"Image": models, code, and papers
Hierarchical ResNeXt Models for Breast Cancer Histology Image Classification
Oct 21, 2018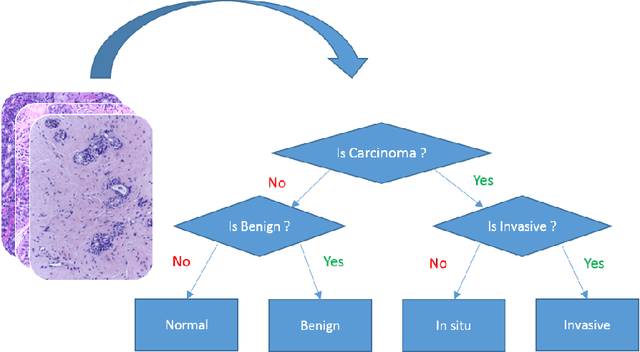
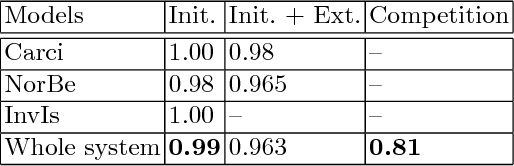
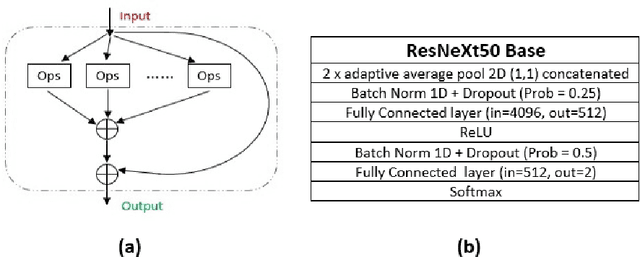
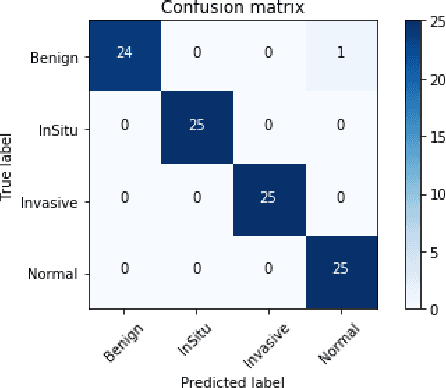
Microscopic histology image analysis is a cornerstone in early detection of breast cancer. However these images are very large and manual analysis is error prone and very time consuming. Thus automating this process is in high demand. We proposed a hierarchical system of convolutional neural networks (CNN) that classifies automatically patches of these images into four pathologies: normal, benign, in situ carcinoma and invasive carcinoma. We evaluated our system on the BACH challenge dataset of image-wise classification and a small dataset that we used to extend it. Using a train/test split of 75%/25%, we achieved an accuracy rate of 0.99 on the test split for the BACH dataset and 0.96 on that of the extension. On the test of the BACH challenge, we've reached an accuracy of 0.81 which rank us to the 8th out of 51 teams.
RankSRGAN: Generative Adversarial Networks with Ranker for Image Super-Resolution
Aug 18, 2019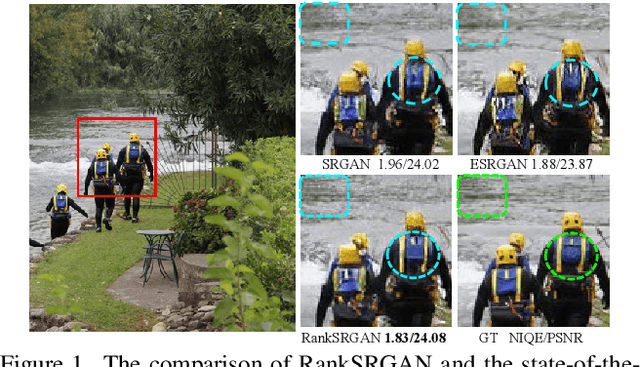
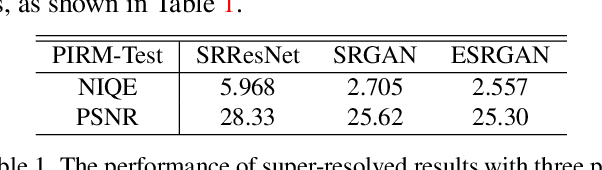
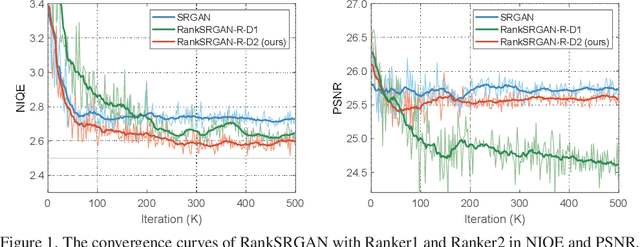
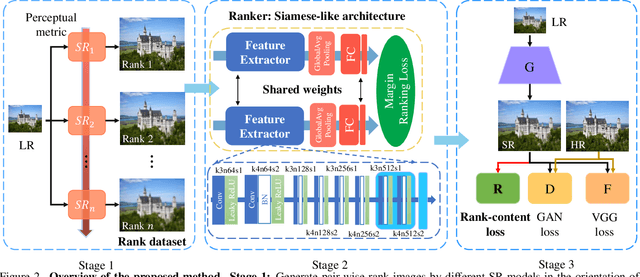
Generative Adversarial Networks (GAN) have demonstrated the potential to recover realistic details for single image super-resolution (SISR). To further improve the visual quality of super-resolved results, PIRM2018-SR Challenge employed perceptual metrics to assess the perceptual quality, such as PI, NIQE, and Ma. However, existing methods cannot directly optimize these indifferentiable perceptual metrics, which are shown to be highly correlated with human ratings. To address the problem, we propose Super-Resolution Generative Adversarial Networks with Ranker (RankSRGAN) to optimize generator in the direction of perceptual metrics. Specifically, we first train a Ranker which can learn the behavior of perceptual metrics and then introduce a novel rank-content loss to optimize the perceptual quality. The most appealing part is that the proposed method can combine the strengths of different SR methods to generate better results. Extensive experiments show that RankSRGAN achieves visually pleasing results and reaches state-of-the-art performance in perceptual metrics. Project page: https://wenlongzhang0724.github.io/Projects/RankSRGAN
Sandwich Batch Normalization
Feb 22, 2021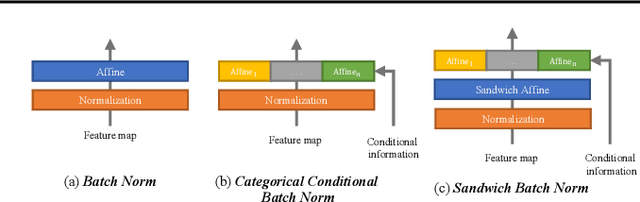
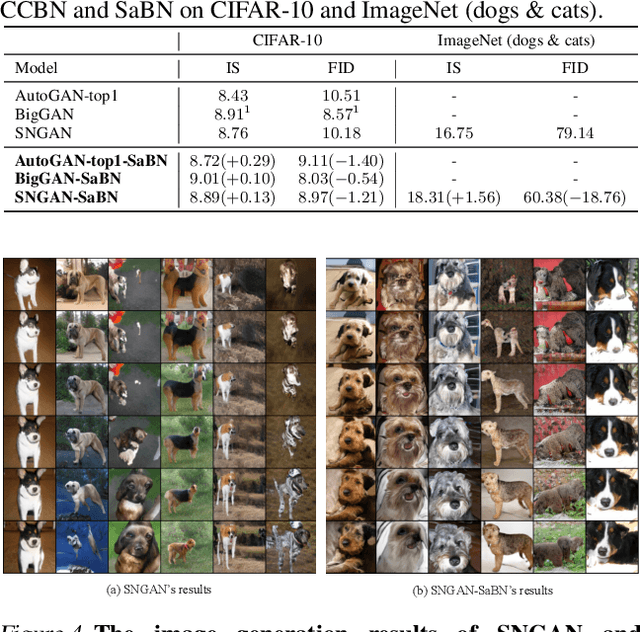
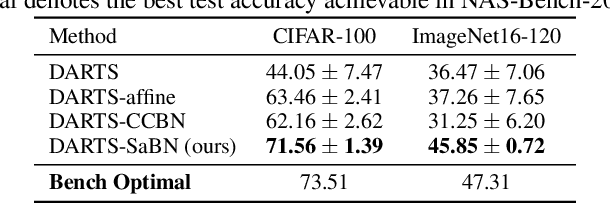

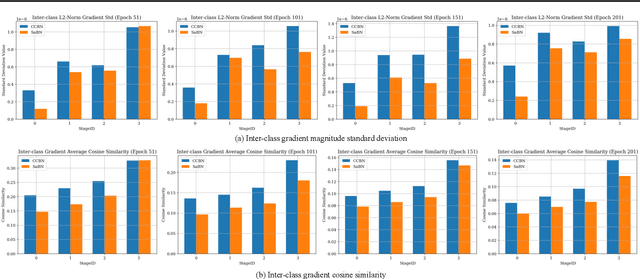

We present Sandwich Batch Normalization (SaBN), an embarrassingly easy improvement of Batch Normalization (BN) with only a few lines of code changes. SaBN is motivated by addressing the inherent feature distribution heterogeneity that one can be identified in many tasks, which can arise from data heterogeneity (multiple input domains) or model heterogeneity (dynamic architectures, model conditioning, etc.). Our SaBN factorizes the BN affine layer into one shared sandwich affine layer, cascaded by several parallel independent affine layers. Concrete analysis reveals that, during optimization, SaBN promotes balanced gradient norms while still preserving diverse gradient directions: a property that many application tasks seem to favor. We demonstrate the prevailing effectiveness of SaBN as a drop-in replacement in four tasks: $\textbf{conditional image generation}$, $\textbf{neural architecture search}$ (NAS), $\textbf{adversarial training}$, and $\textbf{arbitrary style transfer}$. Leveraging SaBN immediately achieves better Inception Score and FID on CIFAR-10 and ImageNet conditional image generation with three state-of-the-art GANs; boosts the performance of a state-of-the-art weight-sharing NAS algorithm significantly on NAS-Bench-201; substantially improves the robust and standard accuracies for adversarial defense; and produces superior arbitrary stylized results. We also provide visualizations and analysis to help understand why SaBN works. Codes are available at https://github.com/VITA-Group/Sandwich-Batch-Normalization.
Wide & Deep neural network model for patch aggregation in CNN-based prostate cancer detection systems
May 20, 2021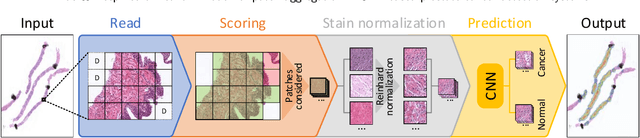
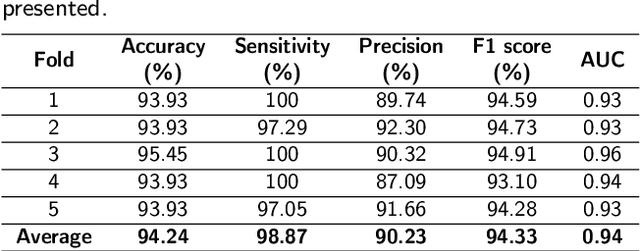
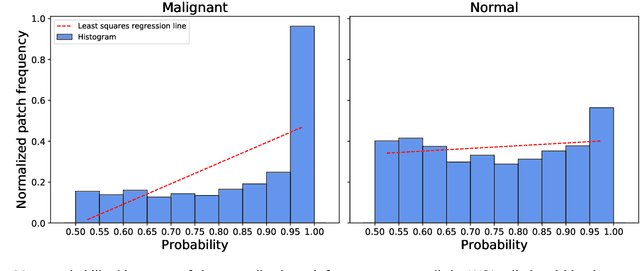
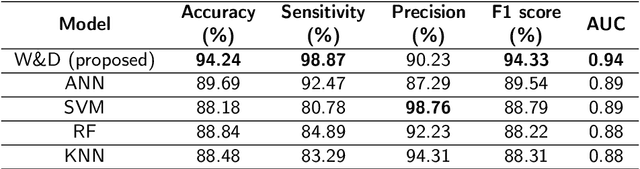
Prostate cancer (PCa) is one of the most commonly diagnosed cancer and one of the leading causes of death among men, with almost 1.41 million new cases and around 375,000 deaths in 2020. Artificial Intelligence algorithms have had a huge impact in medical image analysis, including digital histopathology, where Convolutional Neural Networks (CNNs) are used to provide a fast and accurate diagnosis, supporting experts in this task. To perform an automatic diagnosis, prostate tissue samples are first digitized into gigapixel-resolution whole-slide images. Due to the size of these images, neural networks cannot use them as input and, therefore, small subimages called patches are extracted and predicted, obtaining a patch-level classification. In this work, a novel patch aggregation method based on a custom Wide & Deep neural network model is presented, which performs a slide-level classification using the patch-level classes obtained from a CNN. The malignant tissue ratio, a 10-bin malignant probability histogram, the least squares regression line of the histogram, and the number of malignant connected components are used by the proposed model to perform the classification. An accuracy of 94.24% and a sensitivity of 98.87% were achieved, proving that the proposed system could aid pathologists by speeding up the screening process and, thus, contribute to the fight against PCa.
FlipReID: Closing the Gap between Training and Inference in Person Re-Identification
May 12, 2021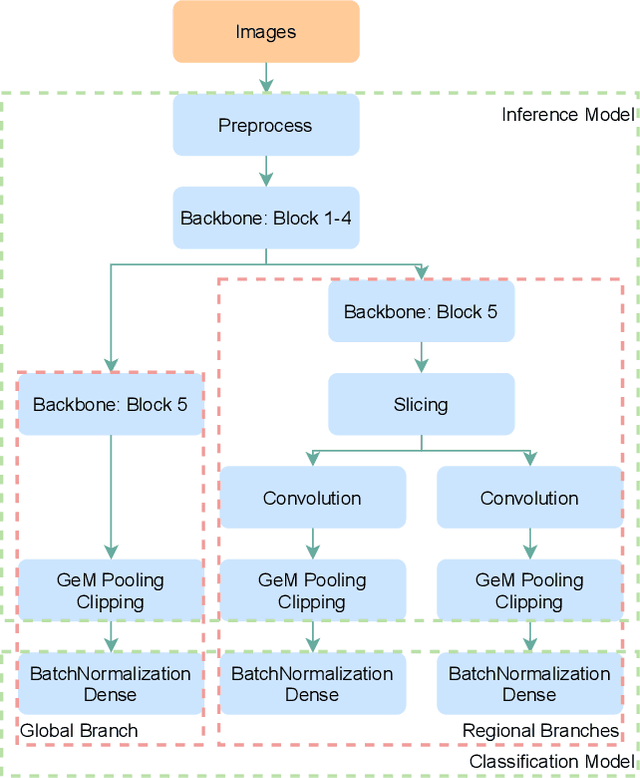
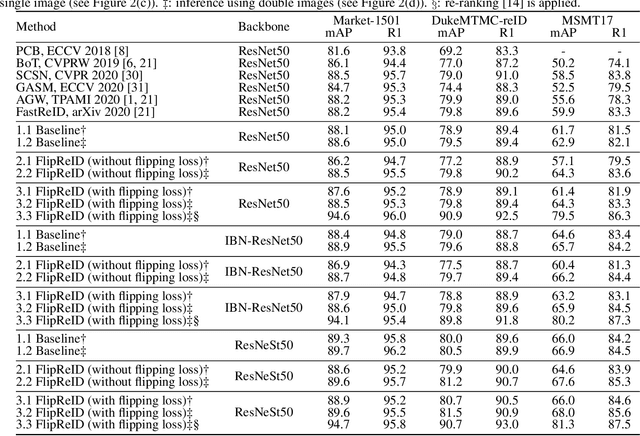
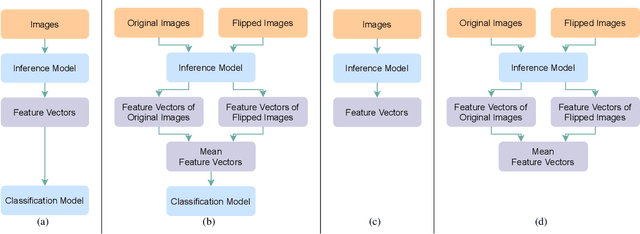
Since neural networks are data-hungry, incorporating data augmentation in training is a widely adopted technique that enlarges datasets and improves generalization. On the other hand, aggregating predictions of multiple augmented samples (i.e., test-time augmentation) could boost performance even further. In the context of person re-identification models, it is common practice to extract embeddings for both the original images and their horizontally flipped variants. The final representation is the mean of the aforementioned feature vectors. However, such scheme results in a gap between training and inference, i.e., the mean feature vectors calculated in inference are not part of the training pipeline. In this study, we devise the FlipReID structure with the flipping loss to address this issue. More specifically, models using the FlipReID structure are trained on the original images and the flipped images simultaneously, and incorporating the flipping loss minimizes the mean squared error between feature vectors of corresponding image pairs. Extensive experiments show that our method brings consistent improvements. In particular, we set a new record for MSMT17 which is the largest person re-identification dataset. The source code is available at https://github.com/nixingyang/FlipReID.
Robust Brain Magnetic Resonance Image Segmentation for Hydrocephalus Patients: Hard and Soft Attention
Jan 12, 2020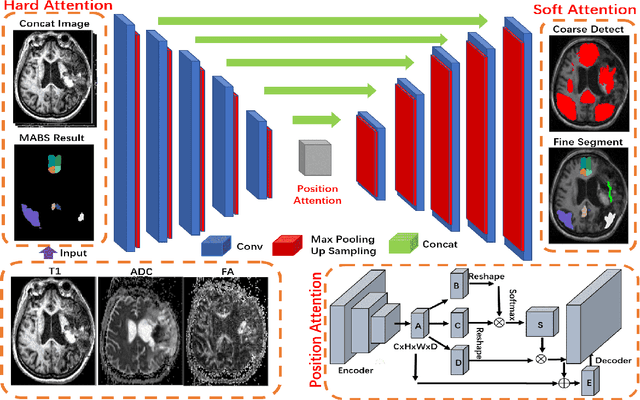
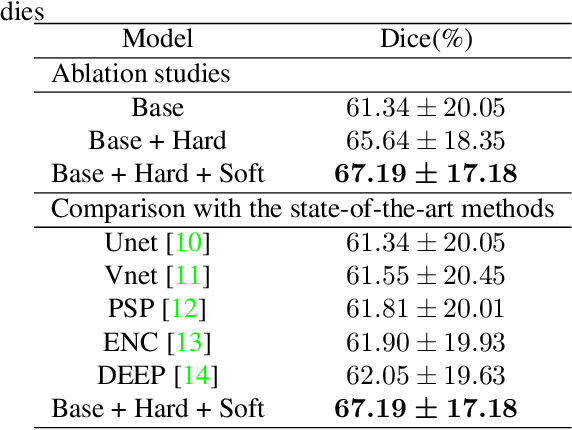
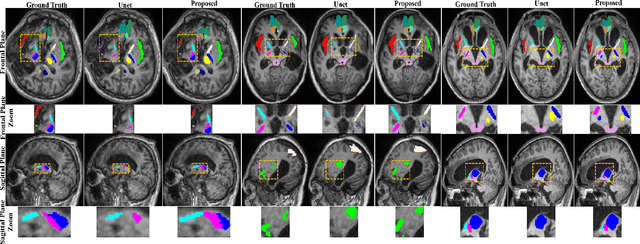
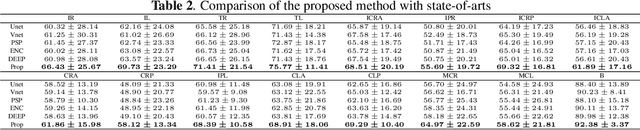
Brain magnetic resonance (MR) segmentation for hydrocephalus patients is considered as a challenging work. Encoding the variation of the brain anatomical structures from different individuals cannot be easily achieved. The task becomes even more difficult especially when the image data from hydrocephalus patients are considered, which often have large deformations and differ significantly from the normal subjects. Here, we propose a novel strategy with hard and soft attention modules to solve the segmentation problems for hydrocephalus MR images. Our main contributions are three-fold: 1) the hard-attention module generates coarse segmentation map using multi-atlas-based method and the VoxelMorph tool, which guides subsequent segmentation process and improves its robustness; 2) the soft-attention module incorporates position attention to capture precise context information, which further improves the segmentation accuracy; 3) we validate our method by segmenting insula, thalamus and many other regions-of-interests (ROIs) that are critical to quantify brain MR images of hydrocephalus patients in real clinical scenario. The proposed method achieves much improved robustness and accuracy when segmenting all 17 consciousness-related ROIs with high variations for different subjects. To the best of our knowledge, this is the first work to employ deep learning for solving the brain segmentation problems of hydrocephalus patients.
Rank-One Prior: Toward Real-Time Scene Recovery
Apr 07, 2021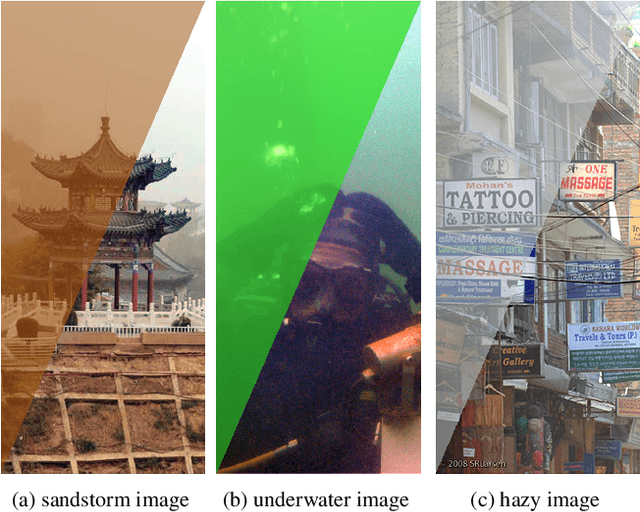
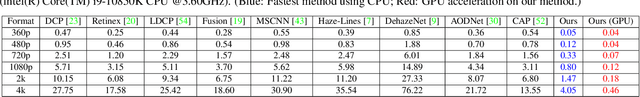
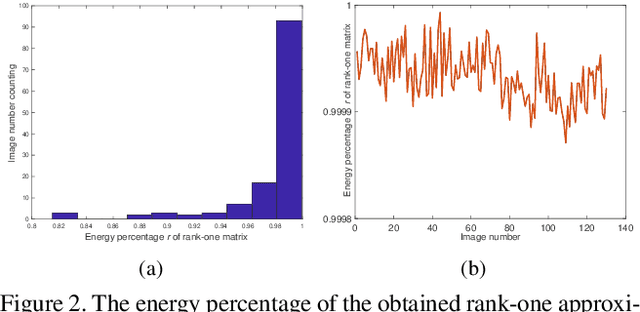
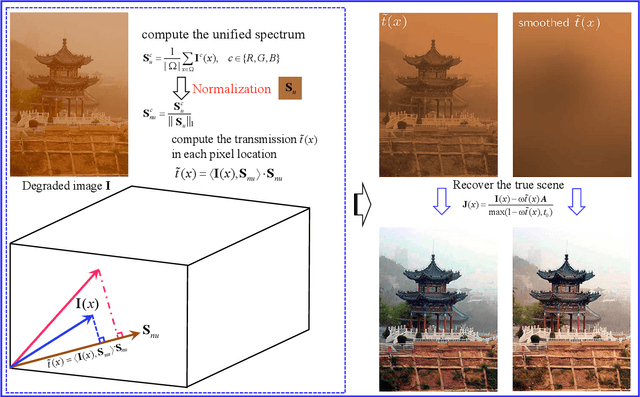
Scene recovery is a fundamental imaging task for several practical applications, e.g., video surveillance and autonomous vehicles, etc. To improve visual quality under different weather/imaging conditions, we propose a real-time light correction method to recover the degraded scenes in the cases of sandstorms, underwater, and haze. The heart of our work is that we propose an intensity projection strategy to estimate the transmission. This strategy is motivated by a straightforward rank-one transmission prior. The complexity of transmission estimation is $O(N)$ where $N$ is the size of the single image. Then we can recover the scene in real-time. Comprehensive experiments on different types of weather/imaging conditions illustrate that our method outperforms competitively several state-of-the-art imaging methods in terms of efficiency and robustness.
Learning Converged Propagations with Deep Prior Ensemble for Image Enhancement
Oct 09, 2018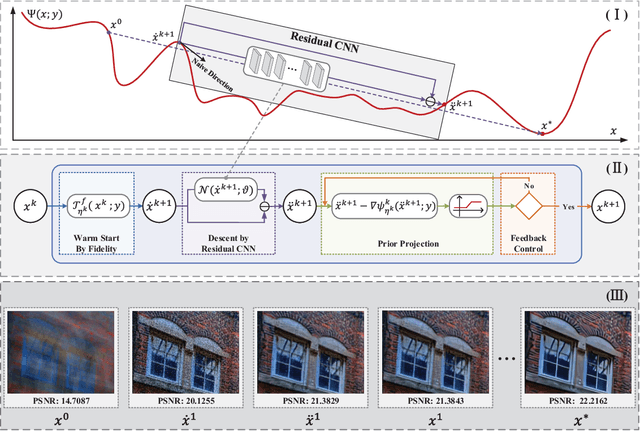



Enhancing visual qualities of images plays very important roles in various vision and learning applications. In the past few years, both knowledge-driven maximum a posterior (MAP) with prior modelings and fully data-dependent convolutional neural network (CNN) techniques have been investigated to address specific enhancement tasks. In this paper, by exploiting the advantages of these two types of mechanisms within a complementary propagation perspective, we propose a unified framework, named deep prior ensemble (DPE), for solving various image enhancement tasks. Specifically, we first establish the basic propagation scheme based on the fundamental image modeling cues and then introduce residual CNNs to help predicting the propagation direction at each stage. By designing prior projections to perform feedback control, we theoretically prove that even with experience-inspired CNNs, DPE is definitely converged and the output will always satisfy our fundamental task constraints. The main advantage against conventional optimization-based MAP approaches is that our descent directions are learned from collected training data, thus are much more robust to unwanted local minimums. While, compared with existing CNN type networks, which are often designed in heuristic manners without theoretical guarantees, DPE is able to gain advantages from rich task cues investigated on the bases of domain knowledges. Therefore, DPE actually provides a generic ensemble methodology to integrate both knowledge and data-based cues for different image enhancement tasks. More importantly, our theoretical investigations verify that the feedforward propagations of DPE are properly controlled toward our desired solution. Experimental results demonstrate that the proposed DPE outperforms state-of-the-arts on a variety of image enhancement tasks in terms of both quantitative measure and visual perception quality.
A Deeper Look into Convolutions via Pruning
Feb 04, 2021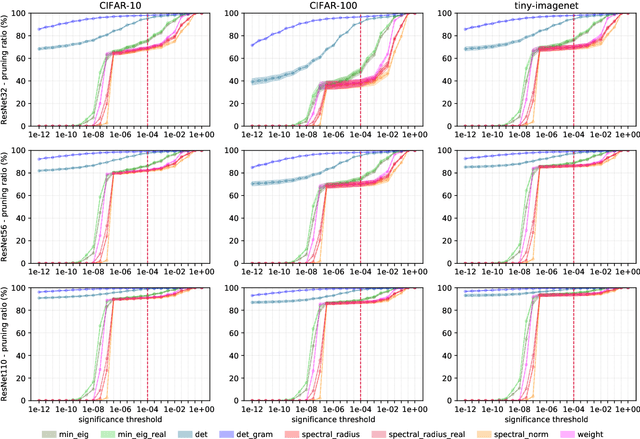
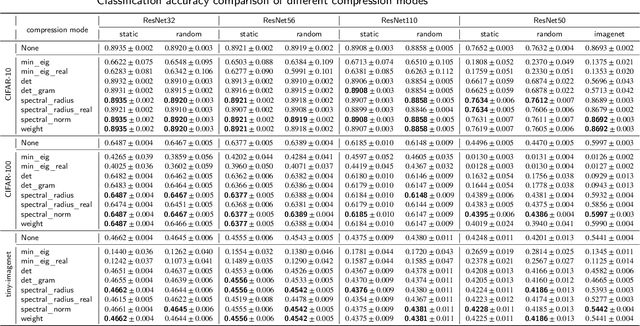
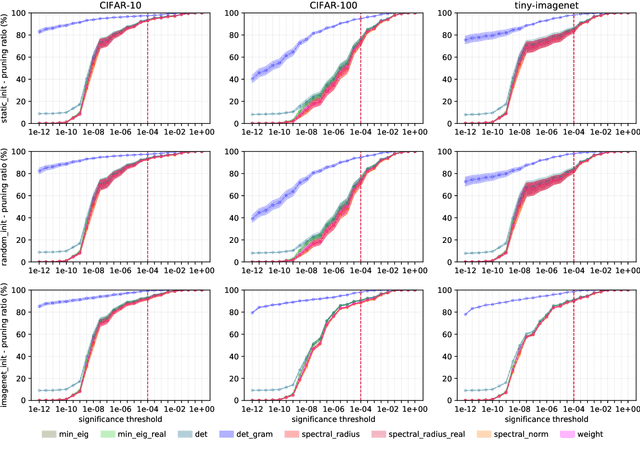
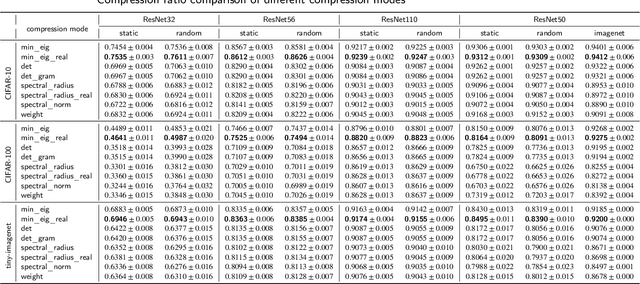
Convolutional neural networks (CNNs) are able to attain better visual recognition performance than fully connected neural networks despite having much less parameters due to their parameter sharing principle. Hence, modern architectures are designed to contain a very small number of fully-connected layers, often at the end, after multiple layers of convolutions. It is interesting to observe that we can replace large fully-connected layers with relatively small groups of tiny matrices applied on the entire image. Moreover, although this strategy already reduces the number of parameters, most of the convolutions can be eliminated as well, without suffering any loss in recognition performance. However, there is no solid recipe to detect this hidden subset of convolutional neurons that is responsible for the majority of the recognition work. Hence, in this work, we use the matrix characteristics based on eigenvalues in addition to the classical weight-based importance assignment approach for pruning to shed light on the internal mechanisms of a widely used family of CNNs, namely residual neural networks (ResNets), for the image classification problem using CIFAR-10, CIFAR-100 and Tiny ImageNet datasets.
Region-specific Dictionary Learning-based Low-dose Thoracic CT Reconstruction
Oct 20, 2020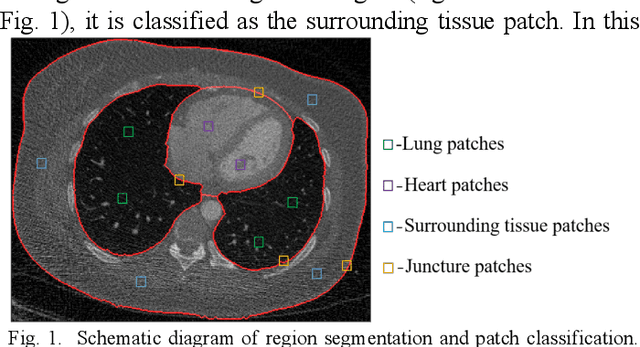
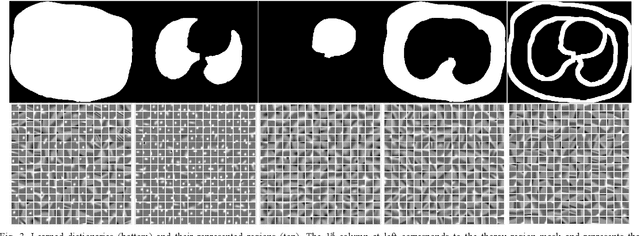
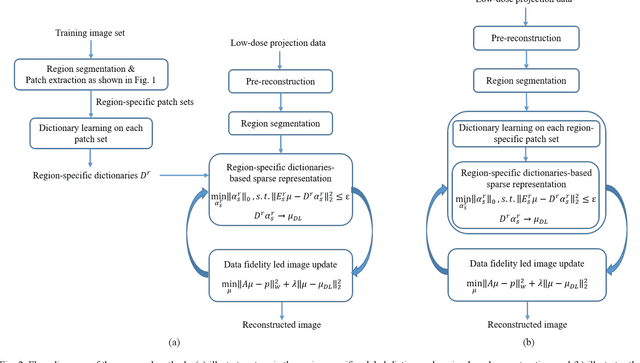

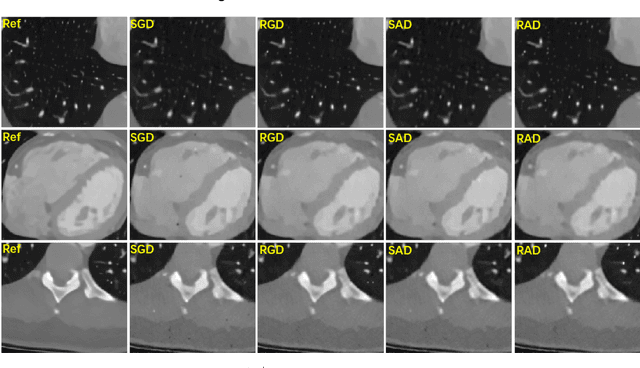
This paper presents a dictionary learning-based method with region-specific image patches to maximize the utility of the powerful sparse data processing technique for CT image reconstruction. Considering heterogeneous distributions of image features and noise in CT, region-specific customization of dictionaries is utilized in iterative reconstruction. Thoracic CT images are partitioned into several regions according to their structural and noise characteristics. Dictionaries specific to each region are then learned from the segmented thoracic CT images and applied to subsequent image reconstruction of the region. Parameters for dictionary learning and sparse representation are determined according to the structural and noise properties of each region. The proposed method results in better performance than the conventional reconstruction based on a single dictionary in recovering structures and suppressing noise in both simulation and human CT imaging. Quantitatively, the simulation study shows maximum improvement of image quality for the whole thorax can achieve 4.88% and 11.1% in terms of the Structure-SIMilarity (SSIM) and Root-Mean-Square Error (RMSE) indices, respectively. For human imaging data, it is found that the structures in the lungs and heart can be better recovered, while simultaneously decreasing noise around the vertebra effectively. The proposed strategy takes into account inherent regional differences inside of the reconstructed object and leads to improved images. The method can be readily extended to CT imaging of other anatomical regions and other applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge