Daguang Xu
Swin UNETR: Swin Transformers for Semantic Segmentation of Brain Tumors in MRI Images
Jan 04, 2022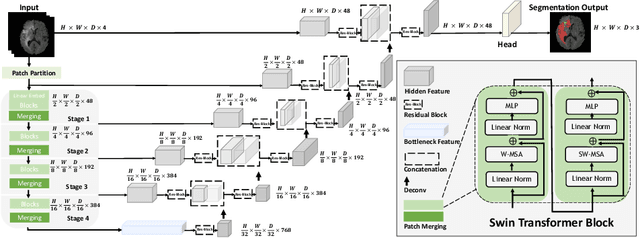

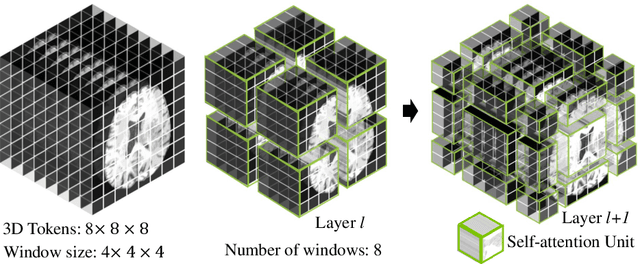
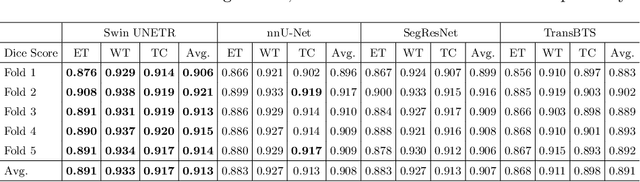
Abstract:Semantic segmentation of brain tumors is a fundamental medical image analysis task involving multiple MRI imaging modalities that can assist clinicians in diagnosing the patient and successively studying the progression of the malignant entity. In recent years, Fully Convolutional Neural Networks (FCNNs) approaches have become the de facto standard for 3D medical image segmentation. The popular "U-shaped" network architecture has achieved state-of-the-art performance benchmarks on different 2D and 3D semantic segmentation tasks and across various imaging modalities. However, due to the limited kernel size of convolution layers in FCNNs, their performance of modeling long-range information is sub-optimal, and this can lead to deficiencies in the segmentation of tumors with variable sizes. On the other hand, transformer models have demonstrated excellent capabilities in capturing such long-range information in multiple domains, including natural language processing and computer vision. Inspired by the success of vision transformers and their variants, we propose a novel segmentation model termed Swin UNEt TRansformers (Swin UNETR). Specifically, the task of 3D brain tumor semantic segmentation is reformulated as a sequence to sequence prediction problem wherein multi-modal input data is projected into a 1D sequence of embedding and used as an input to a hierarchical Swin transformer as the encoder. The swin transformer encoder extracts features at five different resolutions by utilizing shifted windows for computing self-attention and is connected to an FCNN-based decoder at each resolution via skip connections. We have participated in BraTS 2021 segmentation challenge, and our proposed model ranks among the top-performing approaches in the validation phase. Code: https://monai.io/research/swin-unetr
HyperSegNAS: Bridging One-Shot Neural Architecture Search with 3D Medical Image Segmentation using HyperNet
Dec 20, 2021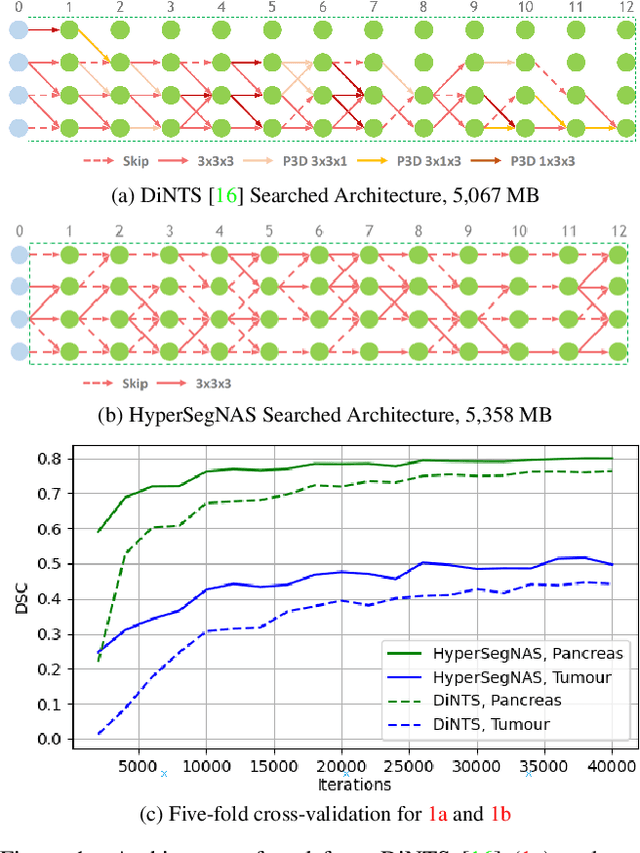
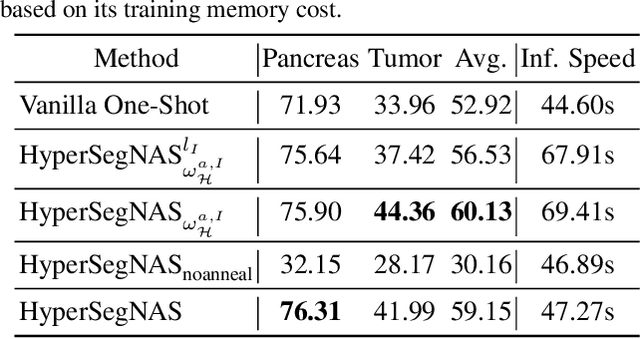
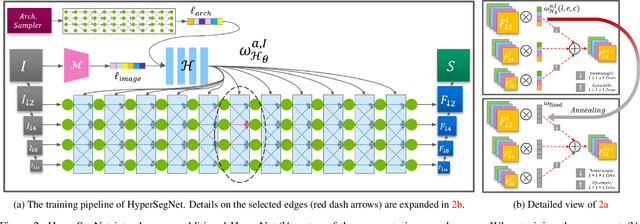
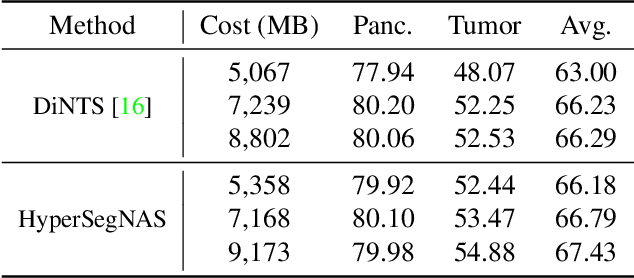
Abstract:Semantic segmentation of 3D medical images is a challenging task due to the high variability of the shape and pattern of objects (such as organs or tumors). Given the recent success of deep learning in medical image segmentation, Neural Architecture Search (NAS) has been introduced to find high-performance 3D segmentation network architectures. However, because of the massive computational requirements of 3D data and the discrete optimization nature of architecture search, previous NAS methods require a long search time or necessary continuous relaxation, and commonly lead to sub-optimal network architectures. While one-shot NAS can potentially address these disadvantages, its application in the segmentation domain has not been well studied in the expansive multi-scale multi-path search space. To enable one-shot NAS for medical image segmentation, our method, named HyperSegNAS, introduces a HyperNet to assist super-net training by incorporating architecture topology information. Such a HyperNet can be removed once the super-net is trained and introduces no overhead during architecture search. We show that HyperSegNAS yields better performing and more intuitive architectures compared to the previous state-of-the-art (SOTA) segmentation networks; furthermore, it can quickly and accurately find good architecture candidates under different computing constraints. Our method is evaluated on public datasets from the Medical Segmentation Decathlon (MSD) challenge, and achieves SOTA performances.
Self-Supervised Pre-Training of Swin Transformers for 3D Medical Image Analysis
Nov 29, 2021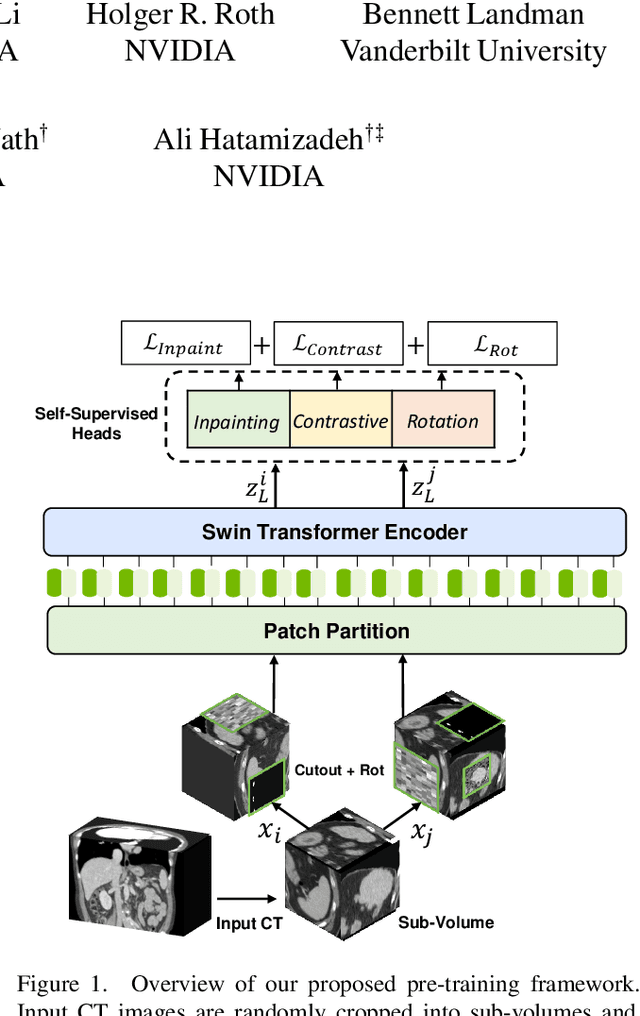
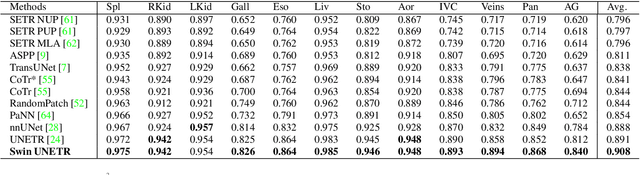
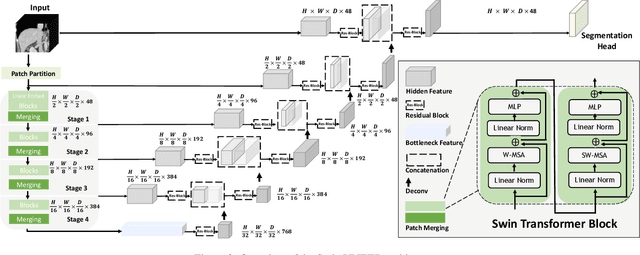
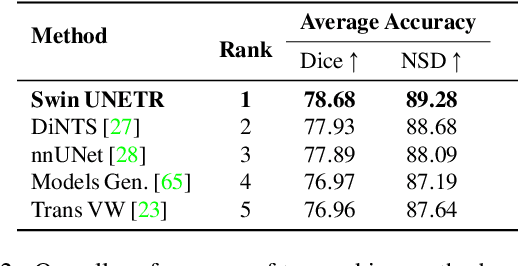
Abstract:Vision Transformers (ViT)s have shown great performance in self-supervised learning of global and local representations that can be transferred to downstream applications. Inspired by these results, we introduce a novel self-supervised learning framework with tailored proxy tasks for medical image analysis. Specifically, we propose: (i) a new 3D transformer-based model, dubbed Swin UNEt TRansformers (Swin UNETR), with a hierarchical encoder for self-supervised pre-training; (ii) tailored proxy tasks for learning the underlying pattern of human anatomy. We demonstrate successful pre-training of the proposed model on 5,050 publicly available computed tomography (CT) images from various body organs. The effectiveness of our approach is validated by fine-tuning the pre-trained models on the Beyond the Cranial Vault (BTCV) Segmentation Challenge with 13 abdominal organs and segmentation tasks from the Medical Segmentation Decathlon (MSD) dataset. Our model is currently the state-of-the-art (i.e. ranked 1st) on the public test leaderboards of both MSD and BTCV datasets. Code: https://monai.io/research/swin-unetr
T-AutoML: Automated Machine Learning for Lesion Segmentation using Transformers in 3D Medical Imaging
Nov 15, 2021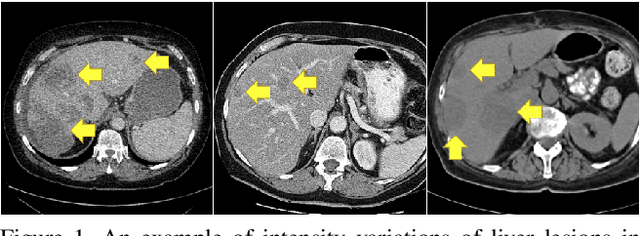
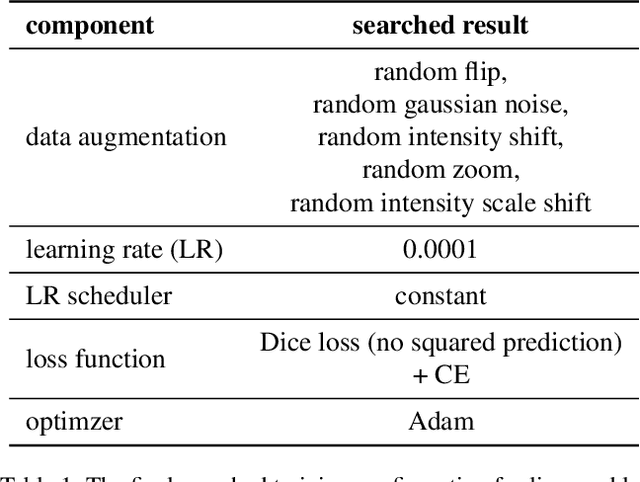
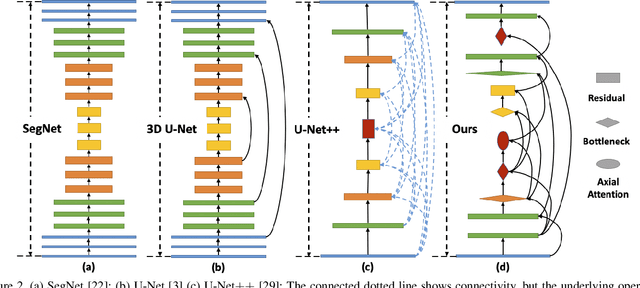
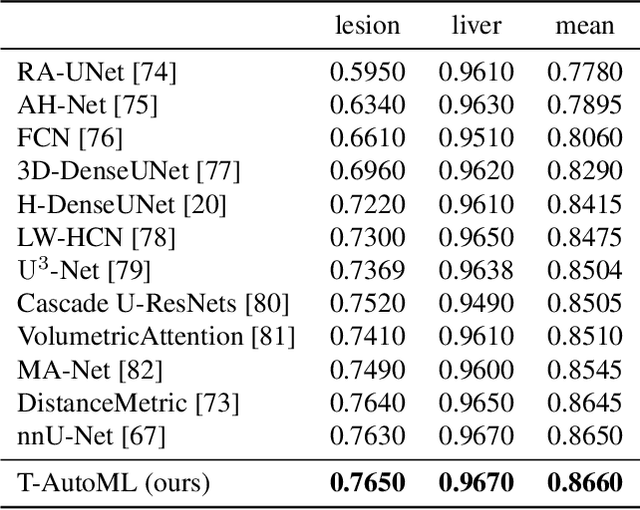
Abstract:Lesion segmentation in medical imaging has been an important topic in clinical research. Researchers have proposed various detection and segmentation algorithms to address this task. Recently, deep learning-based approaches have significantly improved the performance over conventional methods. However, most state-of-the-art deep learning methods require the manual design of multiple network components and training strategies. In this paper, we propose a new automated machine learning algorithm, T-AutoML, which not only searches for the best neural architecture, but also finds the best combination of hyper-parameters and data augmentation strategies simultaneously. The proposed method utilizes the modern transformer model, which is introduced to adapt to the dynamic length of the search space embedding and can significantly improve the ability of the search. We validate T-AutoML on several large-scale public lesion segmentation data-sets and achieve state-of-the-art performance.
Accounting for Dependencies in Deep Learning Based Multiple Instance Learning for Whole Slide Imaging
Nov 01, 2021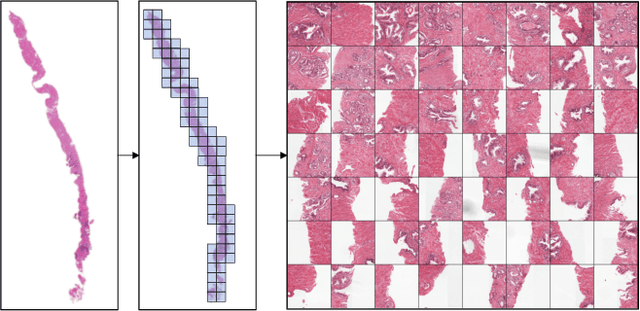
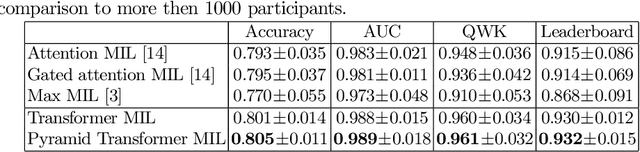
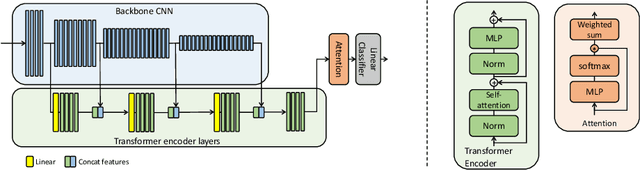
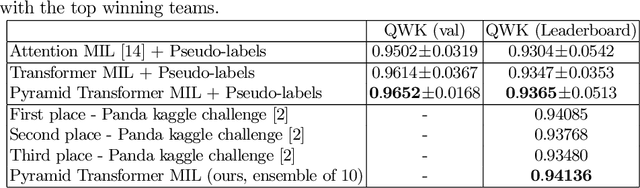
Abstract:Multiple instance learning (MIL) is a key algorithm for classification of whole slide images (WSI). Histology WSIs can have billions of pixels, which create enormous computational and annotation challenges. Typically, such images are divided into a set of patches (a bag of instances), where only bag-level class labels are provided. Deep learning based MIL methods calculate instance features using convolutional neural network (CNN). Our proposed approach is also deep learning based, with the following two contributions: Firstly, we propose to explicitly account for dependencies between instances during training by embedding self-attention Transformer blocks to capture dependencies between instances. For example, a tumor grade may depend on the presence of several particular patterns at different locations in WSI, which requires to account for dependencies between patches. Secondly, we propose an instance-wise loss function based on instance pseudo-labels. We compare the proposed algorithm to multiple baseline methods, evaluate it on the PANDA challenge dataset, the largest publicly available WSI dataset with over 11K images, and demonstrate state-of-the-art results.
MedPerf: Open Benchmarking Platform for Medical Artificial Intelligence using Federated Evaluation
Oct 08, 2021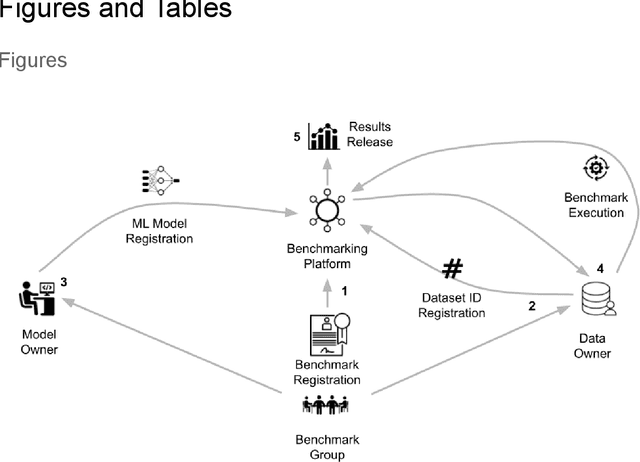
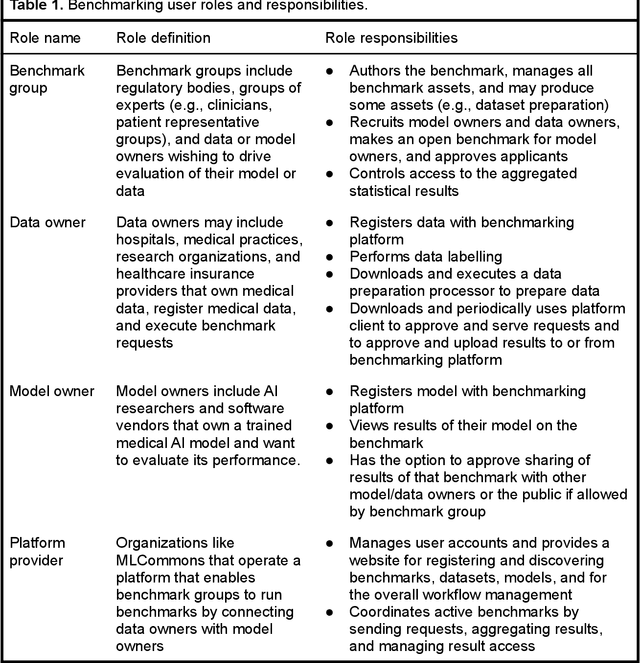
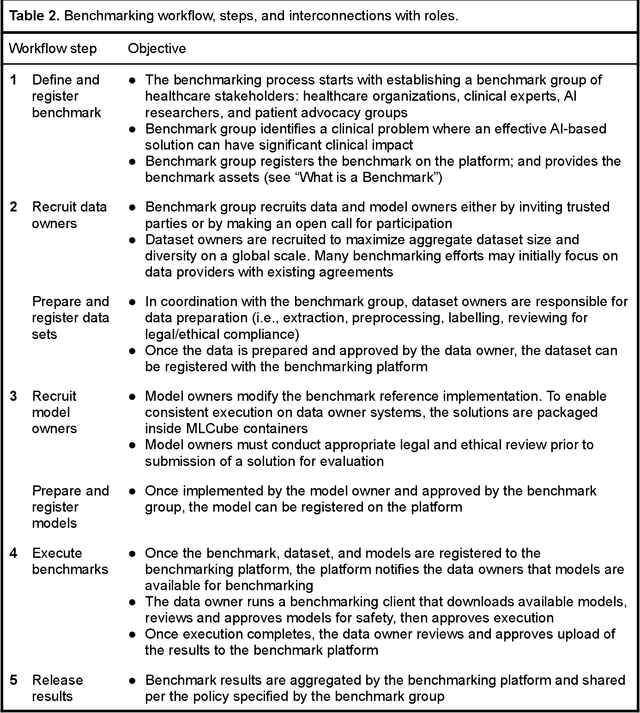
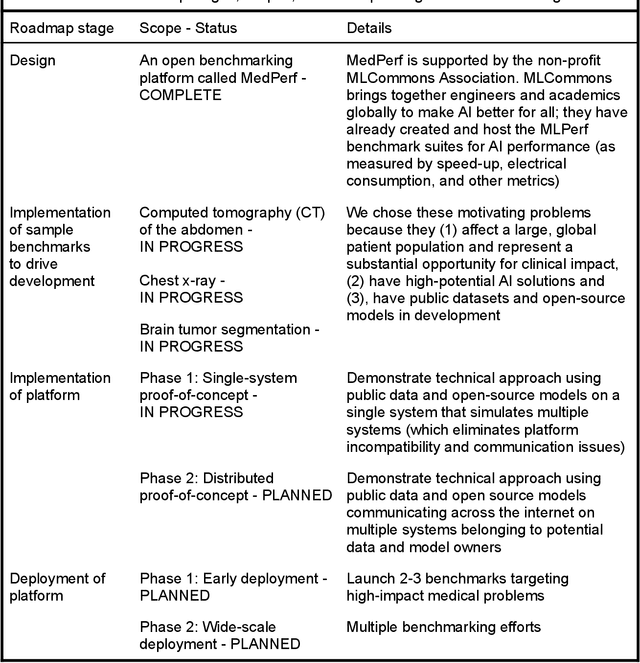
Abstract:Medical AI has tremendous potential to advance healthcare by supporting the evidence-based practice of medicine, personalizing patient treatment, reducing costs, and improving provider and patient experience. We argue that unlocking this potential requires a systematic way to measure the performance of medical AI models on large-scale heterogeneous data. To meet this need, we are building MedPerf, an open framework for benchmarking machine learning in the medical domain. MedPerf will enable federated evaluation in which models are securely distributed to different facilities for evaluation, thereby empowering healthcare organizations to assess and verify the performance of AI models in an efficient and human-supervised process, while prioritizing privacy. We describe the current challenges healthcare and AI communities face, the need for an open platform, the design philosophy of MedPerf, its current implementation status, and our roadmap. We call for researchers and organizations to join us in creating the MedPerf open benchmarking platform.
Improving Pneumonia Localization via Cross-Attention on Medical Images and Reports
Oct 06, 2021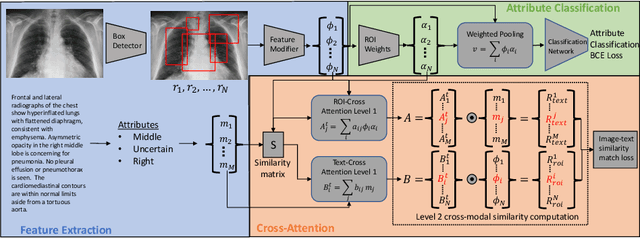
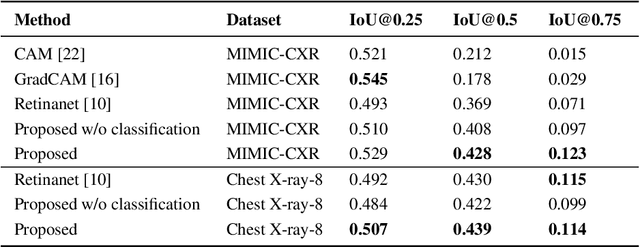
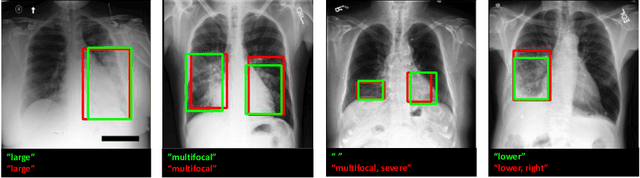
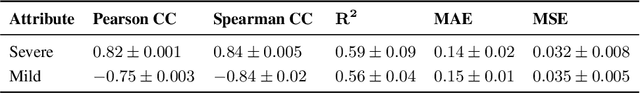
Abstract:Localization and characterization of diseases like pneumonia are primary steps in a clinical pipeline, facilitating detailed clinical diagnosis and subsequent treatment planning. Additionally, such location annotated datasets can provide a pathway for deep learning models to be used for downstream tasks. However, acquiring quality annotations is expensive on human resources and usually requires domain expertise. On the other hand, medical reports contain a plethora of information both about pneumonia characteristics and its location. In this paper, we propose a novel weakly-supervised attention-driven deep learning model that leverages encoded information in medical reports during training to facilitate better localization. Our model also performs classification of attributes that are associated to pneumonia and extracted from medical reports for supervision. Both the classification and localization are trained in conjunction and once trained, the model can be utilized for both the localization and characterization of pneumonia using only the input image. In this paper, we explore and analyze the model using chest X-ray datasets and demonstrate qualitatively and quantitatively that the introduction of textual information improves pneumonia localization. We showcase quantitative results on two datasets, MIMIC-CXR and Chest X-ray-8, and we also showcase severity characterization on the COVID-19 dataset.
Multi-task Federated Learning for Heterogeneous Pancreas Segmentation
Aug 19, 2021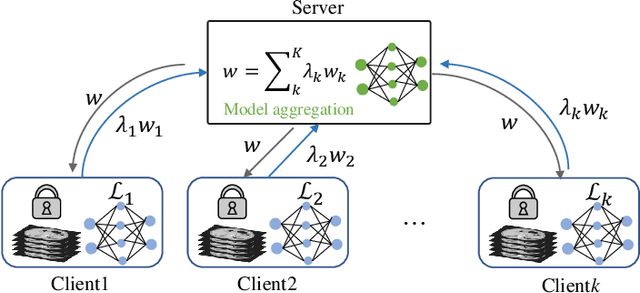
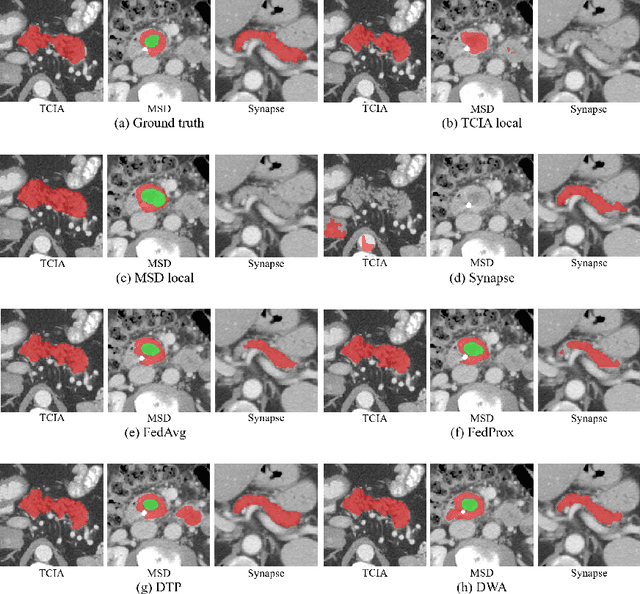
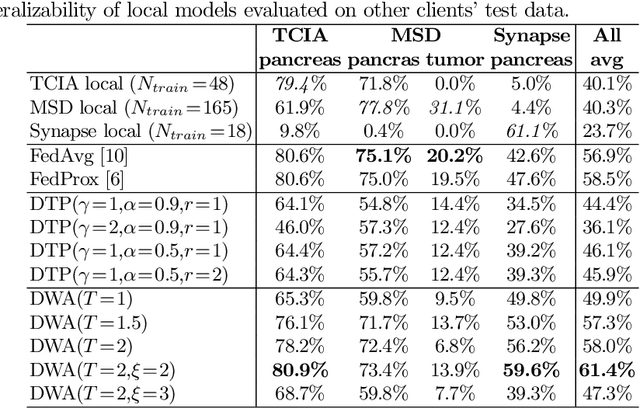
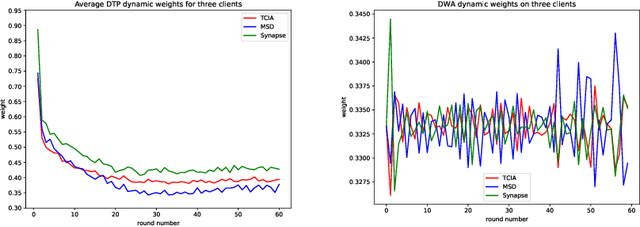

Abstract:Federated learning (FL) for medical image segmentation becomes more challenging in multi-task settings where clients might have different categories of labels represented in their data. For example, one client might have patient data with "healthy'' pancreases only while datasets from other clients may contain cases with pancreatic tumors. The vanilla federated averaging algorithm makes it possible to obtain more generalizable deep learning-based segmentation models representing the training data from multiple institutions without centralizing datasets. However, it might be sub-optimal for the aforementioned multi-task scenarios. In this paper, we investigate heterogeneous optimization methods that show improvements for the automated segmentation of pancreas and pancreatic tumors in abdominal CT images with FL settings.
Federated Whole Prostate Segmentation in MRI with Personalized Neural Architectures
Jul 16, 2021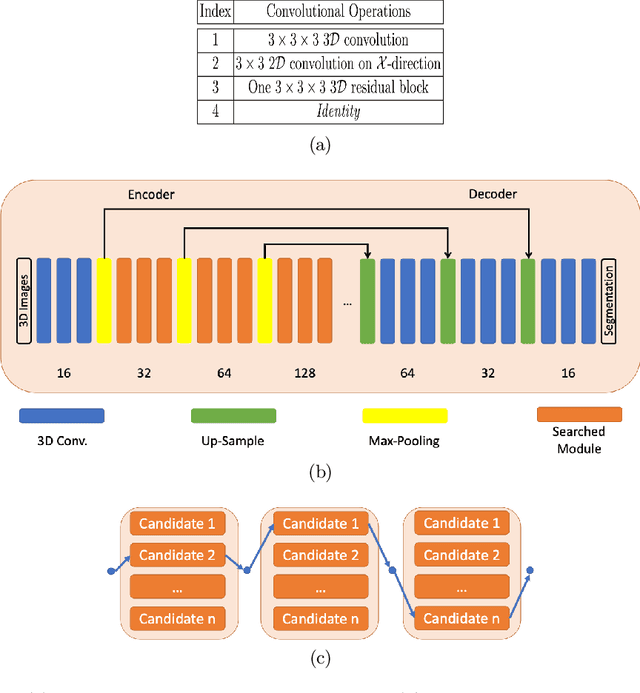
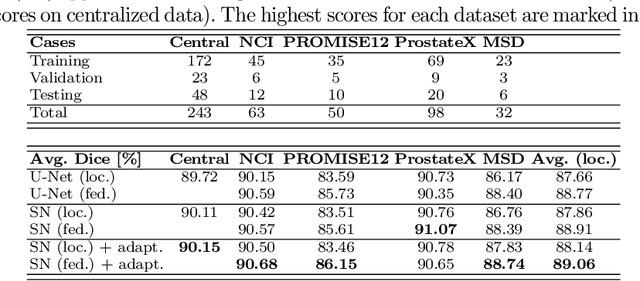
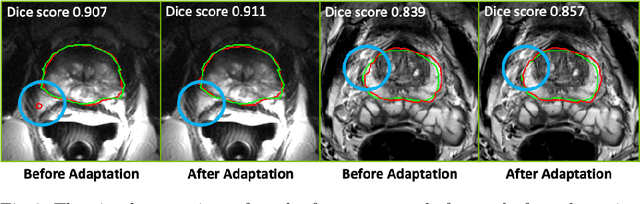
Abstract:Building robust deep learning-based models requires diverse training data, ideally from several sources. However, these datasets cannot be combined easily because of patient privacy concerns or regulatory hurdles, especially if medical data is involved. Federated learning (FL) is a way to train machine learning models without the need for centralized datasets. Each FL client trains on their local data while only sharing model parameters with a global server that aggregates the parameters from all clients. At the same time, each client's data can exhibit differences and inconsistencies due to the local variation in the patient population, imaging equipment, and acquisition protocols. Hence, the federated learned models should be able to adapt to the local particularities of a client's data. In this work, we combine FL with an AutoML technique based on local neural architecture search by training a "supernet". Furthermore, we propose an adaptation scheme to allow for personalized model architectures at each FL client's site. The proposed method is evaluated on four different datasets from 3D prostate MRI and shown to improve the local models' performance after adaptation through selecting an optimal path through the AutoML supernet.
The Power of Proxy Data and Proxy Networks for Hyper-Parameter Optimization in Medical Image Segmentation
Jul 12, 2021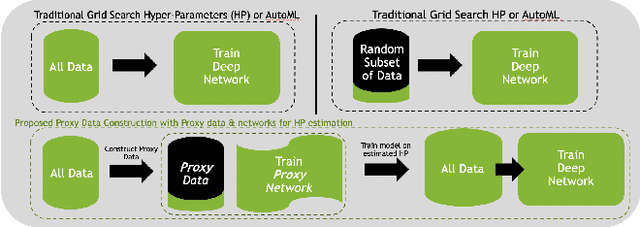
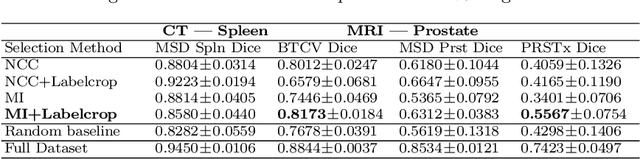
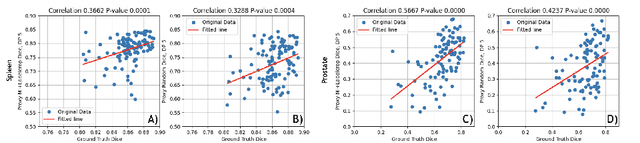
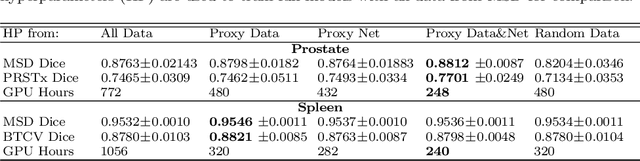
Abstract:Deep learning models for medical image segmentation are primarily data-driven. Models trained with more data lead to improved performance and generalizability. However, training is a computationally expensive process because multiple hyper-parameters need to be tested to find the optimal setting for best performance. In this work, we focus on accelerating the estimation of hyper-parameters by proposing two novel methodologies: proxy data and proxy networks. Both can be useful for estimating hyper-parameters more efficiently. We test the proposed techniques on CT and MR imaging modalities using well-known public datasets. In both cases using one dataset for building proxy data and another data source for external evaluation. For CT, the approach is tested on spleen segmentation with two datasets. The first dataset is from the medical segmentation decathlon (MSD), where the proxy data is constructed, the secondary dataset is utilized as an external validation dataset. Similarly, for MR, the approach is evaluated on prostate segmentation where the first dataset is from MSD and the second dataset is PROSTATEx. First, we show higher correlation to using full data for training when testing on the external validation set using smaller proxy data than a random selection of the proxy data. Second, we show that a high correlation exists for proxy networks when compared with the full network on validation Dice score. Third, we show that the proposed approach of utilizing a proxy network can speed up an AutoML framework for hyper-parameter search by 3.3x, and by 4.4x if proxy data and proxy network are utilized together.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge