Daniel Rueckert
on behalf of the PINNACLE consortium
Towards Cardiac MRI Foundation Models: Comprehensive Visual-Tabular Representations for Whole-Heart Assessment and Beyond
Apr 18, 2025Abstract:Cardiac magnetic resonance imaging is the gold standard for non-invasive cardiac assessment, offering rich spatio-temporal views of the cardiac anatomy and physiology. Patient-level health factors, such as demographics, metabolic, and lifestyle, are known to substantially influence cardiovascular health and disease risk, yet remain uncaptured by CMR alone. To holistically understand cardiac health and to enable the best possible interpretation of an individual's disease risk, CMR and patient-level factors must be jointly exploited within an integrated framework. Recent multi-modal approaches have begun to bridge this gap, yet they often rely on limited spatio-temporal data and focus on isolated clinical tasks, thereby hindering the development of a comprehensive representation for cardiac health evaluation. To overcome these limitations, we introduce ViTa, a step toward foundation models that delivers a comprehensive representation of the heart and a precise interpretation of individual disease risk. Leveraging data from 42,000 UK Biobank participants, ViTa integrates 3D+T cine stacks from short-axis and long-axis views, enabling a complete capture of the cardiac cycle. These imaging data are then fused with detailed tabular patient-level factors, enabling context-aware insights. This multi-modal paradigm supports a wide spectrum of downstream tasks, including cardiac phenotype and physiological feature prediction, segmentation, and classification of cardiac and metabolic diseases within a single unified framework. By learning a shared latent representation that bridges rich imaging features and patient context, ViTa moves beyond traditional, task-specific models toward a universal, patient-specific understanding of cardiac health, highlighting its potential to advance clinical utility and scalability in cardiac analysis.
From Pixels to Histopathology: A Graph-Based Framework for Interpretable Whole Slide Image Analysis
Mar 14, 2025Abstract:The histopathological classification of whole-slide images (WSIs) is a fundamental task in digital pathology; yet it requires extensive time and expertise from specialists. While deep learning methods show promising results, they typically process WSIs by dividing them into artificial patches, which inherently prevents a network from learning from the entire image context, disregards natural tissue structures and compromises interpretability. Our method overcomes this limitation through a novel graph-based framework that constructs WSI graph representations. The WSI-graph efficiently captures essential histopathological information in a compact form. We build tissue representations (nodes) that follow biological boundaries rather than arbitrary patches all while providing interpretable features for explainability. Through adaptive graph coarsening guided by learned embeddings, we progressively merge regions while maintaining discriminative local features and enabling efficient global information exchange. In our method's final step, we solve the diagnostic task through a graph attention network. We empirically demonstrate strong performance on multiple challenging tasks such as cancer stage classification and survival prediction, while also identifying predictive factors using Integrated Gradients. Our implementation is publicly available at https://github.com/HistoGraph31/pix2pathology
Evaluation of Alignment-Regularity Characteristics in Deformable Image Registration
Mar 10, 2025



Abstract:Evaluating deformable image registration (DIR) is challenging due to the inherent trade-off between achieving high alignment accuracy and maintaining deformation regularity. In this work, we introduce a novel evaluation scheme based on the alignment-regularity characteristic (ARC) to systematically capture and analyze this trade-off. We first introduce the ARC curves, which describe the performance of a given registration algorithm as a spectrum measured by alignment and regularity metrics. We further adopt a HyperNetwork-based approach that learns to continuously interpolate across the full regularization range, accelerating the construction and improving the sample density of ARC curves. We empirically demonstrate our evaluation scheme using representative learning-based deformable image registration methods with various network architectures and transformation models on two public datasets. We present a range of findings not evident from existing evaluation practices and provide general recommendations for model evaluation and selection using our evaluation scheme. All code relevant is made publicly available.
Are foundation models useful feature extractors for electroencephalography analysis?
Feb 28, 2025Abstract:The success of foundation models in natural language processing and computer vision has motivated similar approaches for general time series analysis. While these models are effective for a variety of tasks, their applicability in medical domains with limited data remains largely unexplored. To address this, we investigate the effectiveness of foundation models in medical time series analysis involving electroencephalography (EEG). Through extensive experiments on tasks such as age prediction, seizure detection, and the classification of clinically relevant EEG events, we compare their diagnostic accuracy with that of specialised EEG models. Our analysis shows that foundation models extract meaningful EEG features, outperform specialised models even without domain adaptation, and localise task-specific biomarkers. Moreover, we demonstrate that diagnostic accuracy is substantially influenced by architectural choices such as context length. Overall, our study reveals that foundation models with general time series understanding eliminate the dependency on large domain-specific datasets, making them valuable tools for clinical practice.
MedVLM-R1: Incentivizing Medical Reasoning Capability of Vision-Language Models (VLMs) via Reinforcement Learning
Feb 26, 2025


Abstract:Reasoning is a critical frontier for advancing medical image analysis, where transparency and trustworthiness play a central role in both clinician trust and regulatory approval. Although Medical Visual Language Models (VLMs) show promise for radiological tasks, most existing VLMs merely produce final answers without revealing the underlying reasoning. To address this gap, we introduce MedVLM-R1, a medical VLM that explicitly generates natural language reasoning to enhance transparency and trustworthiness. Instead of relying on supervised fine-tuning (SFT), which often suffers from overfitting to training distributions and fails to foster genuine reasoning, MedVLM-R1 employs a reinforcement learning framework that incentivizes the model to discover human-interpretable reasoning paths without using any reasoning references. Despite limited training data (600 visual question answering samples) and model parameters (2B), MedVLM-R1 boosts accuracy from 55.11% to 78.22% across MRI, CT, and X-ray benchmarks, outperforming larger models trained on over a million samples. It also demonstrates robust domain generalization under out-of-distribution tasks. By unifying medical image analysis with explicit reasoning, MedVLM-R1 marks a pivotal step toward trustworthy and interpretable AI in clinical practice.
Interpretable Retinal Disease Prediction Using Biology-Informed Heterogeneous Graph Representations
Feb 23, 2025



Abstract:Interpretability is crucial to enhance trust in machine learning models for medical diagnostics. However, most state-of-the-art image classifiers based on neural networks are not interpretable. As a result, clinicians often resort to known biomarkers for diagnosis, although biomarker-based classification typically performs worse than large neural networks. This work proposes a method that surpasses the performance of established machine learning models while simultaneously improving prediction interpretability for diabetic retinopathy staging from optical coherence tomography angiography (OCTA) images. Our method is based on a novel biology-informed heterogeneous graph representation that models retinal vessel segments, intercapillary areas, and the foveal avascular zone (FAZ) in a human-interpretable way. This graph representation allows us to frame diabetic retinopathy staging as a graph-level classification task, which we solve using an efficient graph neural network. We benchmark our method against well-established baselines, including classical biomarker-based classifiers, convolutional neural networks (CNNs), and vision transformers. Our model outperforms all baselines on two datasets. Crucially, we use our biology-informed graph to provide explanations of unprecedented detail. Our approach surpasses existing methods in precisely localizing and identifying critical vessels or intercapillary areas. In addition, we give informative and human-interpretable attributions to critical characteristics. Our work contributes to the development of clinical decision-support tools in ophthalmology.
MAGO-SP: Detection and Correction of Water-Fat Swaps in Magnitude-Only VIBE MRI
Feb 20, 2025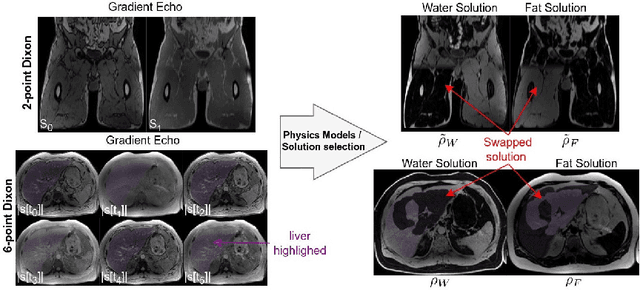
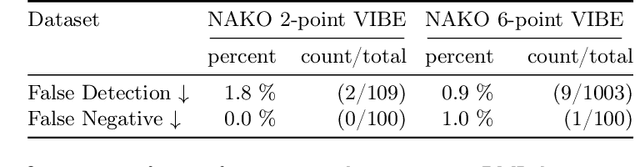
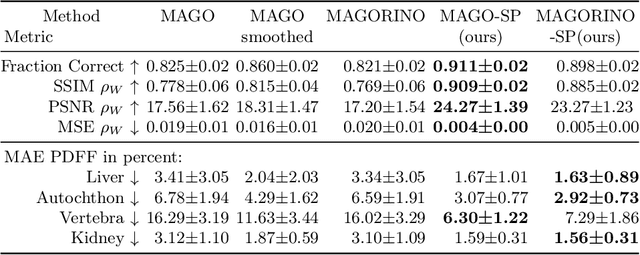
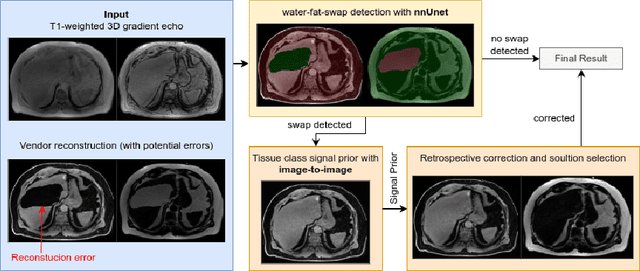

Abstract:Volume Interpolated Breath-Hold Examination (VIBE) MRI generates images suitable for water and fat signal composition estimation. While the two-point VIBE provides water-fat-separated images, the six-point VIBE allows estimation of the effective transversal relaxation rate R2* and the proton density fat fraction (PDFF), which are imaging markers for health and disease. Ambiguity during signal reconstruction can lead to water-fat swaps. This shortcoming challenges the application of VIBE-MRI for automated PDFF analyses of large-scale clinical data and of population studies. This study develops an automated pipeline to detect and correct water-fat swaps in non-contrast-enhanced VIBE images. Our three-step pipeline begins with training a segmentation network to classify volumes as "fat-like" or "water-like," using synthetic water-fat swaps generated by merging fat and water volumes with Perlin noise. Next, a denoising diffusion image-to-image network predicts water volumes as signal priors for correction. Finally, we integrate this prior into a physics-constrained model to recover accurate water and fat signals. Our approach achieves a < 1% error rate in water-fat swap detection for a 6-point VIBE. Notably, swaps disproportionately affect individuals in the Underweight and Class 3 Obesity BMI categories. Our correction algorithm ensures accurate solution selection in chemical phase MRIs, enabling reliable PDFF estimation. This forms a solid technical foundation for automated large-scale population imaging analysis.
Fake It Till You Make It: Using Synthetic Data and Domain Knowledge for Improved Text-Based Learning for LGE Detection
Feb 18, 2025



Abstract:Detection of hyperenhancement from cardiac LGE MRI images is a complex task requiring significant clinical expertise. Although deep learning-based models have shown promising results for the task, they require large amounts of data with fine-grained annotations. Clinical reports generated for cardiac MR studies contain rich, clinically relevant information, including the location, extent and etiology of any scars present. Although recently developed CLIP-based training enables pretraining models with image-text pairs, it requires large amounts of data and further finetuning strategies on downstream tasks. In this study, we use various strategies rooted in domain knowledge to train a model for LGE detection solely using text from clinical reports, on a relatively small clinical cohort of 965 patients. We improve performance through the use of synthetic data augmentation, by systematically creating scar images and associated text. In addition, we standardize the orientation of the images in an anatomy-informed way to enable better alignment of spatial and text features. We also use a captioning loss to enable fine-grained supervision and explore the effect of pretraining of the vision encoder on performance. Finally, ablation studies are carried out to elucidate the contributions of each design component to the overall performance of the model.
SIM: Surface-based fMRI Analysis for Inter-Subject Multimodal Decoding from Movie-Watching Experiments
Jan 27, 2025Abstract:Current AI frameworks for brain decoding and encoding, typically train and test models within the same datasets. This limits their utility for brain computer interfaces (BCI) or neurofeedback, for which it would be useful to pool experiences across individuals to better simulate stimuli not sampled during training. A key obstacle to model generalisation is the degree of variability of inter-subject cortical organisation, which makes it difficult to align or compare cortical signals across participants. In this paper we address this through the use of surface vision transformers, which build a generalisable model of cortical functional dynamics, through encoding the topography of cortical networks and their interactions as a moving image across a surface. This is then combined with tri-modal self-supervised contrastive (CLIP) alignment of audio, video, and fMRI modalities to enable the retrieval of visual and auditory stimuli from patterns of cortical activity (and vice-versa). We validate our approach on 7T task-fMRI data from 174 healthy participants engaged in the movie-watching experiment from the Human Connectome Project (HCP). Results show that it is possible to detect which movie clips an individual is watching purely from their brain activity, even for individuals and movies not seen during training. Further analysis of attention maps reveals that our model captures individual patterns of brain activity that reflect semantic and visual systems. This opens the door to future personalised simulations of brain function. Code & pre-trained models will be made available at https://github.com/metrics-lab/sim, processed data for training will be available upon request at https://gin.g-node.org/Sdahan30/sim.
PARASIDE: An Automatic Paranasal Sinus Segmentation and Structure Analysis Tool for MRI
Jan 24, 2025



Abstract:Chronic rhinosinusitis (CRS) is a common and persistent sinus imflammation that affects 5 - 12\% of the general population. It significantly impacts quality of life and is often difficult to assess due to its subjective nature in clinical evaluation. We introduce PARASIDE, an automatic tool for segmenting air and soft tissue volumes of the structures of the sinus maxillaris, frontalis, sphenodalis and ethmoidalis in T1 MRI. By utilizing that segmentation, we can quantify feature relations that have been observed only manually and subjectively before. We performed an exemplary study and showed both volume and intensity relations between structures and radiology reports. While the soft tissue segmentation is good, the automated annotations of the air volumes are excellent. The average intensity over air structures are consistently below those of the soft tissues, close to perfect separability. Healthy subjects exhibit lower soft tissue volumes and lower intensities. Our developed system is the first automated whole nasal segmentation of 16 structures, and capable of calculating medical relevant features such as the Lund-Mackay score.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge