Chenhui Wang
UCLA Department of Statistics, Los Angeles, CA
Bridging Brain and Semantics: A Hierarchical Framework for Semantically Enhanced fMRI-to-Video Reconstruction
May 14, 2026Abstract:Reconstructing dynamic visual experiences as videos from functional magnetic resonance imaging (fMRI) is pivotal for advancing the understanding of neural processes. However, current fMRI-to-video reconstruction methods are hindered by a semantic gap between noisy fMRI signals and the rich content of videos, stemming from a reliance on incomplete semantic embeddings that neither capture video-specific cues (e.g., actions) nor integrate prior knowledge. To this end, we draw inspiration from the dual-pathway processing mechanism in human brain and introduce CineNeuron, a novel hierarchical framework for semantically enhanced video reconstruction from fMRI signals with two synergistic stages. First, a bottom-up semantic enrichment stage maps fMRI signals to a rich embedding space that comprehensively captures textual semantics, image contents, action concepts, and object categories. Second, a top-down memory integration stage utilizes the proposed Mixture-of-Memories method to dynamically select relevant "memories" from previously seen data and fuse them with the fMRI embedding to refine the video reconstruction. Extensive experimental results on two fMRI-to-video benchmarks demonstrate that CineNeuron surpasses state-of-the-art methods across various metrics.
UniField: A Unified Field-Aware MRI Enhancement Framework
Mar 10, 2026Abstract:Magnetic Resonance Imaging (MRI) field-strength enhancement holds immense value for both clinical diagnostics and advanced research. However, existing methods typically focus on isolated enhancement tasks, such as specific 64mT-to-3T or 3T-to-7T transitions using limited subject cohorts, thereby failing to exploit the shared degradation patterns inherent across different field strengths and severely restricting model generalization. To address this challenge, we propose \methodname, a unified framework integrating multiple modalities and enhancement tasks to mutually promote representation learning by exploiting these shared degradation characteristics. Specifically, our main contributions are threefold. Firstly, to overcome MRI data scarcity and capture continuous anatomical structures, \methodname departs from conventional methods that treat 3D MRI volumes as independent 2D slices. Instead, we directly exploit comprehensive 3D volumetric information by leveraging pre-trained 3D foundation models, thereby embedding generalized and robust structural representations to significantly boost enhancement performance. In addition, to mitigate the spectral bias of mainstream flow-matching models that often over-smooth high-frequency details, we explicitly incorporate the physical mechanisms of magnetic fields to introduce a Field-Aware Spectral Rectification Mechanism (FASRM), tailoring customized spectral corrections to distinct field strengths. Finally, to resolve the fundamental data bottleneck, we organize and publicly release a comprehensive paired multi-field MRI dataset, which is an order of magnitude larger than existing datasets. Extensive experiments demonstrate our method's superiority over state-of-the-art approaches, achieving an average improvement of approximately 1.81 dB in PSNR and 9.47\% in SSIM. Code will be released upon acceptance.
Skill-Evolving Grounded Reasoning for Free-Text Promptable 3D Medical Image Segmentation
Mar 09, 2026Abstract:Free-text promptable 3D medical image segmentation offers an intuitive and clinically flexible interaction paradigm. However, current methods are highly sensitive to linguistic variability: minor changes in phrasing can cause substantial performance degradation despite identical clinical intent. Existing approaches attempt to improve robustness through stronger vision-language fusion or larger vocabularies, yet they lack mechanisms to consistently align ambiguous free-form expressions with anatomically grounded representations. We propose Skill-Evolving grounded Reasoning (SEER), a novel framework for free-text promptable 3D medical image segmentation that explicitly bridges linguistic variability and anatomical precision through a reasoning-driven design. First, we curate the SEER-Trace dataset, which pairs raw clinical requests with image-grounded, skill-tagged reasoning traces, establishing a reproducible benchmark. Second, SEER constructs an evidence-aligned target representation via a vision-language reasoning chain that verifies clinical intent against image-derived anatomical evidence, thereby enforcing semantic consistency before voxel-level decoding. Third, we introduce SEER-Loop, a dynamic skill-evolving strategy that distills high-reward reasoning trajectories into reusable skill artifacts and progressively integrates them into subsequent inference, enabling structured self-refinement and improved robustness to diverse linguistic expressions. Extensive experiments demonstrate superior performance of SEER over state-of-the-art baselines. Under linguistic perturbations, SEER reduces performance variance by 81.94% and improves worst-case Dice by 18.60%.
Brain-WM: Brain Glioblastoma World Model
Mar 08, 2026Abstract:Precise prognostic modeling of glioblastoma (GBM) under varying treatment interventions is essential for optimizing clinical outcomes. While generative AI has shown promise in simulating GBM evolution, existing methods typically treat interventions as static conditional inputs rather than dynamic decision variables. Consequently, they fail to capture the complex, reciprocal interplay between tumor evolution and treatment response. To bridge this gap, we present Brain-WM, a pioneering brain GBM world model that unifies next-step treatment prediction and future MRI generation, thereby capturing the co-evolutionary dynamics between tumor and treatment. Specifically, Brain-WM encodes spatiotemporal dynamics into a shared latent space for joint autoregressive treatment prediction and flow-based future MRI generation. Then, instead of a conventional monolithic framework, Brain-WM adopts a novel Y-shaped Mixture-of-Transformers (MoT) architecture. This design structurally disentangles heterogeneous objectives, successfully leveraging cross-task synergies while preventing feature collapse. Finally, a synergistic multi-timepoint mask alignment objective explicitly anchors latent representations to anatomically grounded tumor structures and progression-aware semantics. Extensive validation on internal and external multi-institutional cohorts demonstrates the superiority of Brain-WM, achieving 91.5% accuracy in treatment planning and SSIMs of 0.8524, 0.8581, and 0.8404 for FLAIR, T1CE, and T2W sequences, respectively. Ultimately, Brain-WM offers a robust clinical sandbox for optimizing patient healthcare. The source code is made available at https://github.com/thibault-wch/Brain-GBM-world-model.
Dynamic Differential Linear Attention: Enhancing Linear Diffusion Transformer for High-Quality Image Generation
Jan 20, 2026Abstract:Diffusion transformers (DiTs) have emerged as a powerful architecture for high-fidelity image generation, yet the quadratic cost of self-attention poses a major scalability bottleneck. To address this, linear attention mechanisms have been adopted to reduce computational cost; unfortunately, the resulting linear diffusion transformers (LiTs) models often come at the expense of generative performance, frequently producing over-smoothed attention weights that limit expressiveness. In this work, we introduce Dynamic Differential Linear Attention (DyDiLA), a novel linear attention formulation that enhances the effectiveness of LiTs by mitigating the oversmoothing issue and improving generation quality. Specifically, the novelty of DyDiLA lies in three key designs: (i) dynamic projection module, which facilitates the decoupling of token representations by learning with dynamically assigned knowledge; (ii) dynamic measure kernel, which provides a better similarity measurement to capture fine-grained semantic distinctions between tokens by dynamically assigning kernel functions for token processing; and (iii) token differential operator, which enables more robust query-to-key retrieval by calculating the differences between the tokens and their corresponding information redundancy produced by dynamic measure kernel. To capitalize on DyDiLA, we introduce a refined LiT, termed DyDi-LiT, that systematically incorporates our advancements. Extensive experiments show that DyDi-LiT consistently outperforms current state-of-the-art (SOTA) models across multiple metrics, underscoring its strong practical potential.
OneRec-V2 Technical Report
Aug 28, 2025



Abstract:Recent breakthroughs in generative AI have transformed recommender systems through end-to-end generation. OneRec reformulates recommendation as an autoregressive generation task, achieving high Model FLOPs Utilization. While OneRec-V1 has shown significant empirical success in real-world deployment, two critical challenges hinder its scalability and performance: (1) inefficient computational allocation where 97.66% of resources are consumed by sequence encoding rather than generation, and (2) limitations in reinforcement learning relying solely on reward models. To address these challenges, we propose OneRec-V2, featuring: (1) Lazy Decoder-Only Architecture: Eliminates encoder bottlenecks, reducing total computation by 94% and training resources by 90%, enabling successful scaling to 8B parameters. (2) Preference Alignment with Real-World User Interactions: Incorporates Duration-Aware Reward Shaping and Adaptive Ratio Clipping to better align with user preferences using real-world feedback. Extensive A/B tests on Kuaishou demonstrate OneRec-V2's effectiveness, improving App Stay Time by 0.467%/0.741% while balancing multi-objective recommendations. This work advances generative recommendation scalability and alignment with real-world feedback, representing a step forward in the development of end-to-end recommender systems.
LangMamba: A Language-driven Mamba Framework for Low-dose CT Denoising with Vision-language Models
Jul 08, 2025Abstract:Low-dose computed tomography (LDCT) reduces radiation exposure but often degrades image quality, potentially compromising diagnostic accuracy. Existing deep learning-based denoising methods focus primarily on pixel-level mappings, overlooking the potential benefits of high-level semantic guidance. Recent advances in vision-language models (VLMs) suggest that language can serve as a powerful tool for capturing structured semantic information, offering new opportunities to improve LDCT reconstruction. In this paper, we introduce LangMamba, a Language-driven Mamba framework for LDCT denoising that leverages VLM-derived representations to enhance supervision from normal-dose CT (NDCT). LangMamba follows a two-stage learning strategy. First, we pre-train a Language-guided AutoEncoder (LangAE) that leverages frozen VLMs to map NDCT images into a semantic space enriched with anatomical information. Second, we synergize LangAE with two key components to guide LDCT denoising: Semantic-Enhanced Efficient Denoiser (SEED), which enhances NDCT-relevant local semantic while capturing global features with efficient Mamba mechanism, and Language-engaged Dual-space Alignment (LangDA) Loss, which ensures that denoised images align with NDCT in both perceptual and semantic spaces. Extensive experiments on two public datasets demonstrate that LangMamba outperforms conventional state-of-the-art methods, significantly improving detail preservation and visual fidelity. Remarkably, LangAE exhibits strong generalizability to unseen datasets, thereby reducing training costs. Furthermore, LangDA loss improves explainability by integrating language-guided insights into image reconstruction and offers a plug-and-play fashion. Our findings shed new light on the potential of language as a supervisory signal to advance LDCT denoising. The code is publicly available on https://github.com/hao1635/LangMamba.
OneRec Technical Report
Jun 16, 2025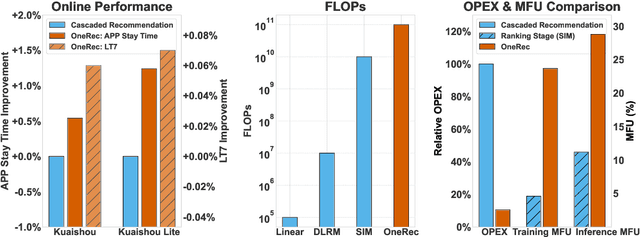

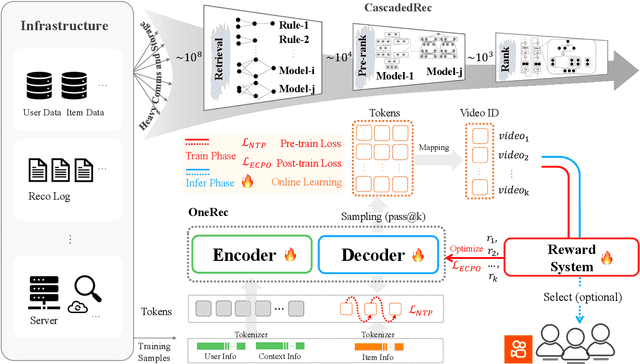
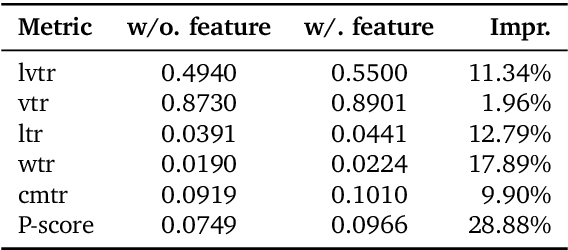
Abstract:Recommender systems have been widely used in various large-scale user-oriented platforms for many years. However, compared to the rapid developments in the AI community, recommendation systems have not achieved a breakthrough in recent years. For instance, they still rely on a multi-stage cascaded architecture rather than an end-to-end approach, leading to computational fragmentation and optimization inconsistencies, and hindering the effective application of key breakthrough technologies from the AI community in recommendation scenarios. To address these issues, we propose OneRec, which reshapes the recommendation system through an end-to-end generative approach and achieves promising results. Firstly, we have enhanced the computational FLOPs of the current recommendation model by 10 $\times$ and have identified the scaling laws for recommendations within certain boundaries. Secondly, reinforcement learning techniques, previously difficult to apply for optimizing recommendations, show significant potential in this framework. Lastly, through infrastructure optimizations, we have achieved 23.7% and 28.8% Model FLOPs Utilization (MFU) on flagship GPUs during training and inference, respectively, aligning closely with the LLM community. This architecture significantly reduces communication and storage overhead, resulting in operating expense that is only 10.6% of traditional recommendation pipelines. Deployed in Kuaishou/Kuaishou Lite APP, it handles 25% of total queries per second, enhancing overall App Stay Time by 0.54% and 1.24%, respectively. Additionally, we have observed significant increases in metrics such as 7-day Lifetime, which is a crucial indicator of recommendation experience. We also provide practical lessons and insights derived from developing, optimizing, and maintaining a production-scale recommendation system with significant real-world impact.
MedITok: A Unified Tokenizer for Medical Image Synthesis and Interpretation
May 25, 2025Abstract:Advanced autoregressive models have reshaped multimodal AI. However, their transformative potential in medical imaging remains largely untapped due to the absence of a unified visual tokenizer -- one capable of capturing fine-grained visual structures for faithful image reconstruction and realistic image synthesis, as well as rich semantics for accurate diagnosis and image interpretation. To this end, we present MedITok, the first unified tokenizer tailored for medical images, encoding both low-level structural details and high-level clinical semantics within a unified latent space. To balance these competing objectives, we introduce a novel two-stage training framework: a visual representation alignment stage that cold-starts the tokenizer reconstruction learning with a visual semantic constraint, followed by a textual semantic representation alignment stage that infuses detailed clinical semantics into the latent space. Trained on the meticulously collected large-scale dataset with over 30 million medical images and 2 million image-caption pairs, MedITok achieves state-of-the-art performance on more than 30 datasets across 9 imaging modalities and 4 different tasks. By providing a unified token space for autoregressive modeling, MedITok supports a wide range of tasks in clinical diagnostics and generative healthcare applications. Model and code will be made publicly available at: https://github.com/Masaaki-75/meditok.
Autoregressive Medical Image Segmentation via Next-Scale Mask Prediction
Feb 28, 2025



Abstract:While deep learning has significantly advanced medical image segmentation, most existing methods still struggle with handling complex anatomical regions. Cascaded or deep supervision-based approaches attempt to address this challenge through multi-scale feature learning but fail to establish sufficient inter-scale dependencies, as each scale relies solely on the features of the immediate predecessor. To this end, we propose the AutoRegressive Segmentation framework via next-scale mask prediction, termed AR-Seg, which progressively predicts the next-scale mask by explicitly modeling dependencies across all previous scales within a unified architecture. AR-Seg introduces three innovations: (1) a multi-scale mask autoencoder that quantizes the mask into multi-scale token maps to capture hierarchical anatomical structures, (2) a next-scale autoregressive mechanism that progressively predicts next-scale masks to enable sufficient inter-scale dependencies, and (3) a consensus-aggregation strategy that combines multiple sampled results to generate a more accurate mask, further improving segmentation robustness. Extensive experimental results on two benchmark datasets with different modalities demonstrate that AR-Seg outperforms state-of-the-art methods while explicitly visualizing the intermediate coarse-to-fine segmentation process.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge