"Image": models, code, and papers
Photorealistic Image Reconstruction from Hybrid Intensity and Event based Sensor
Oct 25, 2018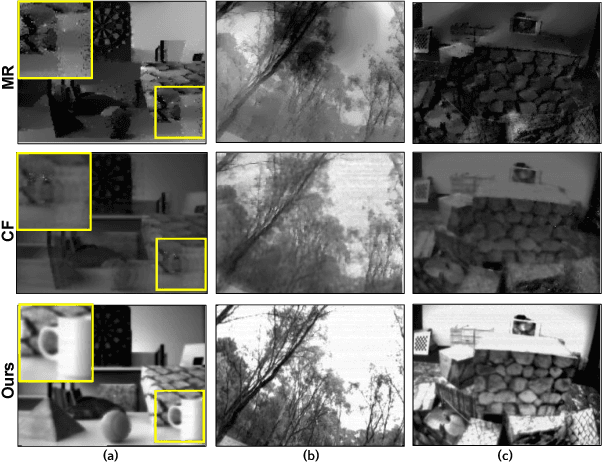
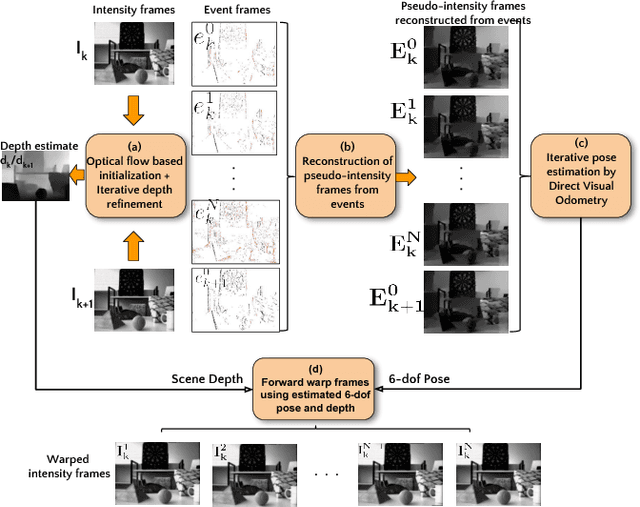
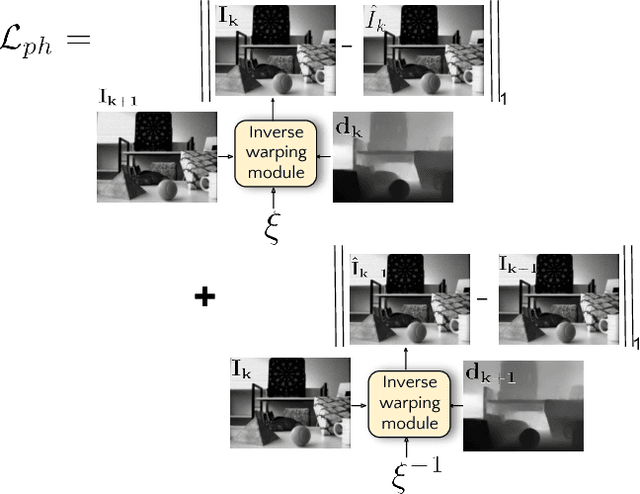
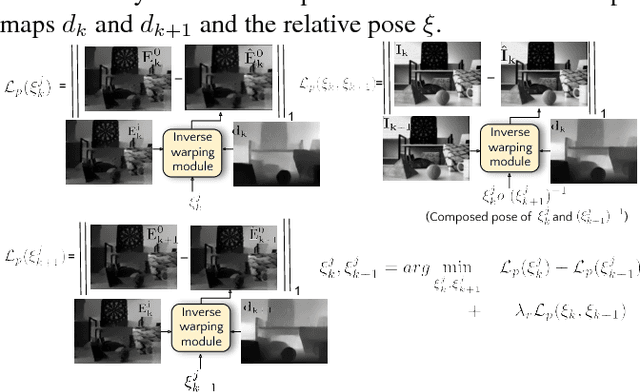
Event sensors output a stream of asynchronous brightness changes (called "events") at a very high temporal rate. Since, these "events" cannot be used directly for traditional computer vision algorithms, many researchers attempted at recovering the intensity information from this stream of events. Although the results are promising, they lack the texture and the consistency of natural videos. We propose to reconstruct photorealistic intensity images from a hybrid sensor consisting of a low frame rate conventional camera along with the event sensor. DAVIS is a commercially available hybrid sensor which bundles the conventional image sensor and the event sensor into a single hardware. To accomplish our task, we use the low frame rate intensity images and warp them to the temporally dense locations of the event data by estimating a spatially dense scene depth and temporally dense sensor ego-motion. We thereby obtain temporally dense photorealisitic images which would have been very difficult by using only either the event sensor or the conventional low frame-rate image sensor. In addition, we also obtain spatio-temporally dense scene flow as a by product of estimating spatially dense depth map and temporally dense sensor ego-motion.
Flexible Bayesian Modelling for Nonlinear Image Registration
Jun 03, 2020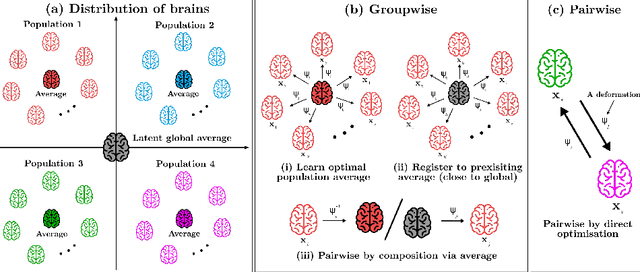
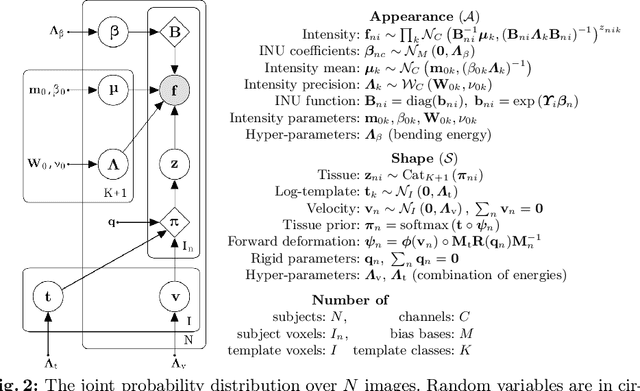
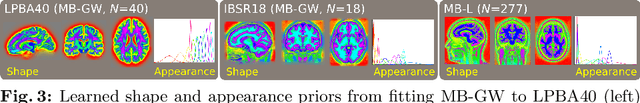
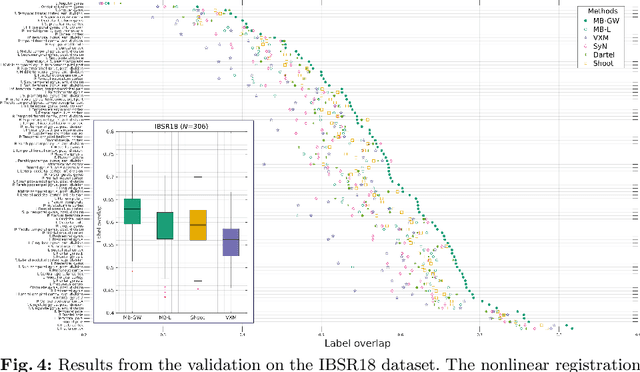
We describe a diffeomorphic registration algorithm that allows groups of images to be accurately aligned to a common space, which we intend to incorporate into the SPM software. The idea is to perform inference in a probabilistic graphical model that accounts for variability in both shape and appearance. The resulting framework is general and entirely unsupervised. The model is evaluated at inter-subject registration of 3D human brain scans. Here, the main modeling assumption is that individual anatomies can be generated by deforming a latent 'average' brain. The method is agnostic to imaging modality and can be applied with no prior processing. We evaluate the algorithm using freely available, manually labelled datasets. In this validation we achieve state-of-the-art results, within reasonable runtimes, against previous state-of-the-art widely used, inter-subject registration algorithms. On the unprocessed dataset, the increase in overlap score is over 17%. These results demonstrate the benefits of using informative computational anatomy frameworks for nonlinear registration.
Bayesian Neural Network Priors Revisited
Feb 12, 2021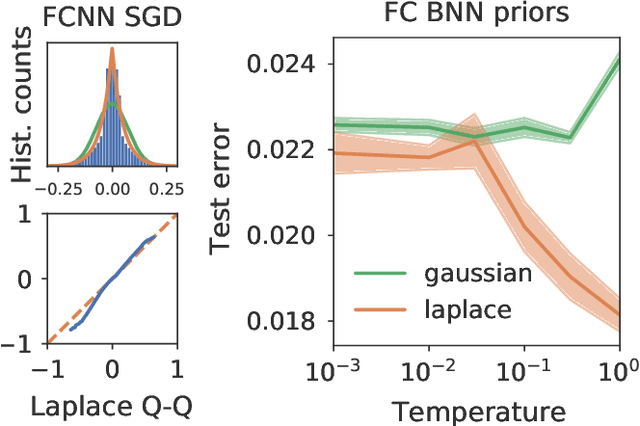
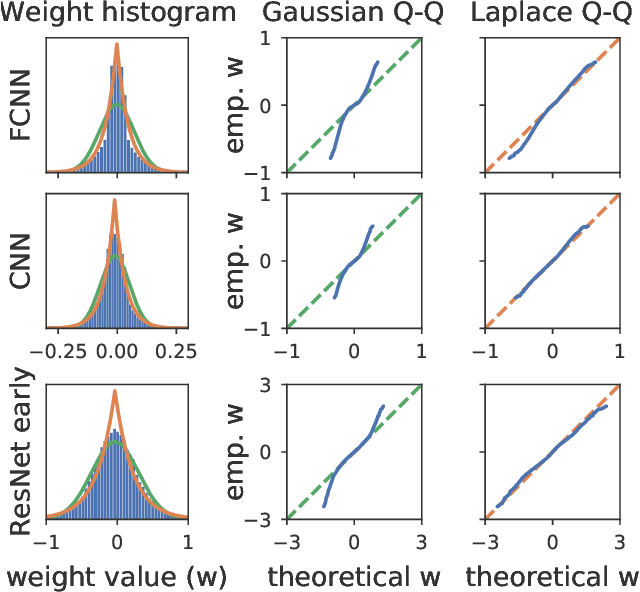
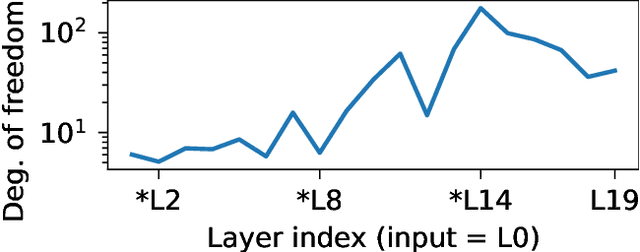
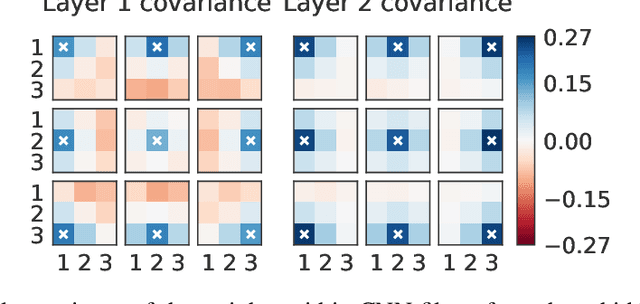
Isotropic Gaussian priors are the de facto standard for modern Bayesian neural network inference. However, such simplistic priors are unlikely to either accurately reflect our true beliefs about the weight distributions, or to give optimal performance. We study summary statistics of neural network weights in different networks trained using SGD. We find that fully connected networks (FCNNs) display heavy-tailed weight distributions, while convolutional neural network (CNN) weights display strong spatial correlations. Building these observations into the respective priors leads to improved performance on a variety of image classification datasets. Moreover, we find that these priors also mitigate the cold posterior effect in FCNNs, while in CNNs we see strong improvements at all temperatures, and hence no reduction in the cold posterior effect.
Covapixels
Oct 18, 2020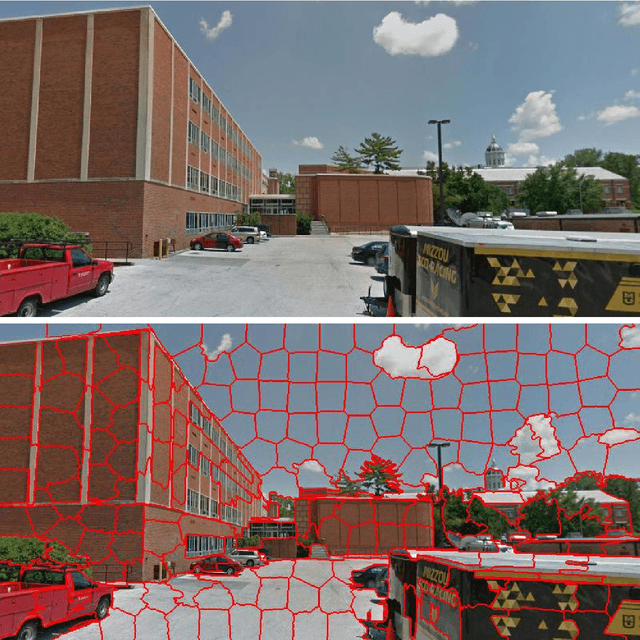
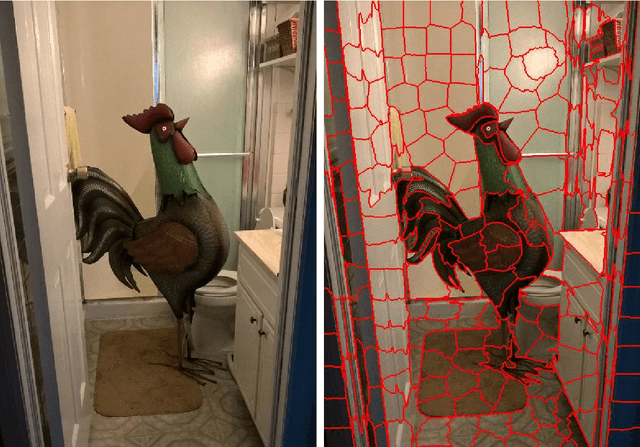
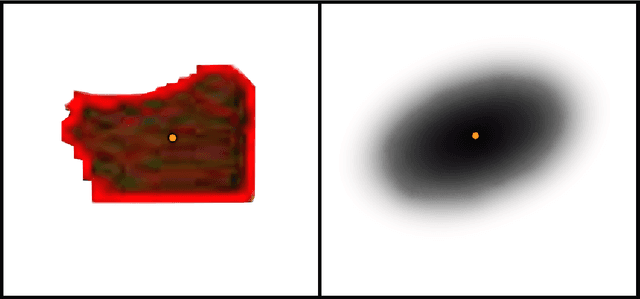


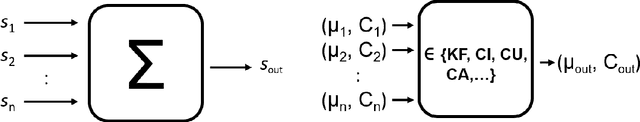
We propose and discuss the summarization of superpixel-type image tiles/patches using mean and covariance information. We refer to the resulting objects as covapixels.
Binary segmentation of medical images using implicit spline representations and deep learning
Feb 25, 2021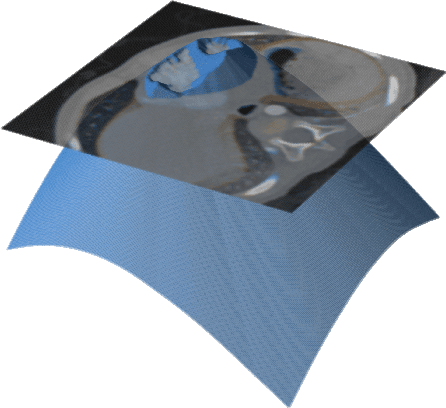
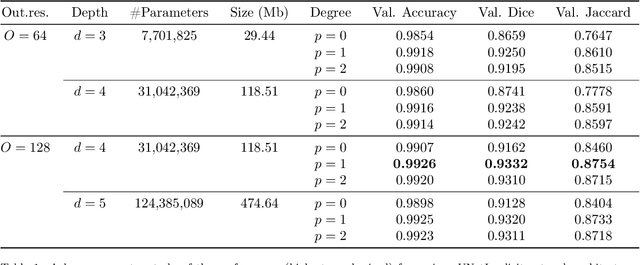
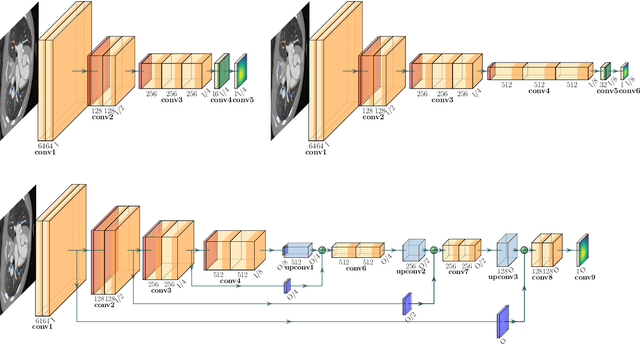
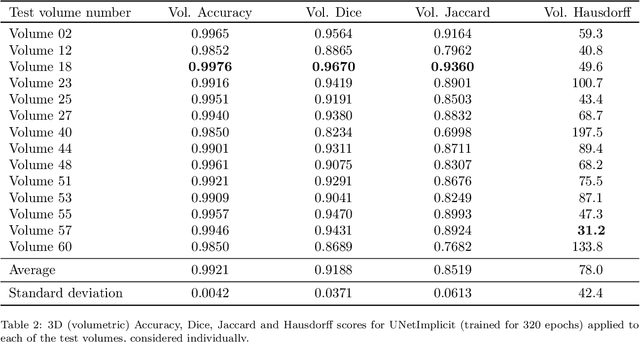
We propose a novel approach to image segmentation based on combining implicit spline representations with deep convolutional neural networks. This is done by predicting the control points of a bivariate spline function whose zero-set represents the segmentation boundary. We adapt several existing neural network architectures and design novel loss functions that are tailored towards providing implicit spline curve approximations. The method is evaluated on a congenital heart disease computed tomography medical imaging dataset. Experiments are carried out by measuring performance in various standard metrics for different networks and loss functions. We determine that splines of bidegree $(1,1)$ with $128\times128$ coefficient resolution performed optimally for $512\times 512$ resolution CT images. For our best network, we achieve an average volumetric test Dice score of almost 92%, which reaches the state of the art for this congenital heart disease dataset.
Hierarchical Amortized Training for Memory-efficient High Resolution 3D GAN
Aug 05, 2020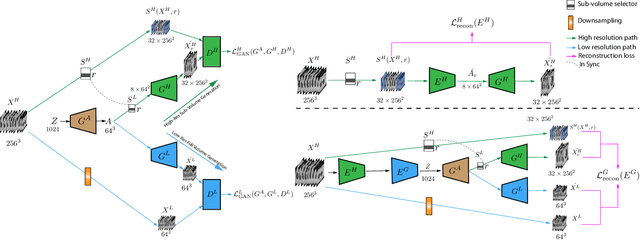
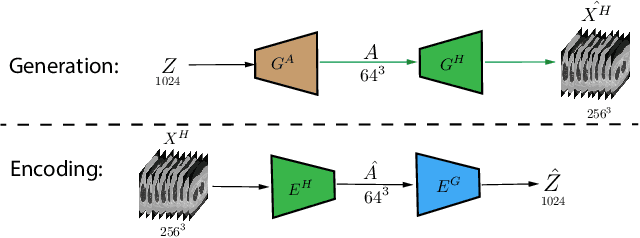
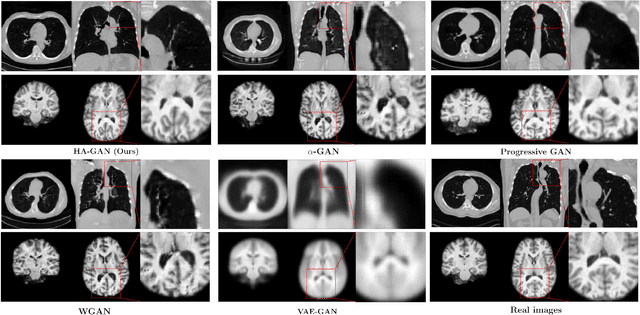
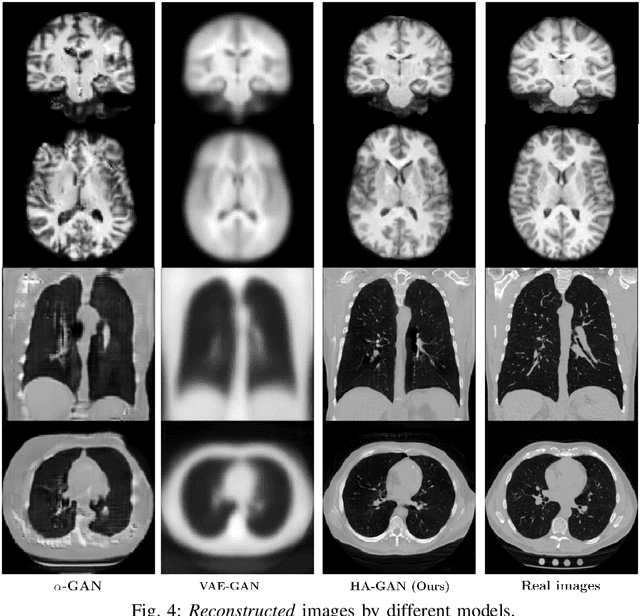
Generative Adversarial Networks (GAN) have many potential medical imaging applications, including data augmentation, domain adaptation, and model explanation. Due to the limited embedded memory of Graphical Processing Units (GPUs), most current 3D GAN models are trained on low-resolution medical images. In this work, we propose a novel end-to-end GAN architecture that can generate high-resolution 3D images. We achieve this goal by separating training and inference. During training, we adopt a hierarchical structure that simultaneously generates a low-resolution version of the image and a randomly selected sub-volume of the high-resolution image. The hierarchical design has two advantages: First, the memory demand for training on high-resolution images is amortized among subvolumes. Furthermore, anchoring the high-resolution subvolumes to a single low-resolution image ensures anatomical consistency between subvolumes. During inference, our model can directly generate full high-resolution images. We also incorporate an encoder with a similar hierarchical structure into the model to extract features from the images. Experiments on 3D thorax CT and brain MRI demonstrate that our approach outperforms state of the art in image generation, image reconstruction, and clinical-relevant variables prediction.
Compressive lensless endoscopy with partial speckle scanning
Apr 22, 2021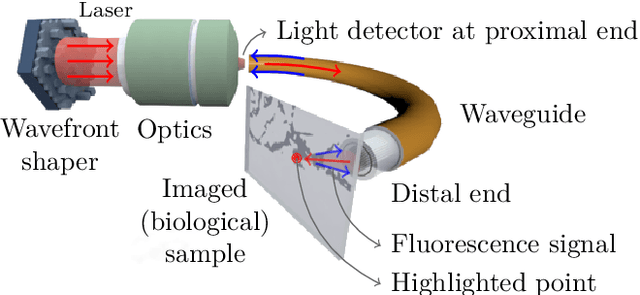
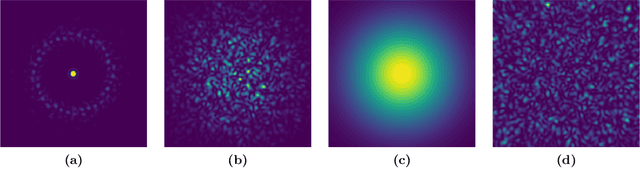
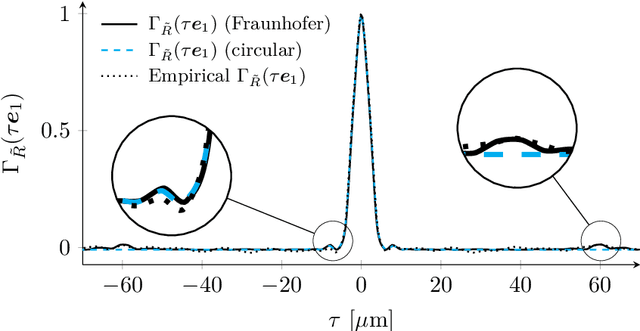
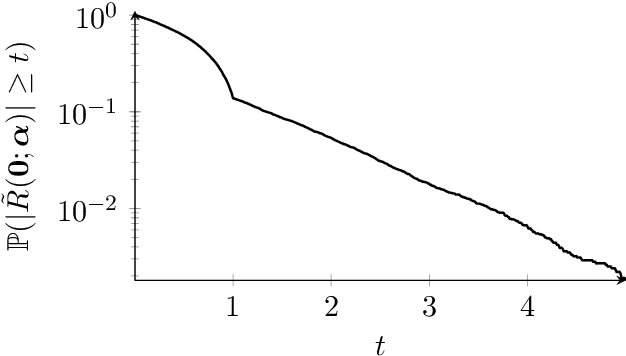
The lensless endoscope (LE) is a promising device to acquire in vivo images at a cellular scale. The tiny size of the probe enables a deep exploration of the tissues. Lensless endoscopy with a multicore fiber (MCF) commonly uses a spatial light modulator (SLM) to coherently combine, at the output of the MCF, few hundreds of beamlets into a focus spot. This spot is subsequently scanned across the sample to generate a fluorescent image. We propose here a novel scanning scheme, partial speckle scanning (PSS), inspired by compressive sensing theory, that avoids the use of an SLM to perform fluorescent imaging in LE with reduced acquisition time. Such a strategy avoids photo-bleaching while keeping high reconstruction quality. We develop our approach on two key properties of the LE: (i) the ability to easily generate speckles, and (ii) the memory effect in MCF that allows to use fast scan mirrors to shift light patterns. First, we show that speckles are sub-exponential random fields. Despite their granular structure, an appropriate choice of the reconstruction parameters makes them good candidates to build efficient sensing matrices. Then, we numerically validate our approach and apply it on experimental data. The proposed sensing technique outperforms conventional raster scanning: higher reconstruction quality is achieved with far fewer observations. For a fixed reconstruction quality, our speckle scanning approach is faster than compressive sensing schemes which require to change the speckle pattern for each observation.
Patch2Pix: Epipolar-Guided Pixel-Level Correspondences
Dec 04, 2020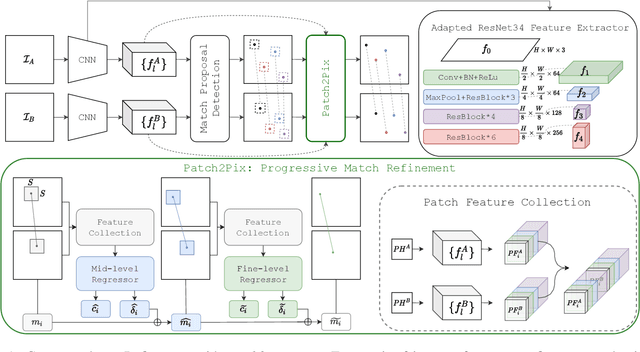
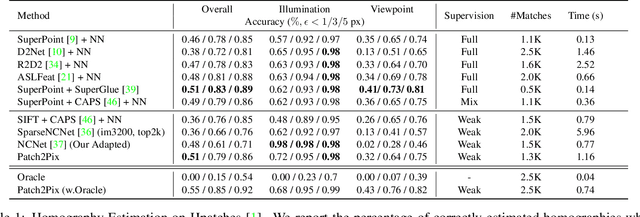
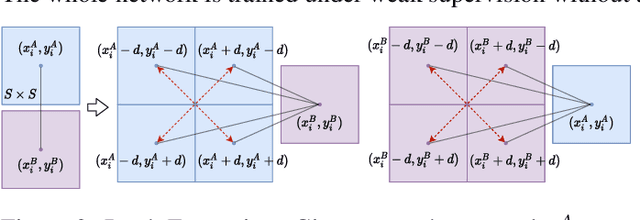
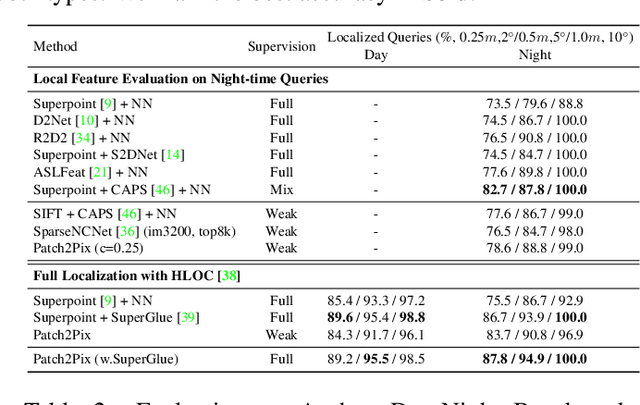
Deep learning has been applied to a classical matching pipeline which typically involves three steps: (i) local feature detection and description, (ii) feature matching, and (iii) outlier rejection. Recently emerged correspondence networks propose to perform those steps inside a single network but suffer from low matching resolution due to the memory bottleneck. In this work, we propose a new perspective to estimate correspondences in a detect-to-refine manner, where we first predict patch-level match proposals and then refine them. We present a novel refinement network Patch2Pix that refines match proposals by regressing pixel-level matches from the local regions defined by those proposals and jointly rejecting outlier matches with confidence scores, which is weakly supervised to learn correspondences that are consistent with the epipolar geometry of an input image pair. We show that our refinement network significantly improves the performance of correspondence networks on image matching, homography estimation, and localization tasks. In addition, we show that our learned refinement generalizes to fully-supervised methods without re-training, which leads us to state-of-the-art localization performance.
Towards a Rigorous Evaluation of Explainability for Multivariate Time Series
Apr 06, 2021
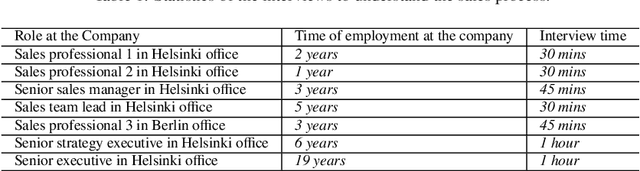
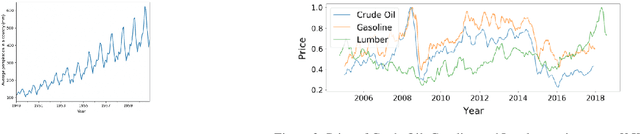
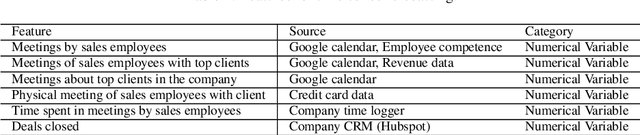
Machine learning-based systems are rapidly gaining popularity and in-line with that there has been a huge research surge in the field of explainability to ensure that machine learning models are reliable, fair, and can be held liable for their decision-making process. Explainable Artificial Intelligence (XAI) methods are typically deployed to debug black-box machine learning models but in comparison to tabular, text, and image data, explainability in time series is still relatively unexplored. The aim of this study was to achieve and evaluate model agnostic explainability in a time series forecasting problem. This work focused on proving a solution for a digital consultancy company aiming to find a data-driven approach in order to understand the effect of their sales related activities on the sales deals closed. The solution involved framing the problem as a time series forecasting problem to predict the sales deals and the explainability was achieved using two novel model agnostic explainability techniques, Local explainable model-agnostic explanations (LIME) and Shapley additive explanations (SHAP) which were evaluated using human evaluation of explainability. The results clearly indicate that the explanations produced by LIME and SHAP greatly helped lay humans in understanding the predictions made by the machine learning model. The presented work can easily be extended to any time
Monte-Carlo Sampling applied to Multiple Instance Learning for Histological Image Classification
Dec 30, 2018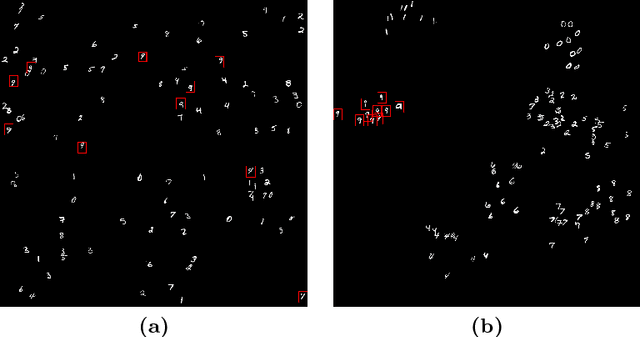
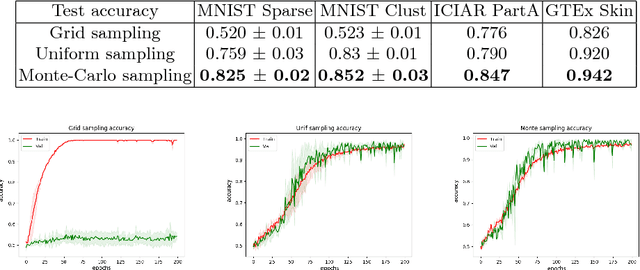
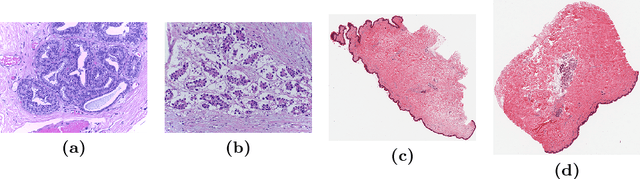

We propose a patch sampling strategy based on a sequential Monte-Carlo method for high resolution image classification in the context of Multiple Instance Learning. When compared with grid sampling and uniform sampling techniques, it achieves higher generalization performance. We validate the strategy on two artificial datasets and two histological datasets for breast cancer and sun exposure classification.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge