Allen Nie
Shammie
LLF-Bench: Benchmark for Interactive Learning from Language Feedback
Dec 13, 2023Abstract:We introduce a new benchmark, LLF-Bench (Learning from Language Feedback Benchmark; pronounced as "elf-bench"), to evaluate the ability of AI agents to interactively learn from natural language feedback and instructions. Learning from language feedback (LLF) is essential for people, largely because the rich information this feedback provides can help a learner avoid much of trial and error and thereby speed up the learning process. Large Language Models (LLMs) have recently enabled AI agents to comprehend natural language -- and hence AI agents can potentially benefit from language feedback during learning like humans do. But existing interactive benchmarks do not assess this crucial capability: they either use numeric reward feedback or require no learning at all (only planning or information retrieval). LLF-Bench is designed to fill this omission. LLF-Bench is a diverse collection of sequential decision-making tasks that includes user recommendation, poem writing, navigation, and robot control. The objective of an agent is to interactively solve these tasks based on their natural-language instructions and the feedback received after taking actions. Crucially, to ensure that the agent actually "learns" from the feedback, LLF-Bench implements several randomization techniques (such as paraphrasing and environment randomization) to ensure that the task isn't familiar to the agent and that the agent is robust to various verbalizations. In addition, LLF-Bench provides a unified OpenAI Gym interface for all its tasks and allows the users to easily configure the information the feedback conveys (among suggestion, explanation, and instantaneous performance) to study how agents respond to different types of feedback. Together, these features make LLF-Bench a unique research platform for developing and testing LLF agents.
MoCa: Measuring Human-Language Model Alignment on Causal and Moral Judgment Tasks
Oct 31, 2023



Abstract:Human commonsense understanding of the physical and social world is organized around intuitive theories. These theories support making causal and moral judgments. When something bad happens, we naturally ask: who did what, and why? A rich literature in cognitive science has studied people's causal and moral intuitions. This work has revealed a number of factors that systematically influence people's judgments, such as the violation of norms and whether the harm is avoidable or inevitable. We collected a dataset of stories from 24 cognitive science papers and developed a system to annotate each story with the factors they investigated. Using this dataset, we test whether large language models (LLMs) make causal and moral judgments about text-based scenarios that align with those of human participants. On the aggregate level, alignment has improved with more recent LLMs. However, using statistical analyses, we find that LLMs weigh the different factors quite differently from human participants. These results show how curated, challenge datasets combined with insights from cognitive science can help us go beyond comparisons based merely on aggregate metrics: we uncover LLMs implicit tendencies and show to what extent these align with human intuitions.
Waypoint Transformer: Reinforcement Learning via Supervised Learning with Intermediate Targets
Jun 24, 2023



Abstract:Despite the recent advancements in offline reinforcement learning via supervised learning (RvS) and the success of the decision transformer (DT) architecture in various domains, DTs have fallen short in several challenging benchmarks. The root cause of this underperformance lies in their inability to seamlessly connect segments of suboptimal trajectories. To overcome this limitation, we present a novel approach to enhance RvS methods by integrating intermediate targets. We introduce the Waypoint Transformer (WT), using an architecture that builds upon the DT framework and conditioned on automatically-generated waypoints. The results show a significant increase in the final return compared to existing RvS methods, with performance on par or greater than existing state-of-the-art temporal difference learning-based methods. Additionally, the performance and stability improvements are largest in the most challenging environments and data configurations, including AntMaze Large Play/Diverse and Kitchen Mixed/Partial.
Reinforcement Learning Tutor Better Supported Lower Performers in a Math Task
Apr 13, 2023



Abstract:Resource limitations make it hard to provide all students with one of the most effective educational interventions: personalized instruction. Reinforcement learning could be a key tool to reduce the development cost and improve the effectiveness of intelligent tutoring software that aims to provide the right support, at the right time, to a student. Here we illustrate that deep reinforcement learning can be used to provide adaptive pedagogical support to students learning about the concept of volume in a narrative storyline software. Using explainable artificial intelligence tools, we extracted interpretable insights about the pedagogical policy learned and demonstrated that the resulting policy had similar performance in a different student population. Most importantly, in both studies, the reinforcement-learning narrative system had the largest benefit for those students with the lowest initial pretest scores, suggesting the opportunity for AI to adapt and provide support for those most in need.
Model-based Offline Reinforcement Learning with Local Misspecification
Jan 26, 2023Abstract:We present a model-based offline reinforcement learning policy performance lower bound that explicitly captures dynamics model misspecification and distribution mismatch and we propose an empirical algorithm for optimal offline policy selection. Theoretically, we prove a novel safe policy improvement theorem by establishing pessimism approximations to the value function. Our key insight is to jointly consider selecting over dynamics models and policies: as long as a dynamics model can accurately represent the dynamics of the state-action pairs visited by a given policy, it is possible to approximate the value of that particular policy. We analyze our lower bound in the LQR setting and also show competitive performance to previous lower bounds on policy selection across a set of D4RL tasks.
Giving Feedback on Interactive Student Programs with Meta-Exploration
Nov 16, 2022



Abstract:Developing interactive software, such as websites or games, is a particularly engaging way to learn computer science. However, teaching and giving feedback on such software is time-consuming -- standard approaches require instructors to manually grade student-implemented interactive programs. As a result, online platforms that serve millions, like Code.org, are unable to provide any feedback on assignments for implementing interactive programs, which critically hinders students' ability to learn. One approach toward automatic grading is to learn an agent that interacts with a student's program and explores states indicative of errors via reinforcement learning. However, existing work on this approach only provides binary feedback of whether a program is correct or not, while students require finer-grained feedback on the specific errors in their programs to understand their mistakes. In this work, we show that exploring to discover errors can be cast as a meta-exploration problem. This enables us to construct a principled objective for discovering errors and an algorithm for optimizing this objective, which provides fine-grained feedback. We evaluate our approach on a set of over 700K real anonymized student programs from a Code.org interactive assignment. Our approach provides feedback with 94.3% accuracy, improving over existing approaches by 17.7% and coming within 1.5% of human-level accuracy. Project web page: https://ezliu.github.io/dreamgrader.
Data-Efficient Pipeline for Offline Reinforcement Learning with Limited Data
Oct 16, 2022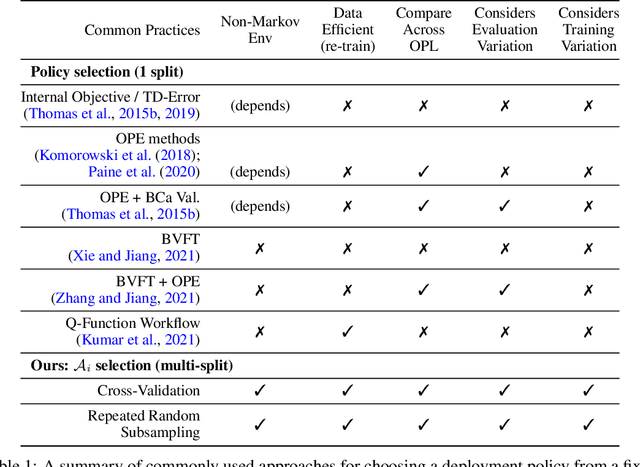
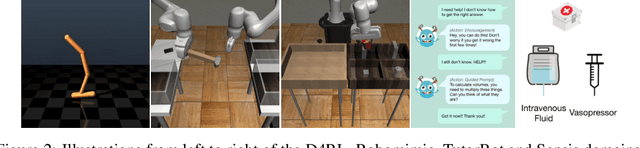
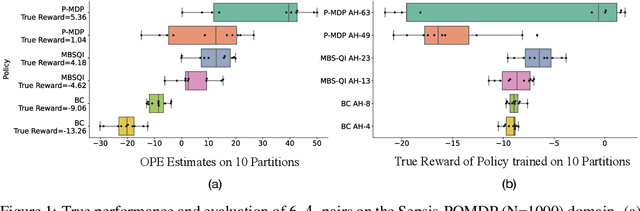

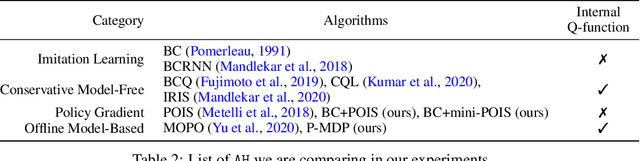
Abstract:Offline reinforcement learning (RL) can be used to improve future performance by leveraging historical data. There exist many different algorithms for offline RL, and it is well recognized that these algorithms, and their hyperparameter settings, can lead to decision policies with substantially differing performance. This prompts the need for pipelines that allow practitioners to systematically perform algorithm-hyperparameter selection for their setting. Critically, in most real-world settings, this pipeline must only involve the use of historical data. Inspired by statistical model selection methods for supervised learning, we introduce a task- and method-agnostic pipeline for automatically training, comparing, selecting, and deploying the best policy when the provided dataset is limited in size. In particular, our work highlights the importance of performing multiple data splits to produce more reliable algorithm-hyperparameter selection. While this is a common approach in supervised learning, to our knowledge, this has not been discussed in detail in the offline RL setting. We show it can have substantial impacts when the dataset is small. Compared to alternate approaches, our proposed pipeline outputs higher-performing deployed policies from a broad range of offline policy learning algorithms and across various simulation domains in healthcare, education, and robotics. This work contributes toward the development of a general-purpose meta-algorithm for automatic algorithm-hyperparameter selection for offline RL.
Beyond the Imitation Game: Quantifying and extrapolating the capabilities of language models
Jun 10, 2022Abstract:Language models demonstrate both quantitative improvement and new qualitative capabilities with increasing scale. Despite their potentially transformative impact, these new capabilities are as yet poorly characterized. In order to inform future research, prepare for disruptive new model capabilities, and ameliorate socially harmful effects, it is vital that we understand the present and near-future capabilities and limitations of language models. To address this challenge, we introduce the Beyond the Imitation Game benchmark (BIG-bench). BIG-bench currently consists of 204 tasks, contributed by 442 authors across 132 institutions. Task topics are diverse, drawing problems from linguistics, childhood development, math, common-sense reasoning, biology, physics, social bias, software development, and beyond. BIG-bench focuses on tasks that are believed to be beyond the capabilities of current language models. We evaluate the behavior of OpenAI's GPT models, Google-internal dense transformer architectures, and Switch-style sparse transformers on BIG-bench, across model sizes spanning millions to hundreds of billions of parameters. In addition, a team of human expert raters performed all tasks in order to provide a strong baseline. Findings include: model performance and calibration both improve with scale, but are poor in absolute terms (and when compared with rater performance); performance is remarkably similar across model classes, though with benefits from sparsity; tasks that improve gradually and predictably commonly involve a large knowledge or memorization component, whereas tasks that exhibit "breakthrough" behavior at a critical scale often involve multiple steps or components, or brittle metrics; social bias typically increases with scale in settings with ambiguous context, but this can be improved with prompting.
Play to Grade: Testing Coding Games as Classifying Markov Decision Process
Oct 27, 2021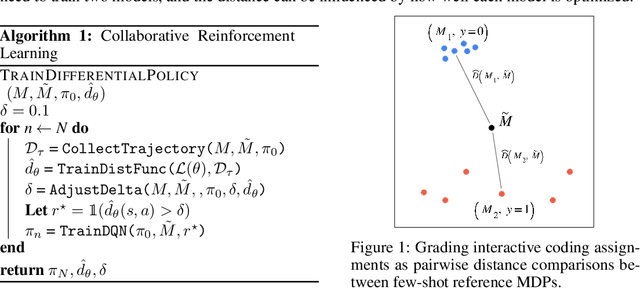
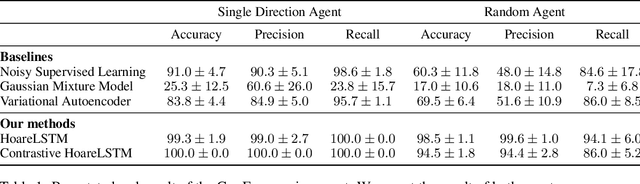
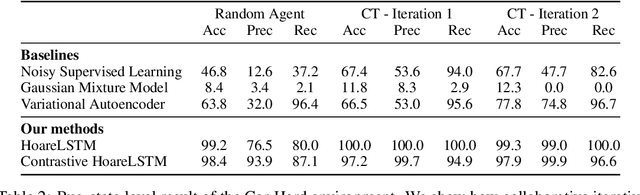
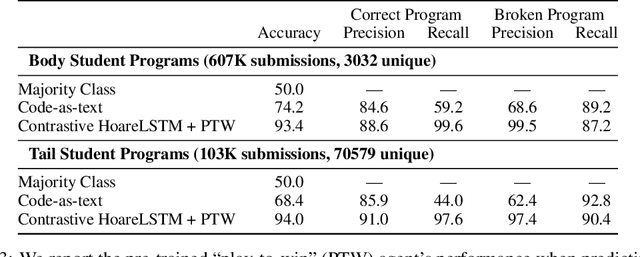
Abstract:Contemporary coding education often presents students with the task of developing programs that have user interaction and complex dynamic systems, such as mouse based games. While pedagogically compelling, there are no contemporary autonomous methods for providing feedback. Notably, interactive programs are impossible to grade by traditional unit tests. In this paper we formalize the challenge of providing feedback to interactive programs as a task of classifying Markov Decision Processes (MDPs). Each student's program fully specifies an MDP where the agent needs to operate and decide, under reasonable generalization, if the dynamics and reward model of the input MDP should be categorized as correct or broken. We demonstrate that by designing a cooperative objective between an agent and an autoregressive model, we can use the agent to sample differential trajectories from the input MDP that allows a classifier to determine membership: Play to Grade. Our method enables an automatic feedback system for interactive code assignments. We release a dataset of 711,274 anonymized student submissions to a single assignment with hand-coded bug labels to support future research.
On the Opportunities and Risks of Foundation Models
Aug 18, 2021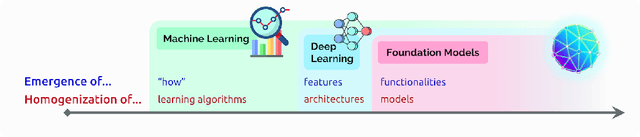
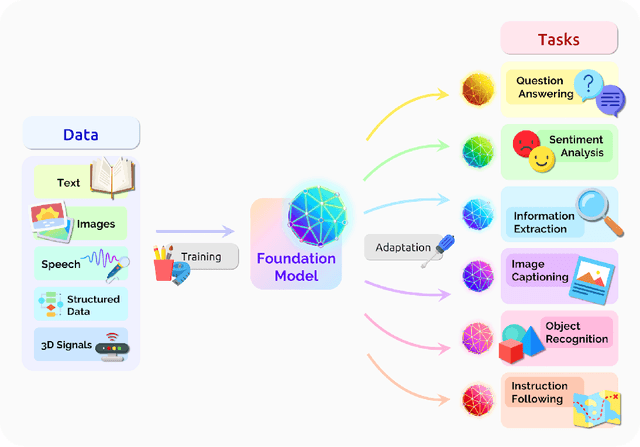
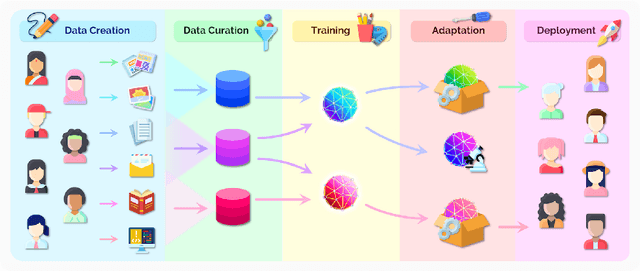
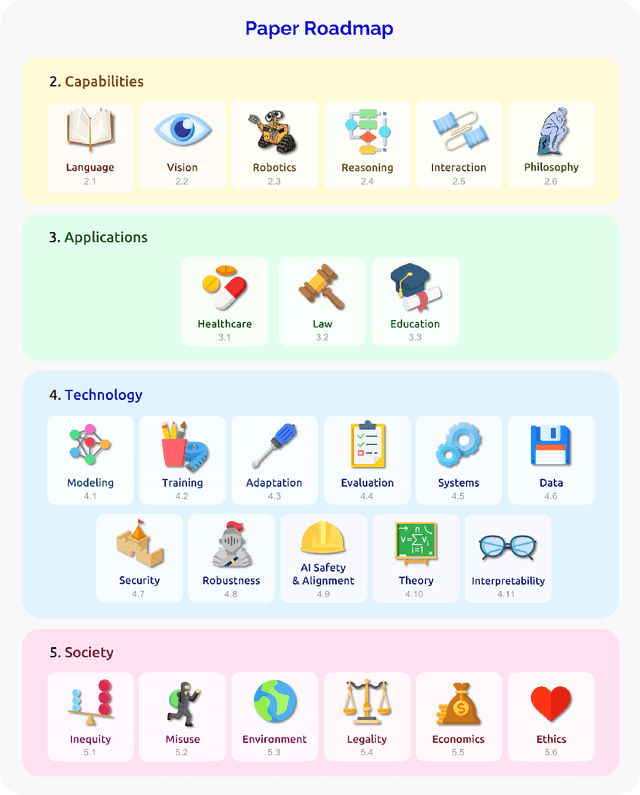
Abstract:AI is undergoing a paradigm shift with the rise of models (e.g., BERT, DALL-E, GPT-3) that are trained on broad data at scale and are adaptable to a wide range of downstream tasks. We call these models foundation models to underscore their critically central yet incomplete character. This report provides a thorough account of the opportunities and risks of foundation models, ranging from their capabilities (e.g., language, vision, robotics, reasoning, human interaction) and technical principles(e.g., model architectures, training procedures, data, systems, security, evaluation, theory) to their applications (e.g., law, healthcare, education) and societal impact (e.g., inequity, misuse, economic and environmental impact, legal and ethical considerations). Though foundation models are based on standard deep learning and transfer learning, their scale results in new emergent capabilities,and their effectiveness across so many tasks incentivizes homogenization. Homogenization provides powerful leverage but demands caution, as the defects of the foundation model are inherited by all the adapted models downstream. Despite the impending widespread deployment of foundation models, we currently lack a clear understanding of how they work, when they fail, and what they are even capable of due to their emergent properties. To tackle these questions, we believe much of the critical research on foundation models will require deep interdisciplinary collaboration commensurate with their fundamentally sociotechnical nature.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge