Daniel Rueckert
on behalf of the PINNACLE consortium
Deformable Image Registration of Dark-Field Chest Radiographs for Local Lung Signal Change Assessment
Jan 18, 2025



Abstract:Dark-field radiography of the human chest has been demonstrated to have promising potential for the analysis of the lung microstructure and the diagnosis of respiratory diseases. However, previous studies of dark-field chest radiographs evaluated the lung signal only in the inspiratory breathing state. Our work aims to add a new perspective to these previous assessments by locally comparing dark-field lung information between different respiratory states. To this end, we discuss suitable image registration methods for dark-field chest radiographs to enable consistent spatial alignment of the lung in distinct breathing states. Utilizing full inspiration and expiration scans from a clinical chronic obstructive pulmonary disease study, we assess the performance of the proposed registration framework and outline applicable evaluation approaches. Our regional characterization of lung dark-field signal changes between the breathing states provides a proof-of-principle that dynamic radiography-based lung function assessment approaches may benefit from considering registered dark-field images in addition to standard plain chest radiographs.
PISCO: Self-Supervised k-Space Regularization for Improved Neural Implicit k-Space Representations of Dynamic MRI
Jan 16, 2025



Abstract:Neural implicit k-space representations (NIK) have shown promising results for dynamic magnetic resonance imaging (MRI) at high temporal resolutions. Yet, reducing acquisition time, and thereby available training data, results in severe performance drops due to overfitting. To address this, we introduce a novel self-supervised k-space loss function $\mathcal{L}_\mathrm{PISCO}$, applicable for regularization of NIK-based reconstructions. The proposed loss function is based on the concept of parallel imaging-inspired self-consistency (PISCO), enforcing a consistent global k-space neighborhood relationship without requiring additional data. Quantitative and qualitative evaluations on static and dynamic MR reconstructions show that integrating PISCO significantly improves NIK representations. Particularly for high acceleration factors (R$\geq$54), NIK with PISCO achieves superior spatio-temporal reconstruction quality compared to state-of-the-art methods. Furthermore, an extensive analysis of the loss assumptions and stability shows PISCO's potential as versatile self-supervised k-space loss function for further applications and architectures. Code is available at: https://github.com/compai-lab/2025-pisco-spieker
TimeFlow: Longitudinal Brain Image Registration and Aging Progression Analysis
Jan 15, 2025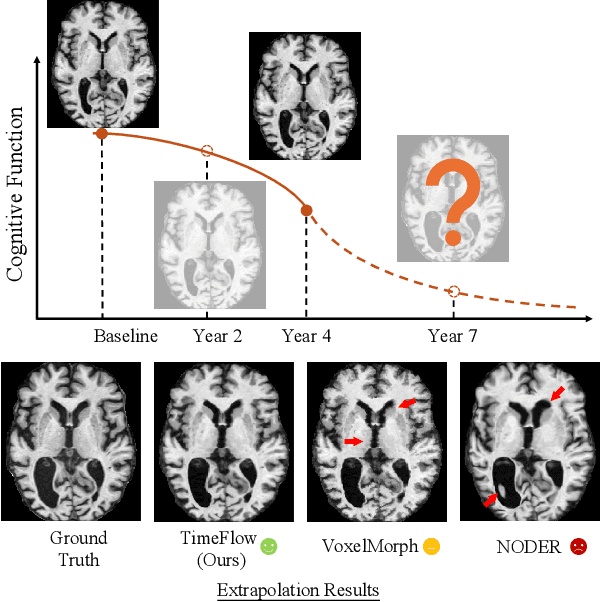


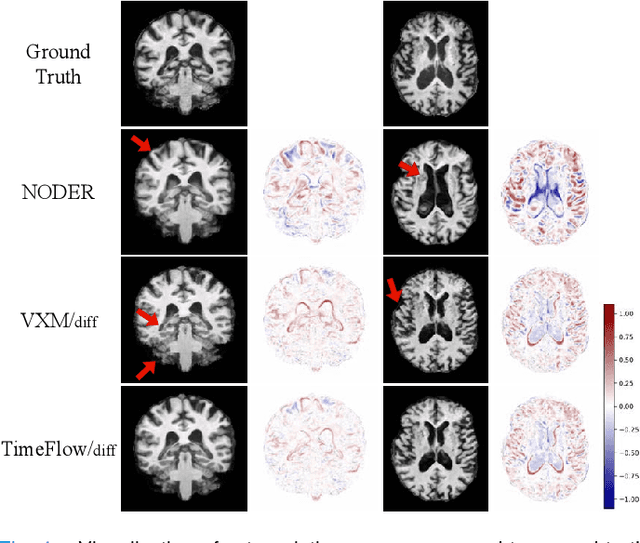
Abstract:Predicting future brain states is crucial for understanding healthy aging and neurodegenerative diseases. Longitudinal brain MRI registration, a cornerstone for such analyses, has long been limited by its inability to forecast future developments, reliance on extensive, dense longitudinal data, and the need to balance registration accuracy with temporal smoothness. In this work, we present \emph{TimeFlow}, a novel framework for longitudinal brain MRI registration that overcomes all these challenges. Leveraging a U-Net architecture with temporal conditioning inspired by diffusion models, TimeFlow enables accurate longitudinal registration and facilitates prospective analyses through future image prediction. Unlike traditional methods that depend on explicit smoothness regularizers and dense sequential data, TimeFlow achieves temporal consistency and continuity without these constraints. Experimental results highlight its superior performance in both future timepoint prediction and registration accuracy compared to state-of-the-art methods. Additionally, TimeFlow supports novel biological brain aging analyses, effectively differentiating neurodegenerative conditions from healthy aging. It eliminates the need for segmentation, thereby avoiding the challenges of non-trivial annotation and inconsistent segmentation errors. TimeFlow paves the way for accurate, data-efficient, and annotation-free prospective analyses of brain aging and chronic diseases.
Efficient Deep Learning-based Forward Solvers for Brain Tumor Growth Models
Jan 14, 2025Abstract:Glioblastoma, a highly aggressive brain tumor, poses major challenges due to its poor prognosis and high morbidity rates. Partial differential equation-based models offer promising potential to enhance therapeutic outcomes by simulating patient-specific tumor behavior for improved radiotherapy planning. However, model calibration remains a bottleneck due to the high computational demands of optimization methods like Monte Carlo sampling and evolutionary algorithms. To address this, we recently introduced an approach leveraging a neural forward solver with gradient-based optimization to significantly reduce calibration time. This approach requires a highly accurate and fully differentiable forward model. We investigate multiple architectures, including (i) an enhanced TumorSurrogate, (ii) a modified nnU-Net, and (iii) a 3D Vision Transformer (ViT). The optimized TumorSurrogate achieved the best overall results, excelling in both tumor outline matching and voxel-level prediction of tumor cell concentration. It halved the MSE relative to the baseline model and achieved the highest Dice score across all tumor cell concentration thresholds. Our study demonstrates significant enhancement in forward solver performance and outlines important future research directions.
Self-Supervised Radiograph Anatomical Region Classification -- How Clean Is Your Real-World Data?
Dec 20, 2024Abstract:Modern deep learning-based clinical imaging workflows rely on accurate labels of the examined anatomical region. Knowing the anatomical region is required to select applicable downstream models and to effectively generate cohorts of high quality data for future medical and machine learning research efforts. However, this information may not be available in externally sourced data or generally contain data entry errors. To address this problem, we show the effectiveness of self-supervised methods such as SimCLR and BYOL as well as supervised contrastive deep learning methods in assigning one of 14 anatomical region classes in our in-house dataset of 48,434 skeletal radiographs. We achieve a strong linear evaluation accuracy of 96.6% with a single model and 97.7% using an ensemble approach. Furthermore, only a few labeled instances (1% of the training set) suffice to achieve an accuracy of 92.2%, enabling usage in low-label and thus low-resource scenarios. Our model can be used to correct data entry mistakes: a follow-up analysis of the test set errors of our best-performing single model by an expert radiologist identified 35% incorrect labels and 11% out-of-domain images. When accounted for, the radiograph anatomical region labelling performance increased -- without and with an ensemble, respectively -- to a theoretical accuracy of 98.0% and 98.8%.
From Model Based to Learned Regularization in Medical Image Registration: A Comprehensive Review
Dec 20, 2024Abstract:Image registration is fundamental in medical imaging applications, such as disease progression analysis or radiation therapy planning. The primary objective of image registration is to precisely capture the deformation between two or more images, typically achieved by minimizing an optimization problem. Due to its inherent ill-posedness, regularization is a key component in driving the solution toward anatomically meaningful deformations. A wide range of regularization methods has been proposed for both conventional and deep learning-based registration. However, the appropriate application of regularization techniques often depends on the specific registration problem, and no one-fits-all method exists. Despite its importance, regularization is often overlooked or addressed with default approaches, assuming existing methods are sufficient. A comprehensive and structured review remains missing. This review addresses this gap by introducing a novel taxonomy that systematically categorizes the diverse range of proposed regularization methods. It highlights the emerging field of learned regularization, which leverages data-driven techniques to automatically derive deformation properties from the data. Moreover, this review examines the transfer of regularization methods from conventional to learning-based registration, identifies open challenges, and outlines future research directions. By emphasizing the critical role of regularization in image registration, we hope to inspire the research community to reconsider regularization strategies in modern registration algorithms and to explore this rapidly evolving field further.
Pitfalls of topology-aware image segmentation
Dec 19, 2024



Abstract:Topological correctness, i.e., the preservation of structural integrity and specific characteristics of shape, is a fundamental requirement for medical imaging tasks, such as neuron or vessel segmentation. Despite the recent surge in topology-aware methods addressing this challenge, their real-world applicability is hindered by flawed benchmarking practices. In this paper, we identify critical pitfalls in model evaluation that include inadequate connectivity choices, overlooked topological artifacts in ground truth annotations, and inappropriate use of evaluation metrics. Through detailed empirical analysis, we uncover these issues' profound impact on the evaluation and ranking of segmentation methods. Drawing from our findings, we propose a set of actionable recommendations to establish fair and robust evaluation standards for topology-aware medical image segmentation methods.
DCRA-Net: Attention-Enabled Reconstruction Model for Dynamic Fetal Cardiac MRI
Dec 19, 2024



Abstract:Dynamic fetal heart magnetic resonance imaging (MRI) presents unique challenges due to the fast heart rate of the fetus compared to adult subjects and uncontrolled fetal motion. This requires high temporal and spatial resolutions over a large field of view, in order to encompass surrounding maternal anatomy. In this work, we introduce Dynamic Cardiac Reconstruction Attention Network (DCRA-Net) - a novel deep learning model that employs attention mechanisms in spatial and temporal domains and temporal frequency representation of data to reconstruct the dynamics of the fetal heart from highly accelerated free-running (non-gated) MRI acquisitions. DCRA-Net was trained on retrospectively undersampled complex-valued cardiac MRIs from 42 fetal subjects and separately from 153 adult subjects, and evaluated on data from 14 fetal and 39 adult subjects respectively. Its performance was compared to L+S and k-GIN methods in both fetal and adult cases for an undersampling factor of 8x. The proposed network performed better than the comparators for both fetal and adult data, for both regular lattice and centrally weighted random undersampling. Aliased signals due to the undersampling were comprehensively resolved, and both the spatial details of the heart and its temporal dynamics were recovered with high fidelity. The highest performance was achieved when using lattice undersampling, data consistency and temporal frequency representation, yielding PSNR of 38 for fetal and 35 for adult cases. Our method is publicly available at https://github.com/denproc/DCRA-Net.
Image registration is a geometric deep learning task
Dec 17, 2024



Abstract:Data-driven deformable image registration methods predominantly rely on operations that process grid-like inputs. However, applying deformable transformations to an image results in a warped space that deviates from a rigid grid structure. Consequently, data-driven approaches with sequential deformations have to apply grid resampling operations between each deformation step. While artifacts caused by resampling are negligible in high-resolution images, the resampling of sparse, high-dimensional feature grids introduces errors that affect the deformation modeling process. Taking inspiration from Lagrangian reference frames of deformation fields, our work introduces a novel paradigm for data-driven deformable image registration that utilizes geometric deep-learning principles to model deformations without grid requirements. Specifically, we model image features as a set of nodes that freely move in Euclidean space, update their coordinates under graph operations, and dynamically readjust their local neighborhoods. We employ this formulation to construct a multi-resolution deformable registration model, where deformation layers iteratively refine the overall transformation at each resolution without intermediate resampling operations on the feature grids. We investigate our method's ability to fully deformably capture large deformations across a number of medical imaging registration tasks. In particular, we apply our approach (GeoReg) to the registration of inter-subject brain MR images and inhale-exhale lung CT images, showing on par performance with the current state-of-the-art methods. We believe our contribution open up avenues of research to reduce the black-box nature of current learned registration paradigms by explicitly modeling the transformation within the architecture.
Subspace Implicit Neural Representations for Real-Time Cardiac Cine MR Imaging
Dec 17, 2024



Abstract:Conventional cardiac cine MRI methods rely on retrospective gating, which limits temporal resolution and the ability to capture continuous cardiac dynamics, particularly in patients with arrhythmias and beat-to-beat variations. To address these challenges, we propose a reconstruction framework based on subspace implicit neural representations for real-time cardiac cine MRI of continuously sampled radial data. This approach employs two multilayer perceptrons to learn spatial and temporal subspace bases, leveraging the low-rank properties of cardiac cine MRI. Initialized with low-resolution reconstructions, the networks are fine-tuned using spoke-specific loss functions to recover spatial details and temporal fidelity. Our method directly utilizes the continuously sampled radial k-space spokes during training, thereby eliminating the need for binning and non-uniform FFT. This approach achieves superior spatial and temporal image quality compared to conventional binned methods at the acceleration rate of 10 and 20, demonstrating potential for high-resolution imaging of dynamic cardiac events and enhancing diagnostic capability.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge