Theodore Barfoot
Average Calibration Losses for Reliable Uncertainty in Medical Image Segmentation
Jun 04, 2025Abstract:Deep neural networks for medical image segmentation are often overconfident, compromising both reliability and clinical utility. In this work, we propose differentiable formulations of marginal L1 Average Calibration Error (mL1-ACE) as an auxiliary loss that can be computed on a per-image basis. We compare both hard- and soft-binning approaches to directly improve pixel-wise calibration. Our experiments on four datasets (ACDC, AMOS, KiTS, BraTS) demonstrate that incorporating mL1-ACE significantly reduces calibration errors, particularly Average Calibration Error (ACE) and Maximum Calibration Error (MCE), while largely maintaining high Dice Similarity Coefficients (DSCs). We find that the soft-binned variant yields the greatest improvements in calibration, over the Dice plus cross-entropy loss baseline, but often compromises segmentation performance, with hard-binned mL1-ACE maintaining segmentation performance, albeit with weaker calibration improvement. To gain further insight into calibration performance and its variability across an imaging dataset, we introduce dataset reliability histograms, an aggregation of per-image reliability diagrams. The resulting analysis highlights improved alignment between predicted confidences and true accuracies. Overall, our approach not only enhances the trustworthiness of segmentation predictions but also shows potential for safer integration of deep learning methods into clinical workflows. We share our code here: https://github.com/cai4cai/Average-Calibration-Losses
Calibration and Uncertainty for multiRater Volume Assessment in multiorgan Segmentation (CURVAS) challenge results
May 13, 2025



Abstract:Deep learning (DL) has become the dominant approach for medical image segmentation, yet ensuring the reliability and clinical applicability of these models requires addressing key challenges such as annotation variability, calibration, and uncertainty estimation. This is why we created the Calibration and Uncertainty for multiRater Volume Assessment in multiorgan Segmentation (CURVAS), which highlights the critical role of multiple annotators in establishing a more comprehensive ground truth, emphasizing that segmentation is inherently subjective and that leveraging inter-annotator variability is essential for robust model evaluation. Seven teams participated in the challenge, submitting a variety of DL models evaluated using metrics such as Dice Similarity Coefficient (DSC), Expected Calibration Error (ECE), and Continuous Ranked Probability Score (CRPS). By incorporating consensus and dissensus ground truth, we assess how DL models handle uncertainty and whether their confidence estimates align with true segmentation performance. Our findings reinforce the importance of well-calibrated models, as better calibration is strongly correlated with the quality of the results. Furthermore, we demonstrate that segmentation models trained on diverse datasets and enriched with pre-trained knowledge exhibit greater robustness, particularly in cases deviating from standard anatomical structures. Notably, the best-performing models achieved high DSC and well-calibrated uncertainty estimates. This work underscores the need for multi-annotator ground truth, thorough calibration assessments, and uncertainty-aware evaluations to develop trustworthy and clinically reliable DL-based medical image segmentation models.
An investigation into the causes of race bias in AI-based cine CMR segmentation
Aug 05, 2024



Abstract:Artificial intelligence (AI) methods are being used increasingly for the automated segmentation of cine cardiac magnetic resonance (CMR) imaging. However, these methods have been shown to be subject to race bias, i.e. they exhibit different levels of performance for different races depending on the (im)balance of the data used to train the AI model. In this paper we investigate the source of this bias, seeking to understand its root cause(s) so that it can be effectively mitigated. We perform a series of classification and segmentation experiments on short-axis cine CMR images acquired from Black and White subjects from the UK Biobank and apply AI interpretability methods to understand the results. In the classification experiments, we found that race can be predicted with high accuracy from the images alone, but less accurately from ground truth segmentations, suggesting that the distributional shift between races, which is often the cause of AI bias, is mostly image-based rather than segmentation-based. The interpretability methods showed that most attention in the classification models was focused on non-heart regions, such as subcutaneous fat. Cropping the images tightly around the heart reduced classification accuracy to around chance level. Similarly, race can be predicted from the latent representations of a biased segmentation model, suggesting that race information is encoded in the model. Cropping images tightly around the heart reduced but did not eliminate segmentation bias. We also investigate the influence of possible confounders on the bias observed.
Average Calibration Error: A Differentiable Loss for Improved Reliability in Image Segmentation
Mar 11, 2024Abstract:Deep neural networks for medical image segmentation often produce overconfident results misaligned with empirical observations. Such miscalibration, challenges their clinical translation. We propose to use marginal L1 average calibration error (mL1-ACE) as a novel auxiliary loss function to improve pixel-wise calibration without compromising segmentation quality. We show that this loss, despite using hard binning, is directly differentiable, bypassing the need for approximate but differentiable surrogate or soft binning approaches. Our work also introduces the concept of dataset reliability histograms which generalises standard reliability diagrams for refined visual assessment of calibration in semantic segmentation aggregated at the dataset level. Using mL1-ACE, we reduce average and maximum calibration error by 45% and 55% respectively, maintaining a Dice score of 87% on the BraTS 2021 dataset. We share our code here: https://github.com/cai4cai/ACE-DLIRIS
Multiple Instance Learning with Auxiliary Task Weighting for Multiple Myeloma Classification
Jul 16, 2021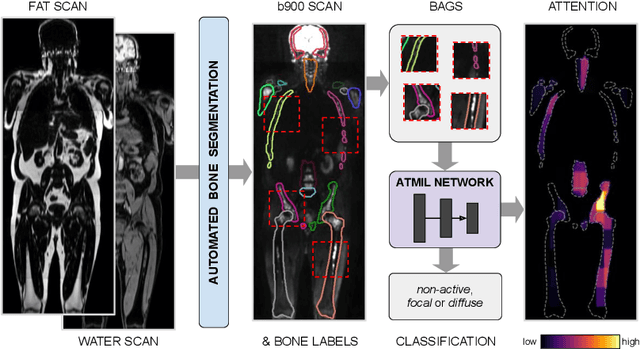
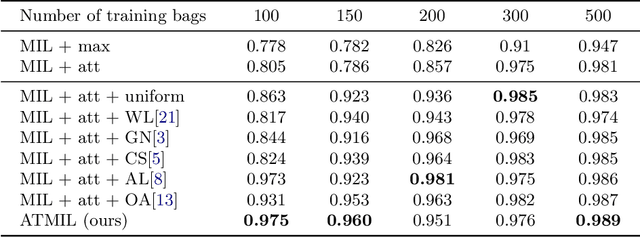
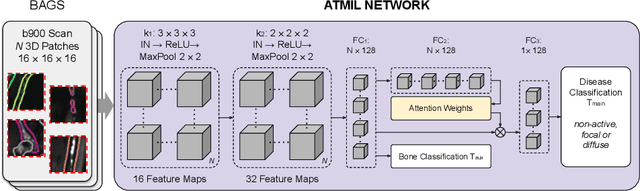
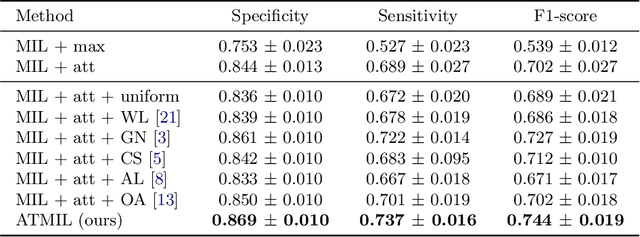
Abstract:Whole body magnetic resonance imaging (WB-MRI) is the recommended modality for diagnosis of multiple myeloma (MM). WB-MRI is used to detect sites of disease across the entire skeletal system, but it requires significant expertise and is time-consuming to report due to the great number of images. To aid radiological reading, we propose an auxiliary task-based multiple instance learning approach (ATMIL) for MM classification with the ability to localize sites of disease. This approach is appealing as it only requires patient-level annotations where an attention mechanism is used to identify local regions with active disease. We borrow ideas from multi-task learning and define an auxiliary task with adaptive reweighting to support and improve learning efficiency in the presence of data scarcity. We validate our approach on both synthetic and real multi-center clinical data. We show that the MIL attention module provides a mechanism to localize bone regions while the adaptive reweighting of the auxiliary task considerably improves the performance.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge