Ben Glocker
Biomedical Image Analysis Group, Department of Computing, Imperial College London
Bayesian analysis of the prevalence bias: learning and predicting from imbalanced data
Jul 31, 2021
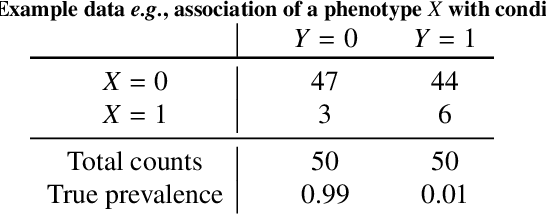
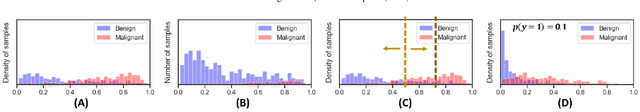
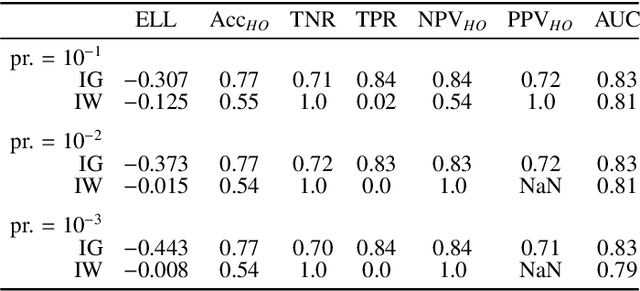
Abstract:Datasets are rarely a realistic approximation of the target population. Say, prevalence is misrepresented, image quality is above clinical standards, etc. This mismatch is known as sampling bias. Sampling biases are a major hindrance for machine learning models. They cause significant gaps between model performance in the lab and in the real world. Our work is a solution to prevalence bias. Prevalence bias is the discrepancy between the prevalence of a pathology and its sampling rate in the training dataset, introduced upon collecting data or due to the practioner rebalancing the training batches. This paper lays the theoretical and computational framework for training models, and for prediction, in the presence of prevalence bias. Concretely a bias-corrected loss function, as well as bias-corrected predictive rules, are derived under the principles of Bayesian risk minimization. The loss exhibits a direct connection to the information gain. It offers a principled alternative to heuristic training losses and complements test-time procedures based on selecting an operating point from summary curves. It integrates seamlessly in the current paradigm of (deep) learning using stochastic backpropagation and naturally with Bayesian models.
A Review of Generative Adversarial Networks in Cancer Imaging: New Applications, New Solutions
Jul 20, 2021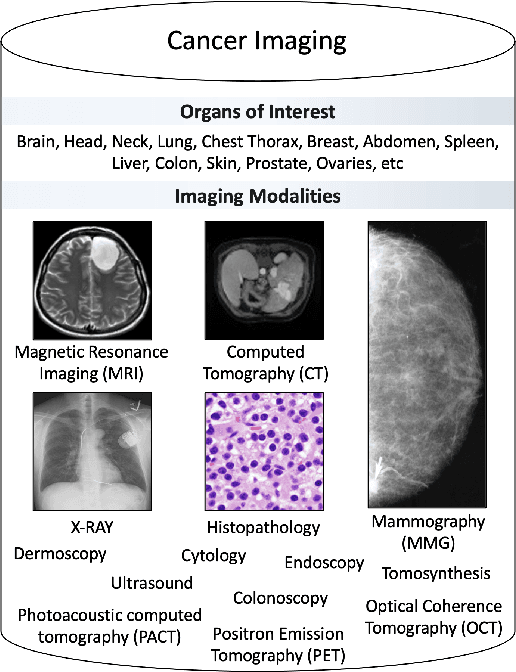
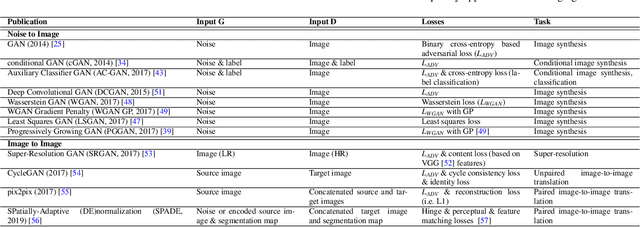
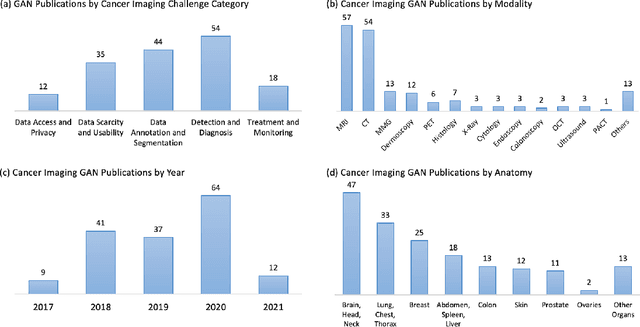
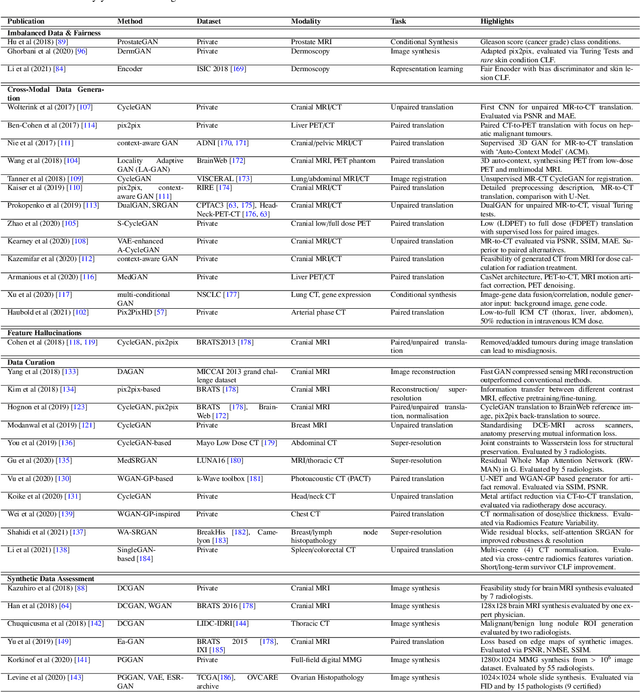
Abstract:Despite technological and medical advances, the detection, interpretation, and treatment of cancer based on imaging data continue to pose significant challenges. These include high inter-observer variability, difficulty of small-sized lesion detection, nodule interpretation and malignancy determination, inter- and intra-tumour heterogeneity, class imbalance, segmentation inaccuracies, and treatment effect uncertainty. The recent advancements in Generative Adversarial Networks (GANs) in computer vision as well as in medical imaging may provide a basis for enhanced capabilities in cancer detection and analysis. In this review, we assess the potential of GANs to address a number of key challenges of cancer imaging, including data scarcity and imbalance, domain and dataset shifts, data access and privacy, data annotation and quantification, as well as cancer detection, tumour profiling and treatment planning. We provide a critical appraisal of the existing literature of GANs applied to cancer imagery, together with suggestions on future research directions to address these challenges. We analyse and discuss 163 papers that apply adversarial training techniques in the context of cancer imaging and elaborate their methodologies, advantages and limitations. With this work, we strive to bridge the gap between the needs of the clinical cancer imaging community and the current and prospective research on GANs in the artificial intelligence community.
Multiple Instance Learning with Auxiliary Task Weighting for Multiple Myeloma Classification
Jul 16, 2021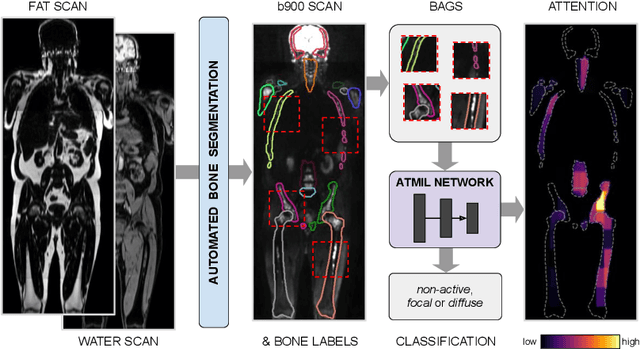
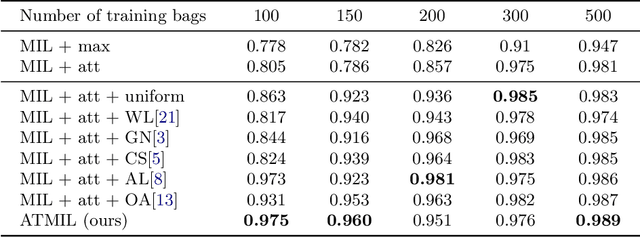
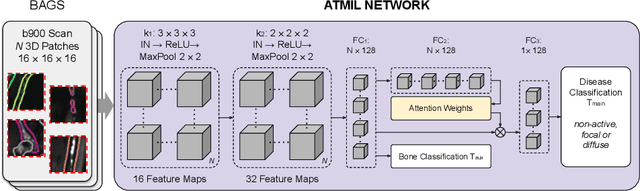
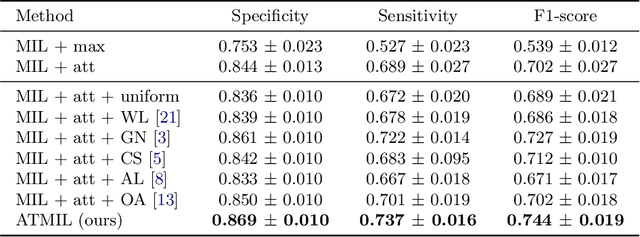
Abstract:Whole body magnetic resonance imaging (WB-MRI) is the recommended modality for diagnosis of multiple myeloma (MM). WB-MRI is used to detect sites of disease across the entire skeletal system, but it requires significant expertise and is time-consuming to report due to the great number of images. To aid radiological reading, we propose an auxiliary task-based multiple instance learning approach (ATMIL) for MM classification with the ability to localize sites of disease. This approach is appealing as it only requires patient-level annotations where an attention mechanism is used to identify local regions with active disease. We borrow ideas from multi-task learning and define an auxiliary task with adaptive reweighting to support and improve learning efficiency in the presence of data scarcity. We validate our approach on both synthetic and real multi-center clinical data. We show that the MIL attention module provides a mechanism to localize bone regions while the adaptive reweighting of the auxiliary task considerably improves the performance.
Learning from Partially Overlapping Labels: Image Segmentation under Annotation Shift
Jul 13, 2021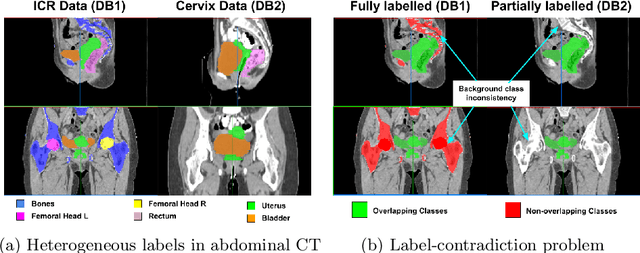
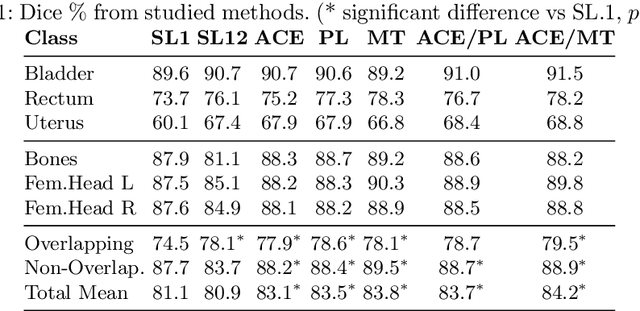
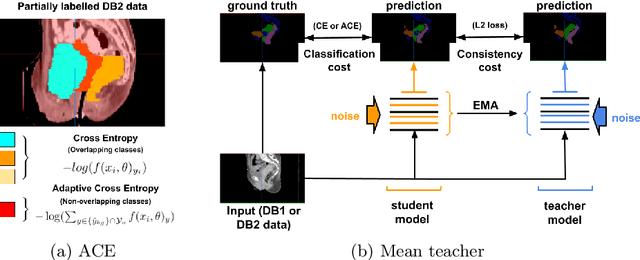
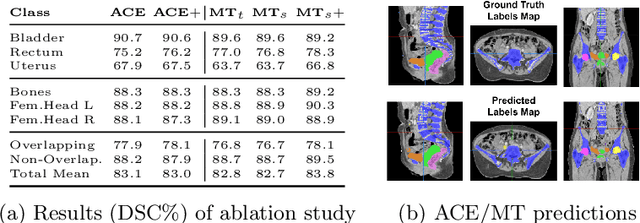
Abstract:Scarcity of high quality annotated images remains a limiting factor for training accurate image segmentation models. While more and more annotated datasets become publicly available, the number of samples in each individual database is often small. Combining different databases to create larger amounts of training data is appealing yet challenging due to the heterogeneity as a result of differences in data acquisition and annotation processes, often yielding incompatible or even conflicting information. In this paper, we investigate and propose several strategies for learning from partially overlapping labels in the context of abdominal organ segmentation. We find that combining a semi-supervised approach with an adaptive cross entropy loss can successfully exploit heterogeneously annotated data and substantially improve segmentation accuracy compared to baseline and alternative approaches.
Detecting Hypo-plastic Left Heart Syndrome in Fetal Ultrasound via Disease-specific Atlas Maps
Jul 06, 2021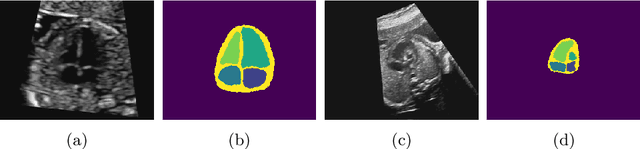
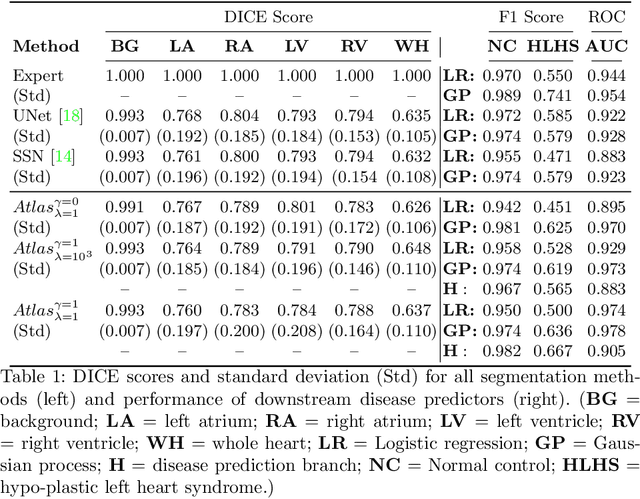
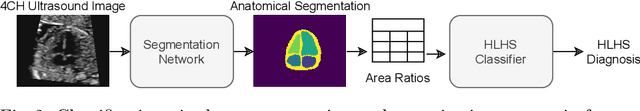
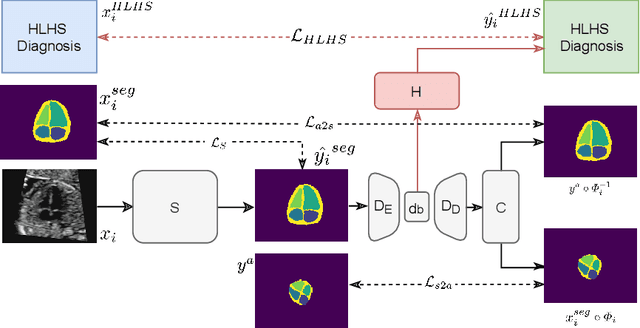
Abstract:Fetal ultrasound screening during pregnancy plays a vital role in the early detection of fetal malformations which have potential long-term health impacts. The level of skill required to diagnose such malformations from live ultrasound during examination is high and resources for screening are often limited. We present an interpretable, atlas-learning segmentation method for automatic diagnosis of Hypo-plastic Left Heart Syndrome (HLHS) from a single `4 Chamber Heart' view image. We propose to extend the recently introduced Image-and-Spatial Transformer Networks (Atlas-ISTN) into a framework that enables sensitising atlas generation to disease. In this framework we can jointly learn image segmentation, registration, atlas construction and disease prediction while providing a maximum level of clinical interpretability compared to direct image classification methods. As a result our segmentation allows diagnoses competitive with expert-derived manual diagnosis and yields an AUC-ROC of 0.978 (1043 cases for training, 260 for validation and 325 for testing).
Confidence-based Out-of-Distribution Detection: A Comparative Study and Analysis
Jul 06, 2021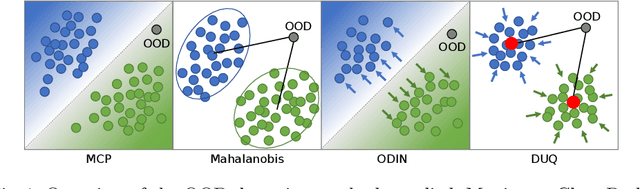
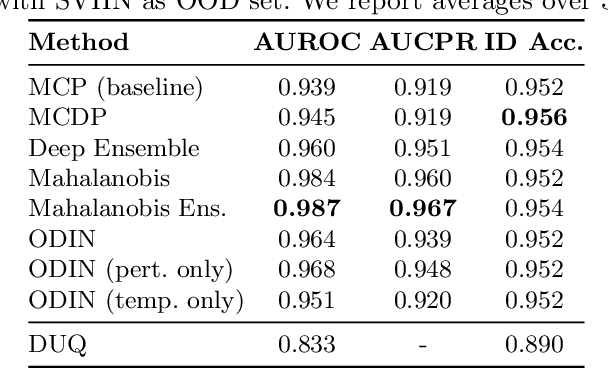
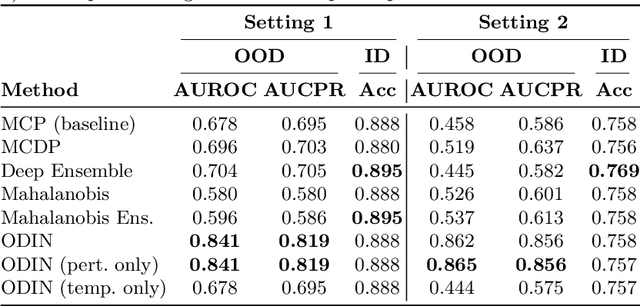
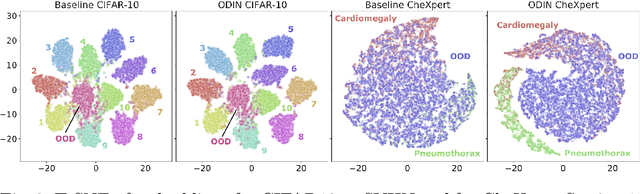
Abstract:Image classification models deployed in the real world may receive inputs outside the intended data distribution. For critical applications such as clinical decision making, it is important that a model can detect such out-of-distribution (OOD) inputs and express its uncertainty. In this work, we assess the capability of various state-of-the-art approaches for confidence-based OOD detection through a comparative study and in-depth analysis. First, we leverage a computer vision benchmark to reproduce and compare multiple OOD detection methods. We then evaluate their capabilities on the challenging task of disease classification using chest X-rays. Our study shows that high performance in a computer vision task does not directly translate to accuracy in a medical imaging task. We analyse factors that affect performance of the methods between the two tasks. Our results provide useful insights for developing the next generation of OOD detection methods.
Distributional Gaussian Process Layers for Outlier Detection in Image Segmentation
Apr 28, 2021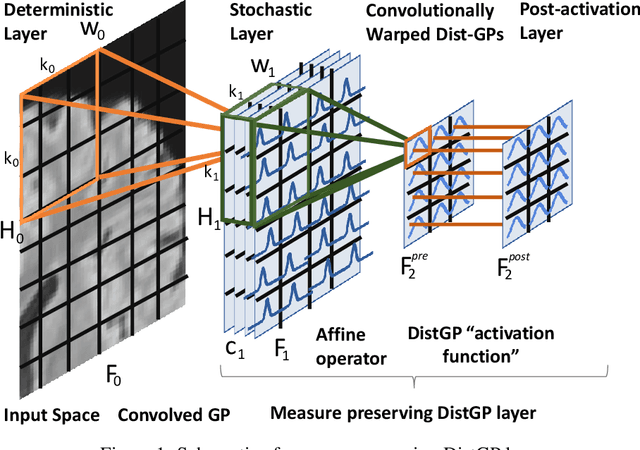
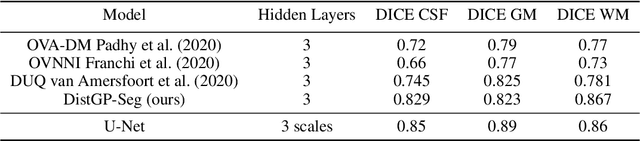

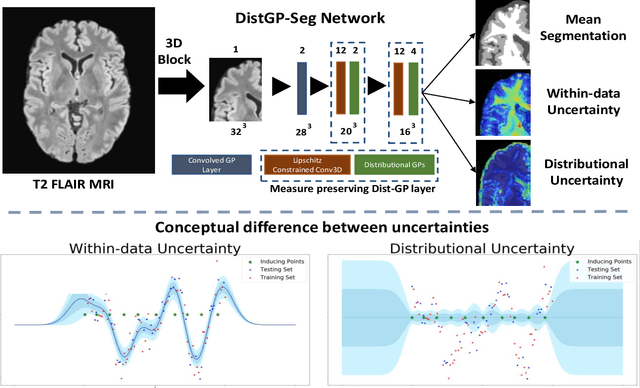
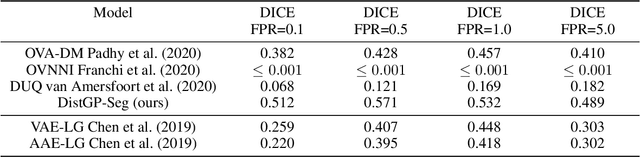
Abstract:We propose a parameter efficient Bayesian layer for hierarchical convolutional Gaussian Processes that incorporates Gaussian Processes operating in Wasserstein-2 space to reliably propagate uncertainty. This directly replaces convolving Gaussian Processes with a distance-preserving affine operator on distributions. Our experiments on brain tissue-segmentation show that the resulting architecture approaches the performance of well-established deterministic segmentation algorithms (U-Net), which has never been achieved with previous hierarchical Gaussian Processes. Moreover, by applying the same segmentation model to out-of-distribution data (i.e., images with pathology such as brain tumors), we show that our uncertainty estimates result in out-of-distribution detection that outperforms the capabilities of previous Bayesian networks and reconstruction-based approaches that learn normative distributions.
Common Limitations of Image Processing Metrics: A Picture Story
Apr 13, 2021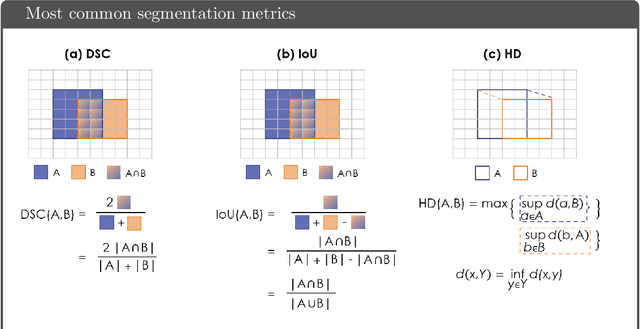
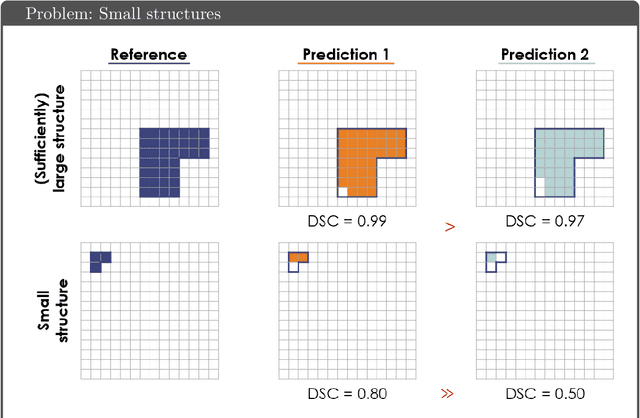
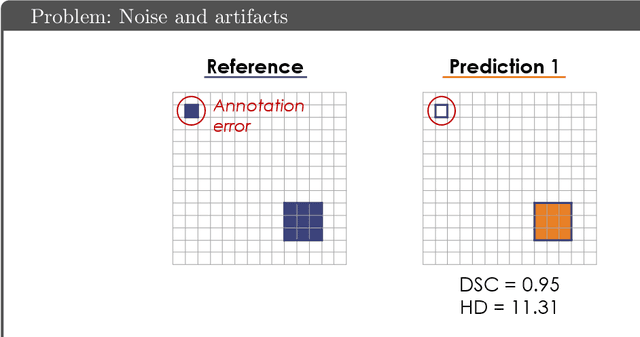
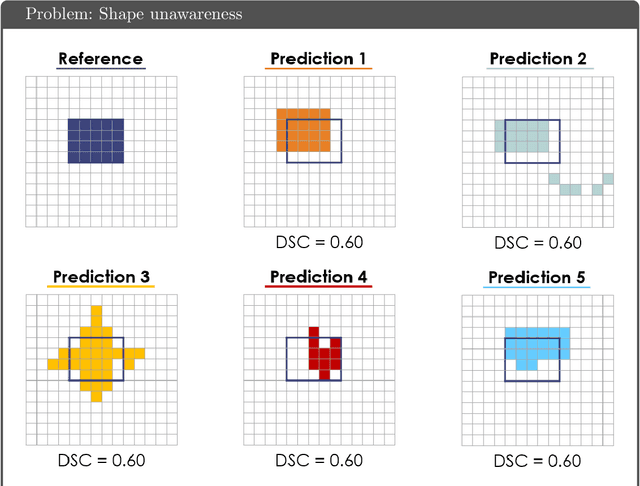
Abstract:While the importance of automatic image analysis is increasing at an enormous pace, recent meta-research revealed major flaws with respect to algorithm validation. Specifically, performance metrics are key for objective, transparent and comparative performance assessment, but relatively little attention has been given to the practical pitfalls when using specific metrics for a given image analysis task. A common mission of several international initiatives is therefore to provide researchers with guidelines and tools to choose the performance metrics in a problem-aware manner. This dynamically updated document has the purpose to illustrate important limitations of performance metrics commonly applied in the field of image analysis. The current version is based on a Delphi process on metrics conducted by an international consortium of image analysis experts.
Spatially Varying Label Smoothing: Capturing Uncertainty from Expert Annotations
Apr 12, 2021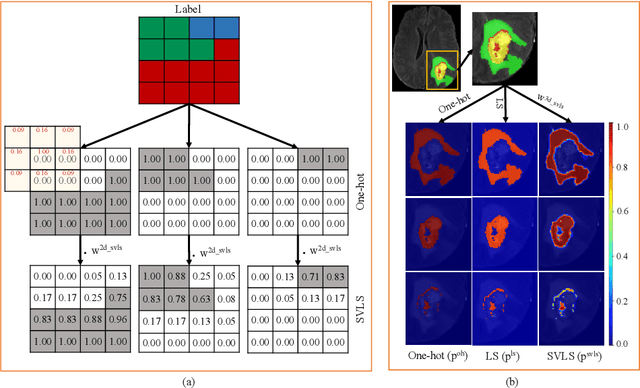
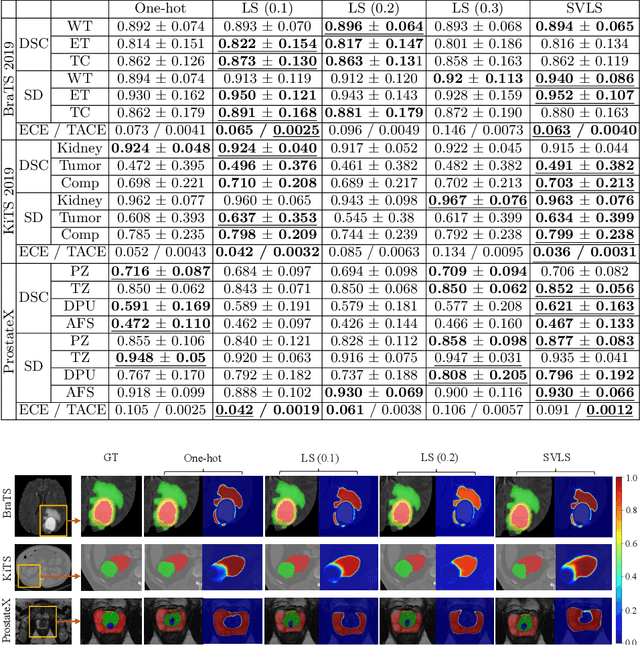
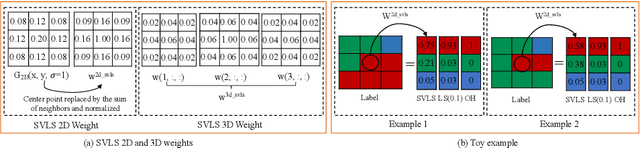
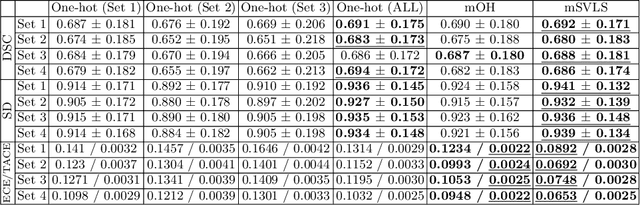
Abstract:The task of image segmentation is inherently noisy due to ambiguities regarding the exact location of boundaries between anatomical structures. We argue that this information can be extracted from the expert annotations at no extra cost, and when integrated into state-of-the-art neural networks, it can lead to improved calibration between soft probabilistic predictions and the underlying uncertainty. We built upon label smoothing (LS) where a network is trained on 'blurred' versions of the ground truth labels which has been shown to be effective for calibrating output predictions. However, LS is not taking the local structure into account and results in overly smoothed predictions with low confidence even for non-ambiguous regions. Here, we propose Spatially Varying Label Smoothing (SVLS), a soft labeling technique that captures the structural uncertainty in semantic segmentation. SVLS also naturally lends itself to incorporate inter-rater uncertainty when multiple labelmaps are available. The proposed approach is extensively validated on four clinical segmentation tasks with different imaging modalities, number of classes and single and multi-rater expert annotations. The results demonstrate that SVLS, despite its simplicity, obtains superior boundary prediction with improved uncertainty and model calibration.
Analyzing Overfitting under Class Imbalance in Neural Networks for Image Segmentation
Feb 20, 2021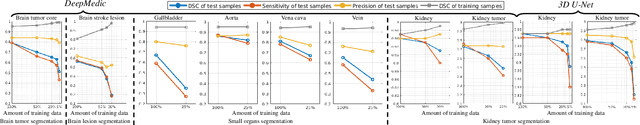
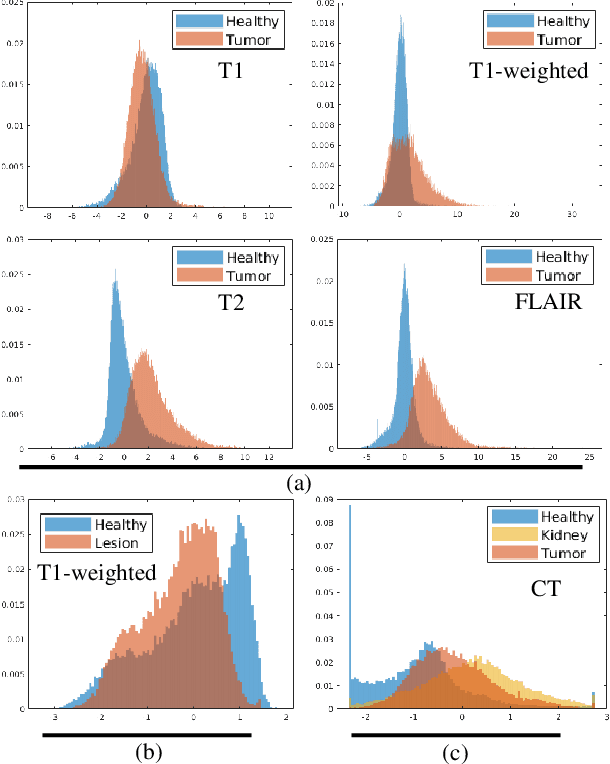
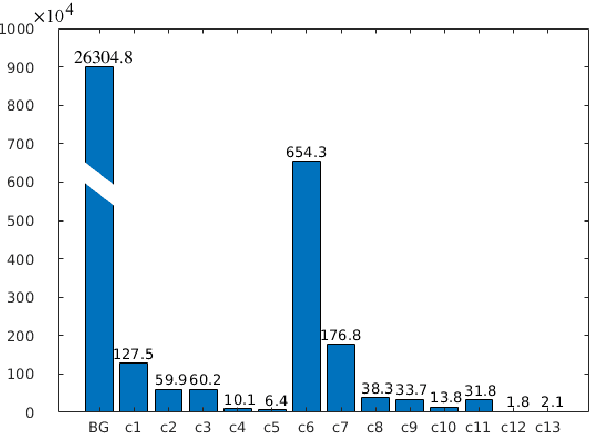
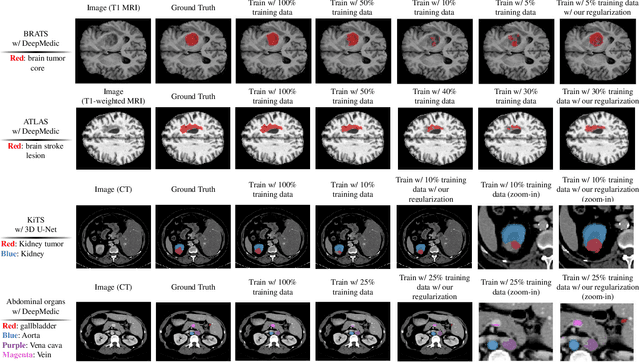
Abstract:Class imbalance poses a challenge for developing unbiased, accurate predictive models. In particular, in image segmentation neural networks may overfit to the foreground samples from small structures, which are often heavily under-represented in the training set, leading to poor generalization. In this study, we provide new insights on the problem of overfitting under class imbalance by inspecting the network behavior. We find empirically that when training with limited data and strong class imbalance, at test time the distribution of logit activations may shift across the decision boundary, while samples of the well-represented class seem unaffected. This bias leads to a systematic under-segmentation of small structures. This phenomenon is consistently observed for different databases, tasks and network architectures. To tackle this problem, we introduce new asymmetric variants of popular loss functions and regularization techniques including a large margin loss, focal loss, adversarial training, mixup and data augmentation, which are explicitly designed to counter logit shift of the under-represented classes. Extensive experiments are conducted on several challenging segmentation tasks. Our results demonstrate that the proposed modifications to the objective function can lead to significantly improved segmentation accuracy compared to baselines and alternative approaches.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge