz networks is still lacking. In this paper, we bridge the gap by studying certified $\ell_\infty$ robustness from a novel perspective of representing Boolean functions. We derive two fundamental impossibility results that hold for any standard Lipschitz network: one for robust classification on finite datasets, and the other for Lipschitz function approximation. These results identify that networks built upon norm-bounded affine layers and Lipschitz activations intrinsically lose expressive power even in the two-dimensional case, and shed light on how recently proposed Lipschitz networks (e.g., GroupSort and $\ell_\infty$-distance nets) bypass these impossibilities by leveraging order statistic functions. Finally, based on these insights, we develop a unified Lipschitz network that generalizes prior works, and design a practical version that can be efficiently trained (making certified robust training free). Extensive experiments show that our approach is scalable, efficient, and consistently yields better certified robustness across multiple datasets and perturbation radii than prior Lipschitz networks.
Adversarial Noises Are Linearly Separable for (Nearly) Random Neural Networks
Jun 09, 2022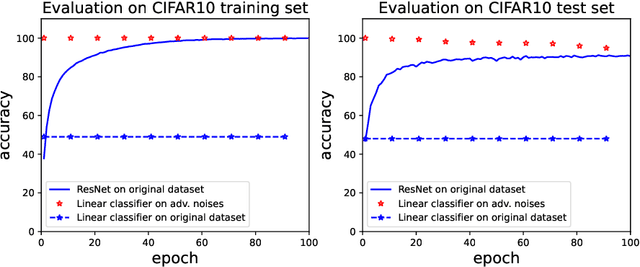
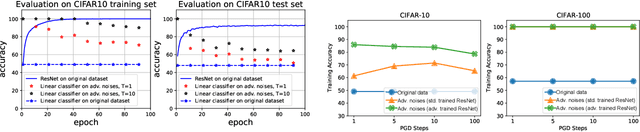
Abstract:Adversarial examples, which are usually generated for specific inputs with a specific model, are ubiquitous for neural networks. In this paper we unveil a surprising property of adversarial noises when they are put together, i.e., adversarial noises crafted by one-step gradient methods are linearly separable if equipped with the corresponding labels. We theoretically prove this property for a two-layer network with randomly initialized entries and the neural tangent kernel setup where the parameters are not far from initialization. The proof idea is to show the label information can be efficiently backpropagated to the input while keeping the linear separability. Our theory and experimental evidence further show that the linear classifier trained with the adversarial noises of the training data can well classify the adversarial noises of the test data, indicating that adversarial noises actually inject a distributional perturbation to the original data distribution. Furthermore, we empirically demonstrate that the adversarial noises may become less linearly separable when the above conditions are compromised while they are still much easier to classify than original features.
Is $L^2$ Physics-Informed Loss Always Suitable for Training Physics-Informed Neural Network?
Jun 04, 2022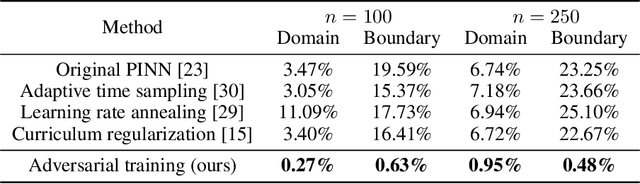
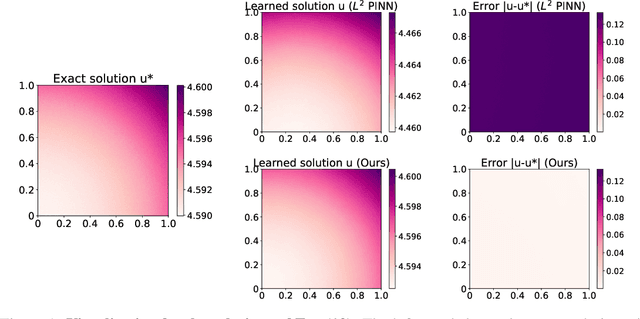
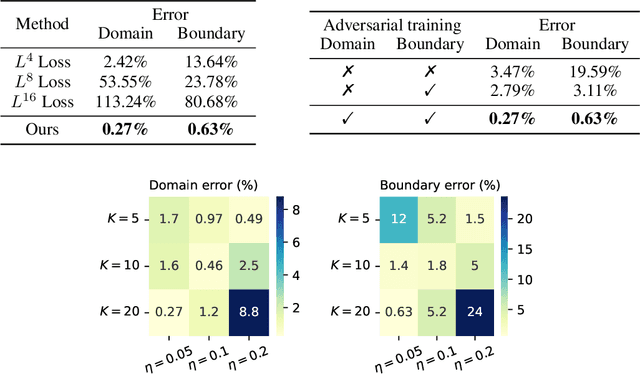
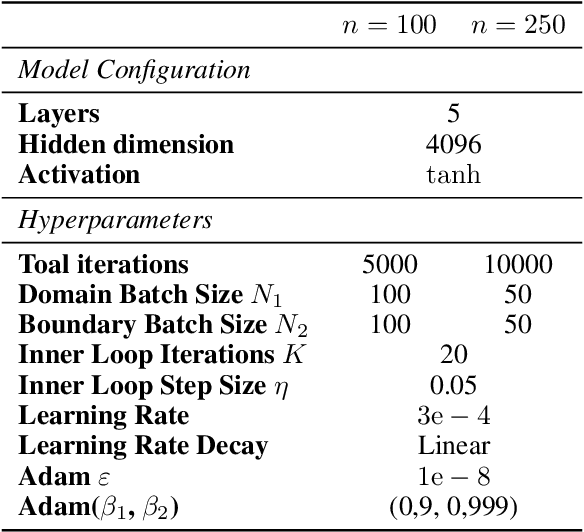
Abstract:The Physics-Informed Neural Network (PINN) approach is a new and promising way to solve partial differential equations using deep learning. The $L^2$ Physics-Informed Loss is the de-facto standard in training Physics-Informed Neural Networks. In this paper, we challenge this common practice by investigating the relationship between the loss function and the approximation quality of the learned solution. In particular, we leverage the concept of stability in the literature of partial differential equation to study the asymptotic behavior of the learned solution as the loss approaches zero. With this concept, we study an important class of high-dimensional non-linear PDEs in optimal control, the Hamilton-Jacobi-Bellman(HJB) Equation, and prove that for general $L^p$ Physics-Informed Loss, a wide class of HJB equation is stable only if $p$ is sufficiently large. Therefore, the commonly used $L^2$ loss is not suitable for training PINN on those equations, while $L^{\infty}$ loss is a better choice. Based on the theoretical insight, we develop a novel PINN training algorithm to minimize the $L^{\infty}$ loss for HJB equations which is in a similar spirit to adversarial training. The effectiveness of the proposed algorithm is empirically demonstrated through experiments.
Your Transformer May Not be as Powerful as You Expect
May 26, 2022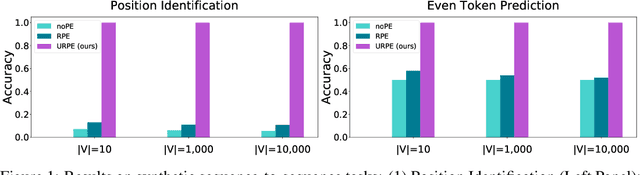
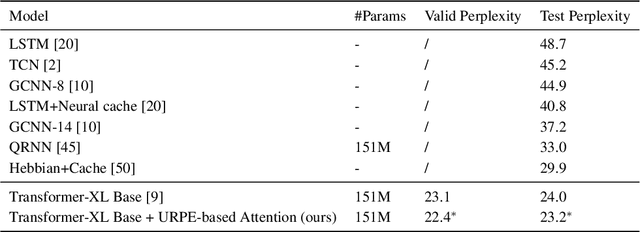
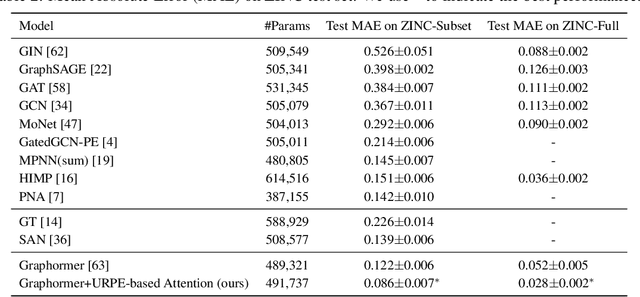
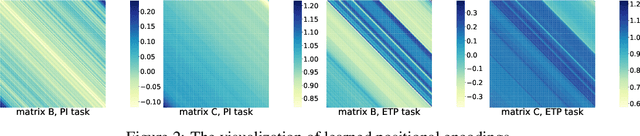
Abstract:Relative Positional Encoding (RPE), which encodes the relative distance between any pair of tokens, is one of the most successful modifications to the original Transformer. As far as we know, theoretical understanding of the RPE-based Transformers is largely unexplored. In this work, we mathematically analyze the power of RPE-based Transformers regarding whether the model is capable of approximating any continuous sequence-to-sequence functions. One may naturally assume the answer is in the affirmative -- RPE-based Transformers are universal function approximators. However, we present a negative result by showing there exist continuous sequence-to-sequence functions that RPE-based Transformers cannot approximate no matter how deep and wide the neural network is. One key reason lies in that most RPEs are placed in the softmax attention that always generates a right stochastic matrix. This restricts the network from capturing positional information in the RPEs and limits its capacity. To overcome the problem and make the model more powerful, we first present sufficient conditions for RPE-based Transformers to achieve universal function approximation. With the theoretical guidance, we develop a novel attention module, called Universal RPE-based (URPE) Attention, which satisfies the conditions. Therefore, the corresponding URPE-based Transformers become universal function approximators. Extensive experiments covering typical architectures and tasks demonstrate that our model is parameter-efficient and can achieve superior performance to strong baselines in a wide range of applications.
METRO: Efficient Denoising Pretraining of Large Scale Autoencoding Language Models with Model Generated Signals
Apr 16, 2022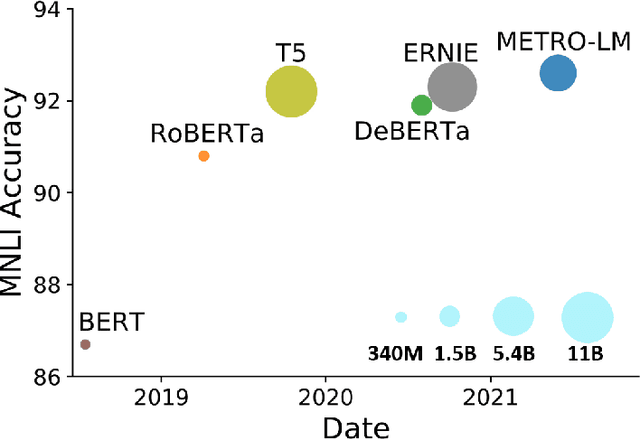
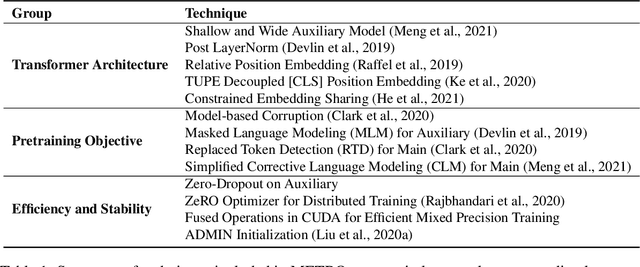
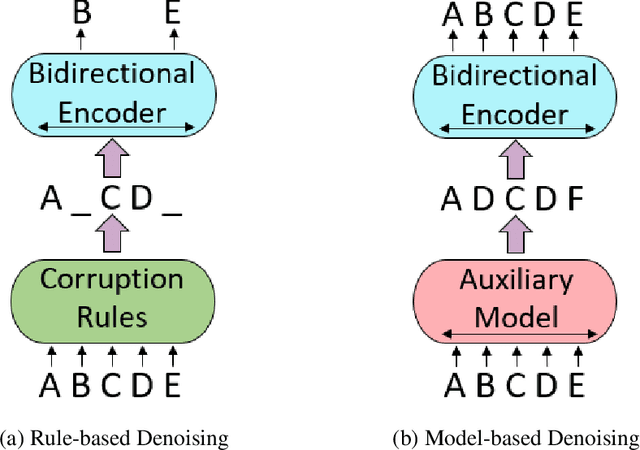
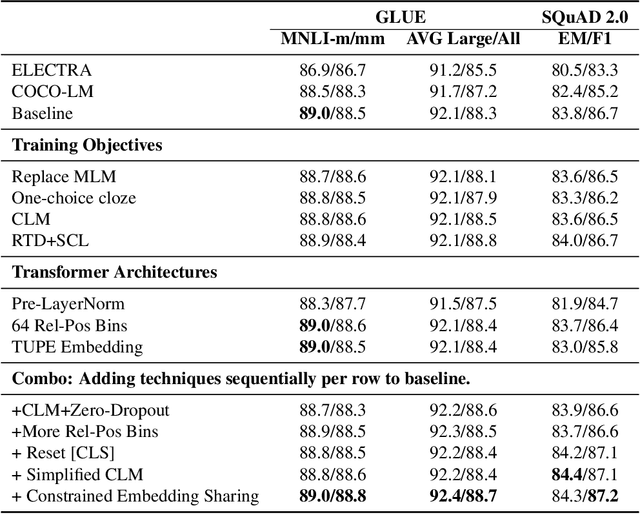
Abstract:We present an efficient method of pretraining large-scale autoencoding language models using training signals generated by an auxiliary model. Originated in ELECTRA, this training strategy has demonstrated sample-efficiency to pretrain models at the scale of hundreds of millions of parameters. In this work, we conduct a comprehensive empirical study, and propose a recipe, namely "Model generated dEnoising TRaining Objective" (METRO), which incorporates some of the best modeling techniques developed recently to speed up, stabilize, and enhance pretrained language models without compromising model effectiveness. The resultant models, METRO-LM, consisting of up to 5.4 billion parameters, achieve new state-of-the-art on the GLUE, SuperGLUE, and SQuAD benchmarks. More importantly, METRO-LM are efficient in that they often outperform previous large models with significantly smaller model sizes and lower pretraining cost.
An Empirical Study of Graphormer on Large-Scale Molecular Modeling Datasets
Mar 14, 2022
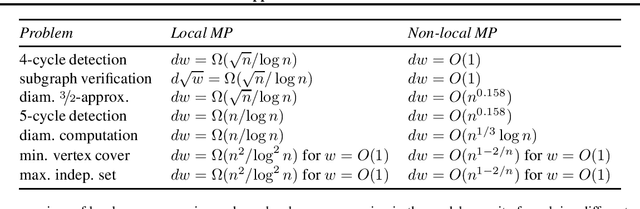
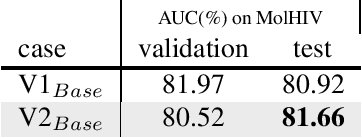
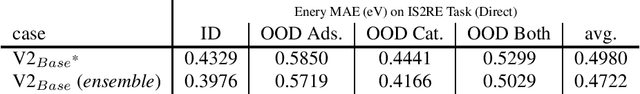

Abstract:This technical note describes the recent updates of Graphormer, including architecture design modifications, and the adaption to 3D molecular dynamics simulation. The "Graphormer-V2" could attain better results on large-scale molecular modeling datasets than the vanilla one, and the performance gain could be consistently obtained on downstream tasks. In addition, we show that with a global receptive field and an adaptive aggregation strategy, Graphormer is more powerful than classic message-passing-based GNNs. Graphormer-V2 achieves much less MAE than the vanilla Graphormer on the PCQM4M quantum chemistry dataset used in KDD Cup 2021, where the latter one won the first place in this competition. In the meanwhile, Graphormer-V2 greatly outperforms the competitors in the recent Open Catalyst Challenge, which is a competition track on NeurIPS 2021 workshop, and aims to model the catalyst-adsorbate reaction system with advanced AI models. All models could be found at \url{https://github.com/Microsoft/Graphormer}.
Benchmarking Graphormer on Large-Scale Molecular Modeling Datasets
Mar 09, 2022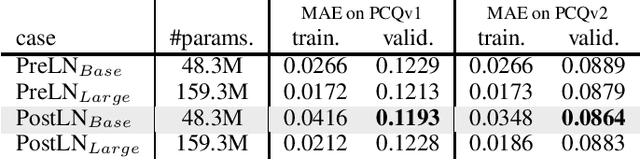
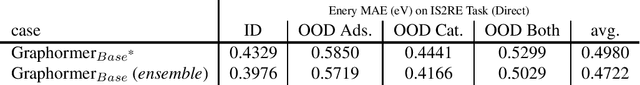
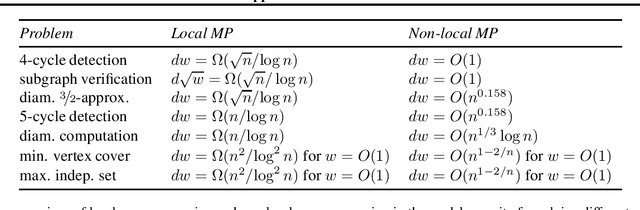
Abstract:This technical note describes the recent updates of Graphormer, including architecture design modifications, and the adaption to 3D molecular dynamics simulation. With these simple modifications, Graphormer could attain better results on large-scale molecular modeling datasets than the vanilla one, and the performance gain could be consistently obtained on 2D and 3D molecular graph modeling tasks. In addition, we show that with a global receptive field and an adaptive aggregation strategy, Graphormer is more powerful than classic message-passing-based GNNs. Empirically, Graphormer could achieve much less MAE than the originally reported results on the PCQM4M quantum chemistry dataset used in KDD Cup 2021. In the meanwhile, it greatly outperforms the competitors in the recent Open Catalyst Challenge, which is a competition track on NeurIPS 2021 workshop, and aims to model the catalyst-adsorbate reaction system with advanced AI models. All codes could be found at https://github.com/Microsoft/Graphormer.
VADOI:Voice-Activity-Detection Overlapping Inference For End-to-end Long-form Speech Recognition
Feb 22, 2022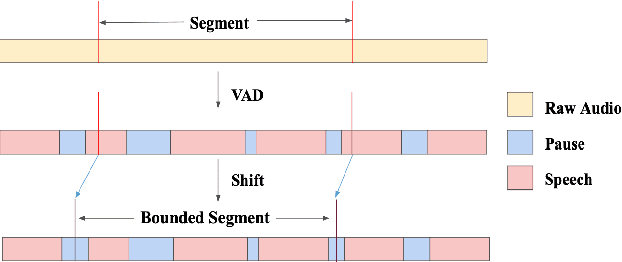
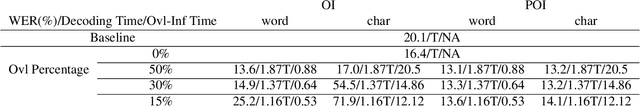
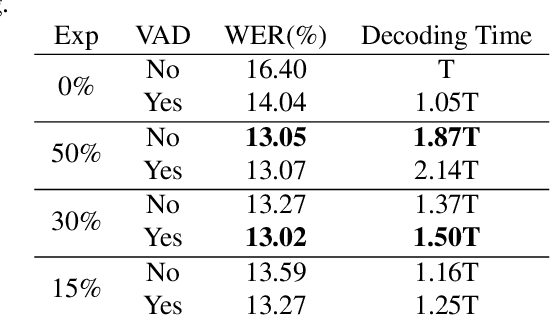
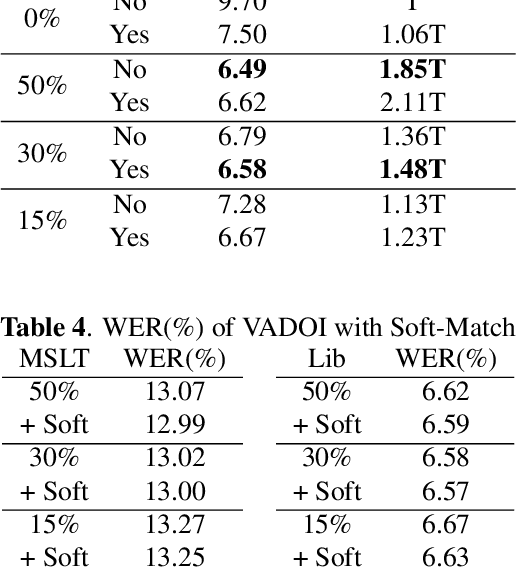
Abstract:While end-to-end models have shown great success on the Automatic Speech Recognition task, performance degrades severely when target sentences are long-form. The previous proposed methods, (partial) overlapping inference are shown to be effective on long-form decoding. For both methods, word error rate (WER) decreases monotonically when overlapping percentage decreases. Setting aside computational cost, the setup with 50% overlapping during inference can achieve the best performance. However, a lower overlapping percentage has an advantage of fast inference speed. In this paper, we first conduct comprehensive experiments comparing overlapping inference and partial overlapping inference with various configurations. We then propose Voice-Activity-Detection Overlapping Inference to provide a trade-off between WER and computation cost. Results show that the proposed method can achieve a 20% relative computation cost reduction on Librispeech and Microsoft Speech Language Translation long-form corpus while maintaining the WER performance when comparing to the best performing overlapping inference algorithm. We also propose Soft-Match to compensate for similar words mis-aligned problem.
Learning Physics-Informed Neural Networks without Stacked Back-propagation
Feb 18, 2022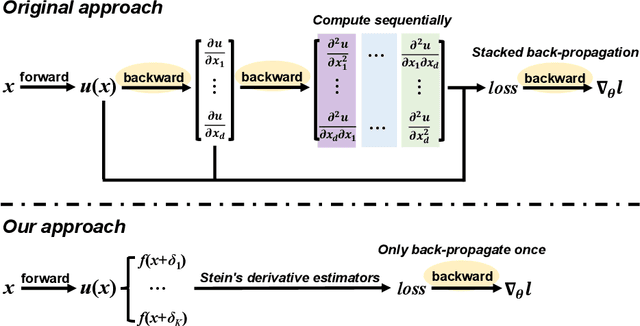
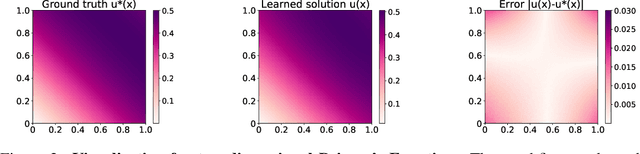
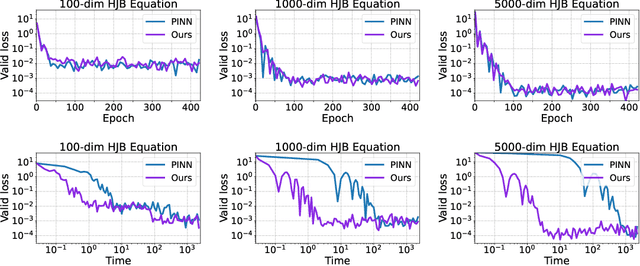
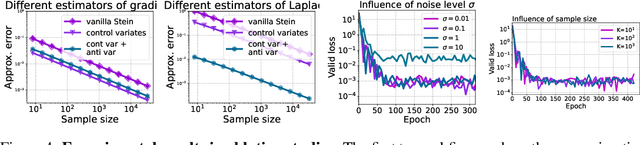
Abstract:Physics-Informed Neural Network (PINN) has become a commonly used machine learning approach to solve partial differential equations (PDE). But, facing high-dimensional second-order PDE problems, PINN will suffer from severe scalability issues since its loss includes second-order derivatives, the computational cost of which will grow along with the dimension during stacked back-propagation. In this paper, we develop a novel approach that can significantly accelerate the training of Physics-Informed Neural Networks. In particular, we parameterize the PDE solution by the Gaussian smoothed model and show that, derived from Stein's Identity, the second-order derivatives can be efficiently calculated without back-propagation. We further discuss the model capacity and provide variance reduction methods to address key limitations in the derivative estimation. Experimental results show that our proposed method can achieve competitive error compared to standard PINN training but is two orders of magnitude faster.
HousE: Knowledge Graph Embedding with Householder Parameterization
Feb 16, 2022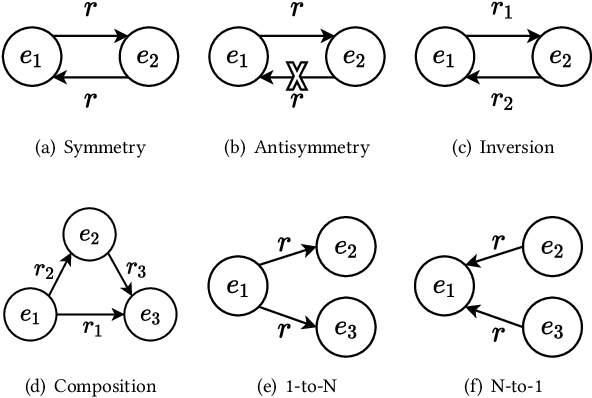
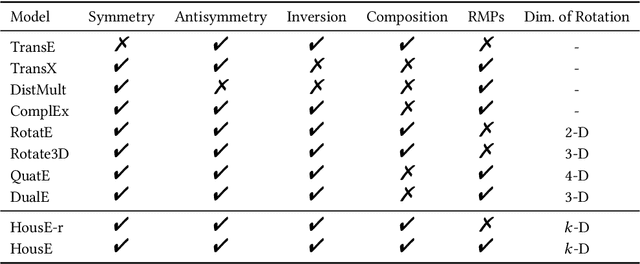
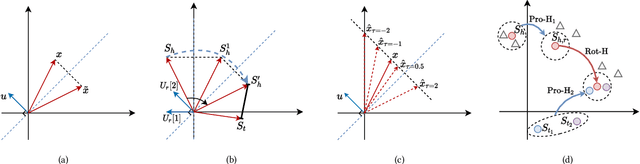
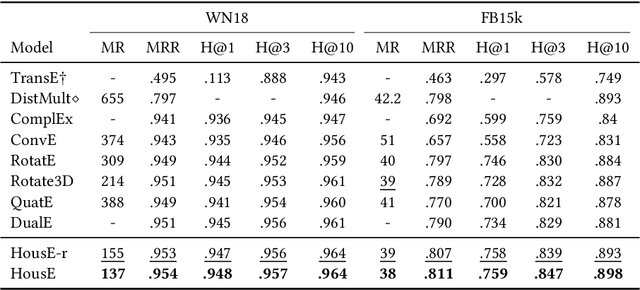
Abstract:The effectiveness of knowledge graph embedding (KGE) largely depends on the ability to model intrinsic relation patterns and mapping properties. However, existing approaches can only capture some of them with insufficient modeling capacity. In this work, we propose a more powerful KGE framework named HousE, which involves a novel parameterization based on two kinds of Householder transformations: (1) Householder rotations to achieve superior capacity of modeling relation patterns; (2) Householder projections to handle sophisticated relation mapping properties. Theoretically, HousE is capable of modeling crucial relation patterns and mapping properties simultaneously. Besides, HousE is a generalization of existing rotation-based models while extending the rotations to high-dimensional spaces. Empirically, HousE achieves new state-of-the-art performance on five benchmark datasets. Our code is available at https://github.com/anrep/HousE.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge