"Image": models, code, and papers
From One to Many: Dynamic Cross Attention Networks for LiDAR and Camera Fusion
Sep 25, 2022
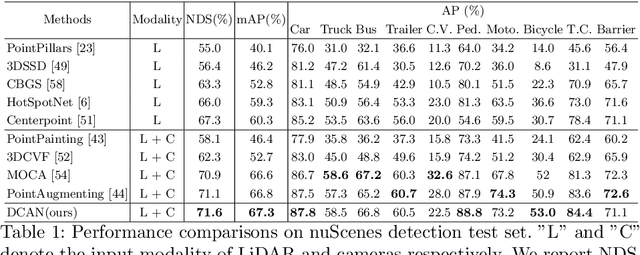
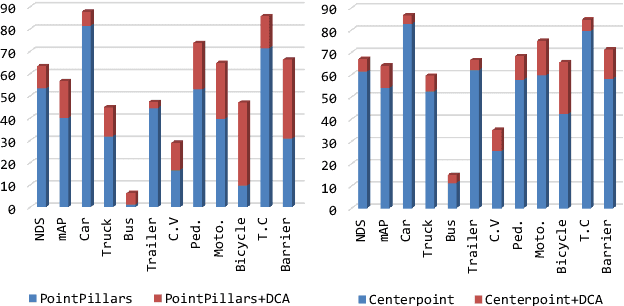
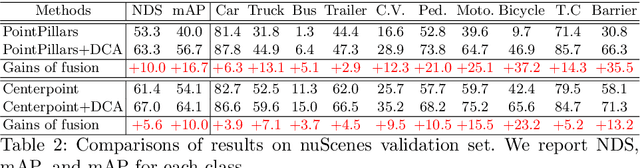
LiDAR and cameras are two complementary sensors for 3D perception in autonomous driving. LiDAR point clouds have accurate spatial and geometry information, while RGB images provide textural and color data for context reasoning. To exploit LiDAR and cameras jointly, existing fusion methods tend to align each 3D point to only one projected image pixel based on calibration, namely one-to-one mapping. However, the performance of these approaches highly relies on the calibration quality, which is sensitive to the temporal and spatial synchronization of sensors. Therefore, we propose a Dynamic Cross Attention (DCA) module with a novel one-to-many cross-modality mapping that learns multiple offsets from the initial projection towards the neighborhood and thus develops tolerance to calibration error. Moreover, a \textit{dynamic query enhancement} is proposed to perceive the model-independent calibration, which further strengthens DCA's tolerance to the initial misalignment. The whole fusion architecture named Dynamic Cross Attention Network (DCAN) exploits multi-level image features and adapts to multiple representations of point clouds, which allows DCA to serve as a plug-in fusion module. Extensive experiments on nuScenes and KITTI prove DCA's effectiveness. The proposed DCAN outperforms state-of-the-art methods on the nuScenes detection challenge.
Leveraging Image Complexity in Macro-Level Neural Network Design for Medical Image Segmentation
Dec 21, 2021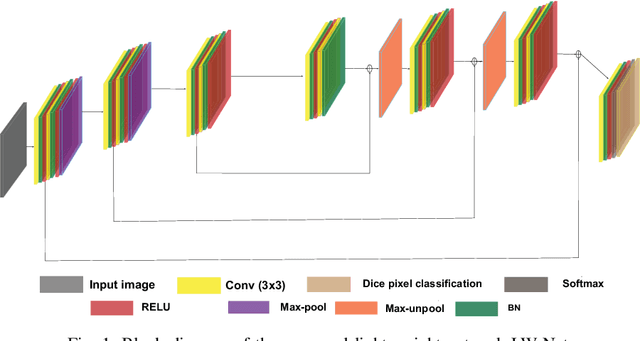
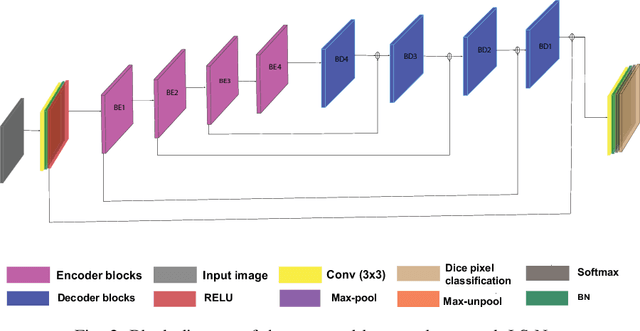
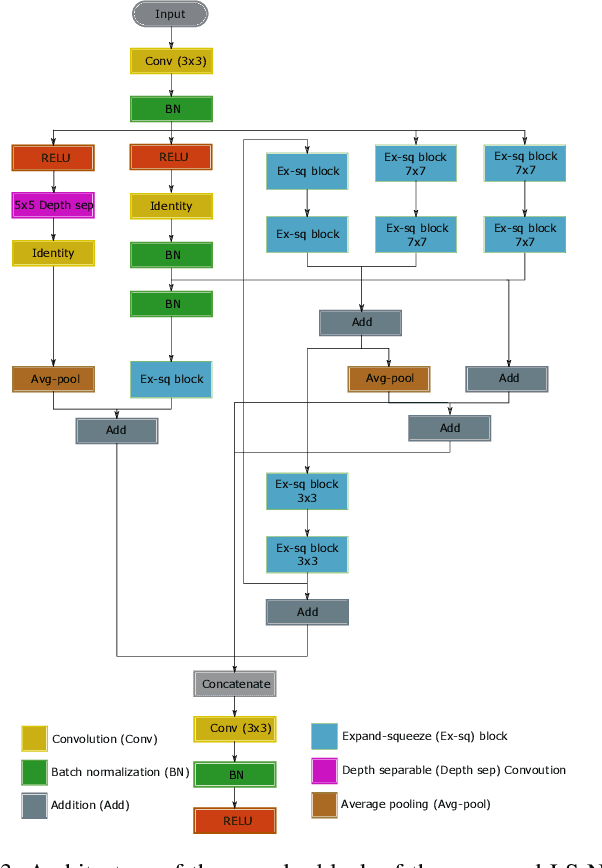
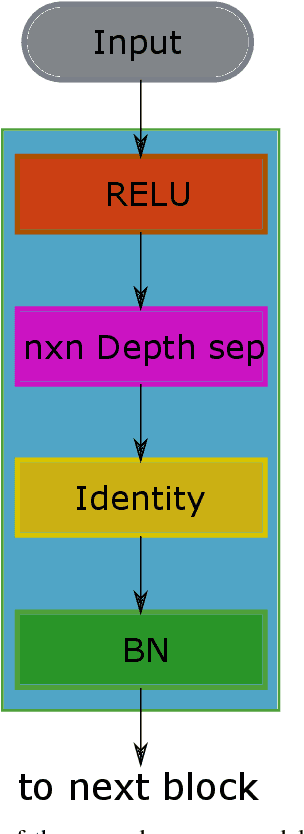
Recent progress in encoder-decoder neural network architecture design has led to significant performance improvements in a wide range of medical image segmentation tasks. However, state-of-the-art networks for a given task may be too computationally demanding to run on affordable hardware, and thus users often resort to practical workarounds by modifying various macro-level design aspects. Two common examples are downsampling of the input images and reducing the network depth to meet computer memory constraints. In this paper we investigate the effects of these changes on segmentation performance and show that image complexity can be used as a guideline in choosing what is best for a given dataset. We consider four statistical measures to quantify image complexity and evaluate their suitability on ten different public datasets. For the purpose of our experiments we also propose two new encoder-decoder architectures representing shallow and deep networks that are more memory efficient than currently popular networks. Our results suggest that median frequency is the best complexity measure in deciding about an acceptable input downsampling factor and network depth. For high-complexity datasets, a shallow network running on the original images may yield better segmentation results than a deep network running on downsampled images, whereas the opposite may be the case for low-complexity images.
Large-scale Bilingual Language-Image Contrastive Learning
Apr 15, 2022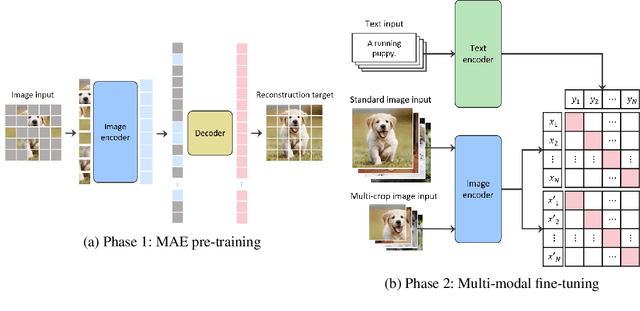
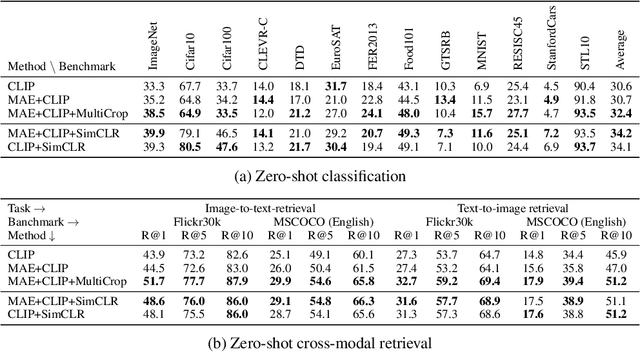
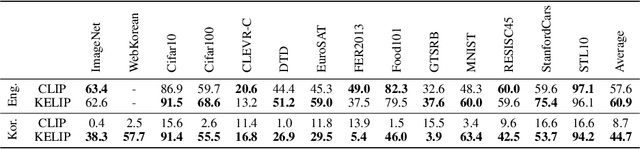
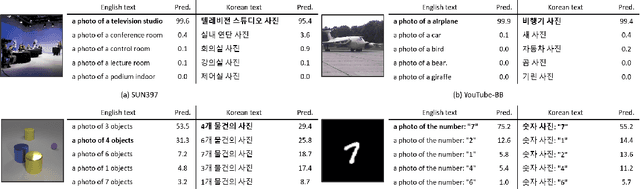
This paper is a technical report to share our experience and findings building a Korean and English bilingual multimodal model. While many of the multimodal datasets focus on English and multilingual multimodal research uses machine-translated texts, employing such machine-translated texts is limited to describing unique expressions, cultural information, and proper noun in languages other than English. In this work, we collect 1.1 billion image-text pairs (708 million Korean and 476 million English) and train a bilingual multimodal model named KELIP. We introduce simple yet effective training schemes, including MAE pre-training and multi-crop augmentation. Extensive experiments demonstrate that a model trained with such training schemes shows competitive performance in both languages. Moreover, we discuss multimodal-related research questions: 1) strong augmentation-based methods can distract the model from learning proper multimodal relations; 2) training multimodal model without cross-lingual relation can learn the relation via visual semantics; 3) our bilingual KELIP can capture cultural differences of visual semantics for the same meaning of words; 4) a large-scale multimodal model can be used for multimodal feature analogy. We hope that this work will provide helpful experience and findings for future research. We provide an open-source pre-trained KELIP.
Alternative design of DeepPDNet in the context of image restoration
Feb 20, 2022
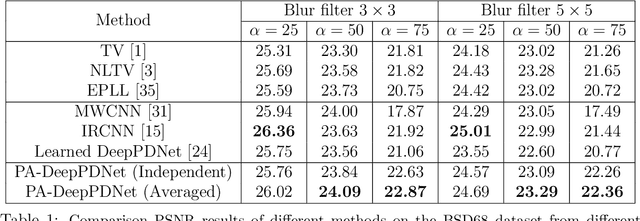
This work designs an image restoration deep network relying on unfolded Chambolle-Pock primal-dual iterations. Each layer of our network is built from Chambolle-Pock iterations when specified for minimizing a sum of a $\ell_2$-norm data-term and an analysis sparse prior. The parameters of our network are the step-sizes of the Chambolle-Pock scheme and the linear operator involved in sparsity-based penalization, including implicitly the regularization parameter. A backpropagation procedure is fully described. Preliminary experiments illustrate the good behavior of such a deep primal-dual network in the context of image restoration on BSD68 database.
Interpreting deep learning output for out-of-distribution detection
Nov 07, 2022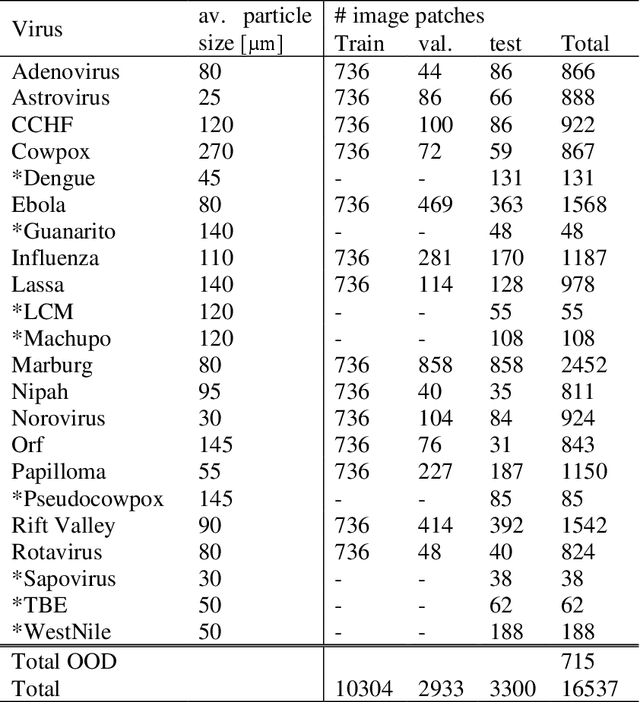
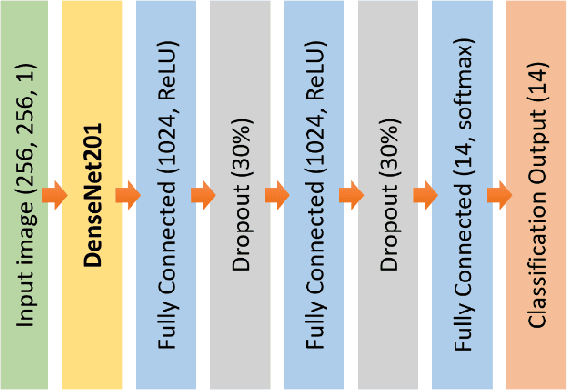
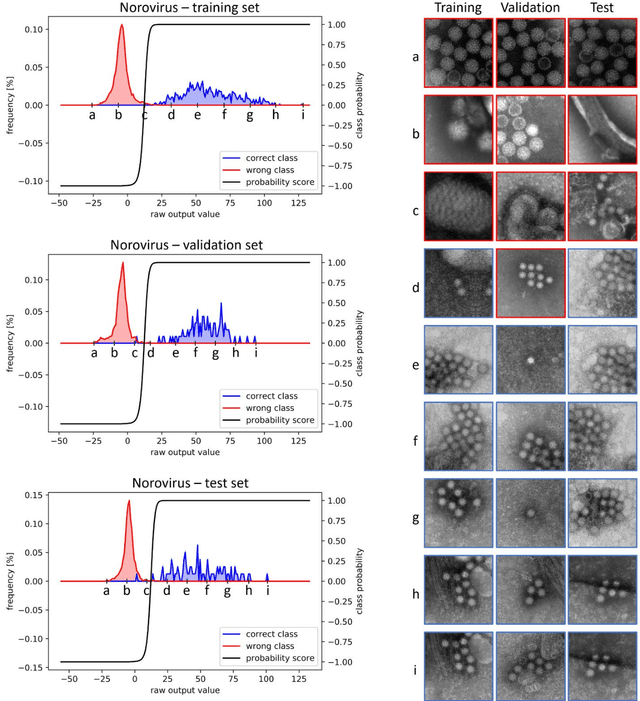
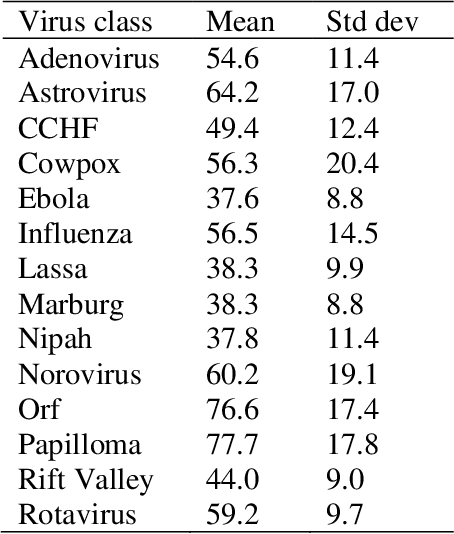
Commonly used AI networks are very self-confident in their predictions, even when the evidence for a certain decision is dubious. The investigation of a deep learning model output is pivotal for understanding its decision processes and assessing its capabilities and limitations. By analyzing the distributions of raw network output vectors, it can be observed that each class has its own decision boundary and, thus, the same raw output value has different support for different classes. Inspired by this fact, we have developed a new method for out-of-distribution detection. The method offers an explanatory step beyond simple thresholding of the softmax output towards understanding and interpretation of the model learning process and its output. Instead of assigning the class label of the highest logit to each new sample presented to the network, it takes the distributions over all classes into consideration. A probability score interpreter (PSI) is created based on the joint logit values in relation to their respective correct vs wrong class distributions. The PSI suggests whether the sample is likely to belong to a specific class, whether the network is unsure, or whether the sample is likely an outlier or unknown type for the network. The simple PSI has the benefit of being applicable on already trained networks. The distributions for correct vs wrong class for each output node are established by simply running the training examples through the trained network. We demonstrate our OOD detection method on a challenging transmission electron microscopy virus image dataset. We simulate a real-world application in which images of virus types unknown to a trained virus classifier, yet acquired with the same procedures and instruments, constitute the OOD samples.
ImageCAS: A Large-Scale Dataset and Benchmark for Coronary Artery Segmentation based on Computed Tomography Angiography Images
Nov 03, 2022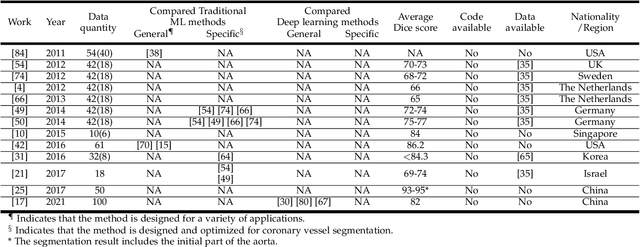
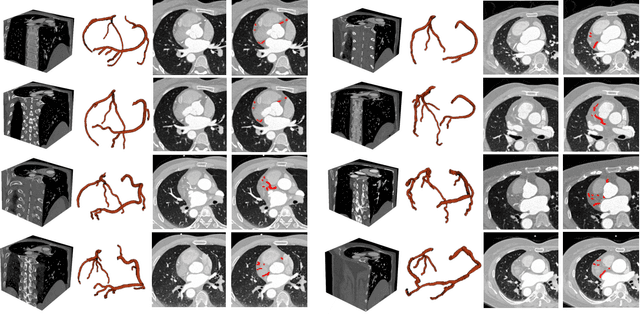
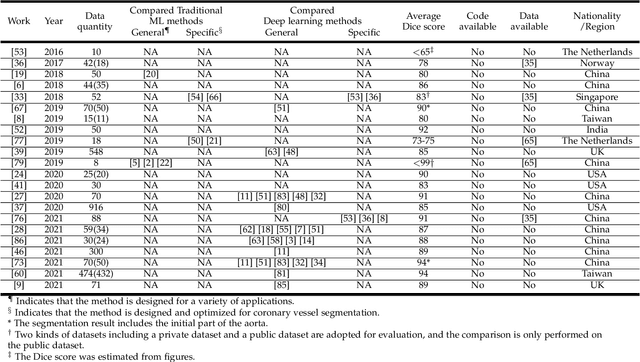

Cardiovascular disease (CVD) accounts for about half of non-communicable diseases. Vessel stenosis in the coronary artery is considered to be the major risk of CVD. Computed tomography angiography (CTA) is one of the widely used noninvasive imaging modalities in coronary artery diagnosis due to its superior image resolution. Clinically, segmentation of coronary arteries is essential for the diagnosis and quantification of coronary artery disease. Recently, a variety of works have been proposed to address this problem. However, on one hand, most works rely on in-house datasets, and only a few works published their datasets to the public which only contain tens of images. On the other hand, their source code have not been published, and most follow-up works have not made comparison with existing works, which makes it difficult to judge the effectiveness of the methods and hinders the further exploration of this challenging yet critical problem in the community. In this paper, we propose a large-scale dataset for coronary artery segmentation on CTA images. In addition, we have implemented a benchmark in which we have tried our best to implement several typical existing methods. Furthermore, we propose a strong baseline method which combines multi-scale patch fusion and two-stage processing to extract the details of vessels. Comprehensive experiments show that the proposed method achieves better performance than existing works on the proposed large-scale dataset. The benchmark and the dataset are published at https://github.com/XiaoweiXu/ImageCAS-A-Large-Scale-Dataset-and-Benchmark-for-Coronary-Artery-Segmentation-based-on-CT.
Unsupervised diffeomorphic cardiac image registration using parameterization of the deformation field
Aug 28, 2022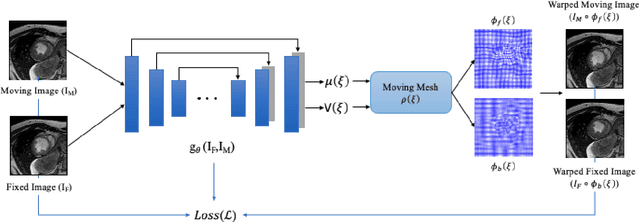
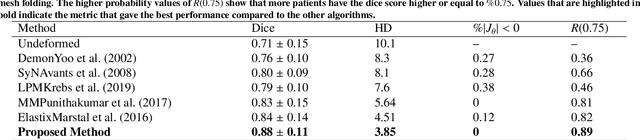

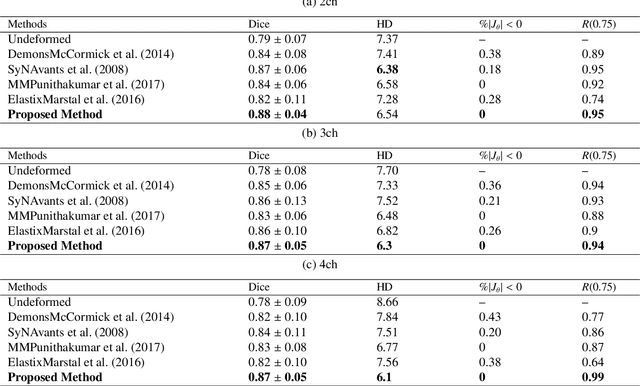
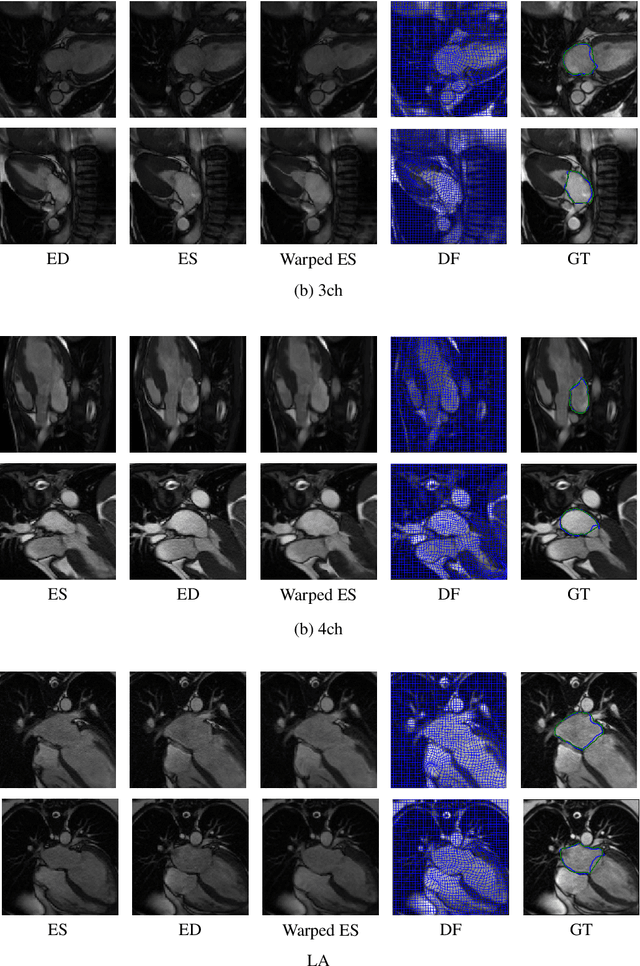
This study proposes an end-to-end unsupervised diffeomorphic deformable registration framework based on moving mesh parameterization. Using this parameterization, a deformation field can be modeled with its transformation Jacobian determinant and curl of end velocity field. The new model of the deformation field has three important advantages; firstly, it relaxes the need for an explicit regularization term and the corresponding weight in the cost function. The smoothness is implicitly embedded in the solution which results in a physically plausible deformation field. Secondly, it guarantees diffeomorphism through explicit constraints applied to the transformation Jacobian determinant to keep it positive. Finally, it is suitable for cardiac data processing, since the nature of this parameterization is to define the deformation field in terms of the radial and rotational components. The effectiveness of the algorithm is investigated by evaluating the proposed method on three different data sets including 2D and 3D cardiac MRI scans. The results demonstrate that the proposed framework outperforms existing learning-based and non-learning-based methods while generating diffeomorphic transformations.
Exploring the GLIDE model for Human Action-effect Prediction
Aug 01, 2022
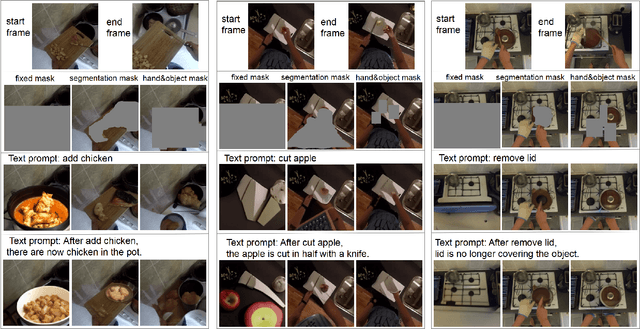



We address the following action-effect prediction task. Given an image depicting an initial state of the world and an action expressed in text, predict an image depicting the state of the world following the action. The prediction should have the same scene context as the input image. We explore the use of the recently proposed GLIDE model for performing this task. GLIDE is a generative neural network that can synthesize (inpaint) masked areas of an image, conditioned on a short piece of text. Our idea is to mask-out a region of the input image where the effect of the action is expected to occur. GLIDE is then used to inpaint the masked region conditioned on the required action. In this way, the resulting image has the same background context as the input image, updated to show the effect of the action. We give qualitative results from experiments using the EPIC dataset of ego-centric videos labelled with actions.
Deep Learning based Super-Resolution for Medical Volume Visualization with Direct Volume Rendering
Oct 14, 2022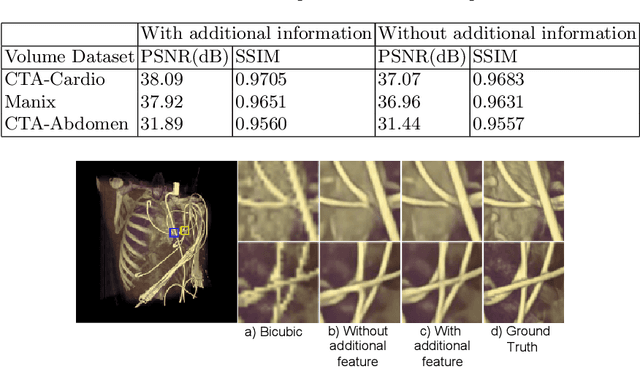
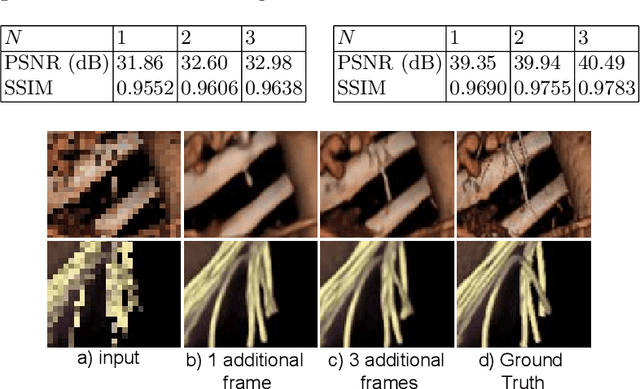
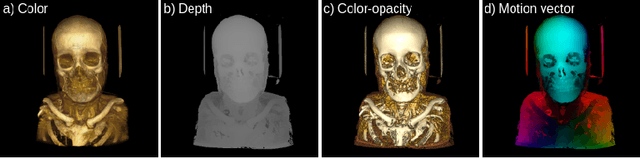
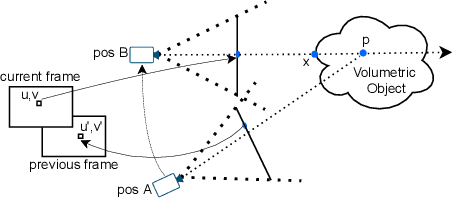


Modern-day display systems demand high-quality rendering. However, rendering at higher resolution requires a large number of data samples and is computationally expensive. Recent advances in deep learning-based image and video super-resolution techniques motivate us to investigate such networks for high-fidelity upscaling of frames rendered at a lower resolution to a higher resolution. While our work focuses on super-resolution of medical volume visualization performed with direct volume rendering, it is also applicable for volume visualization with other rendering techniques. We propose a learning-based technique where our proposed system uses color information along with other supplementary features gathered from our volume renderer to learn efficient upscaling of a low-resolution rendering to a higher-resolution space. Furthermore, to improve temporal stability, we also implement the temporal reprojection technique for accumulating history samples in volumetric rendering.
Automated Learning for Deformable Medical Image Registration by Jointly Optimizing Network Architectures and Objective Functions
Mar 14, 2022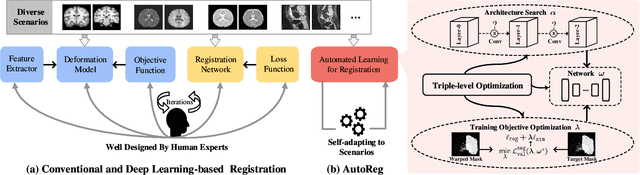
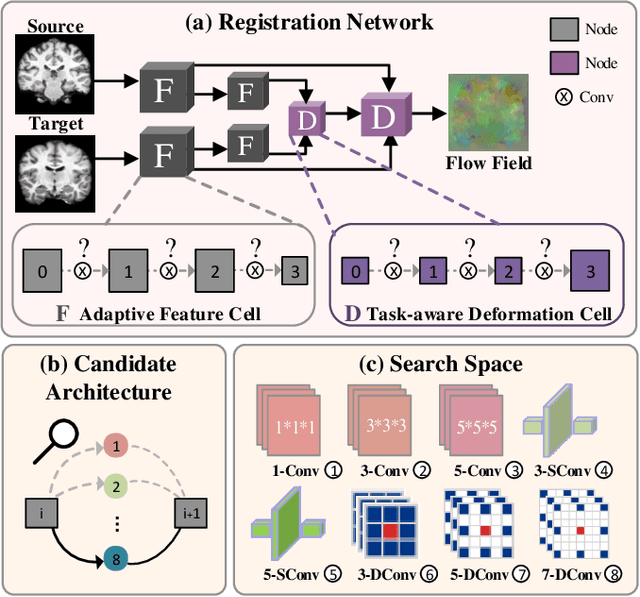
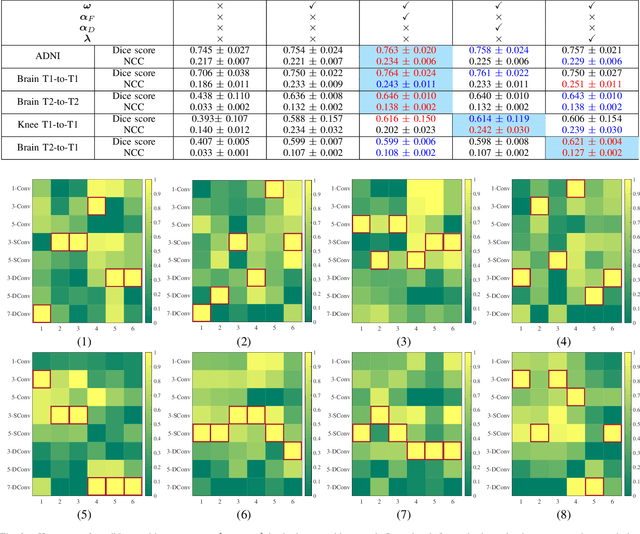
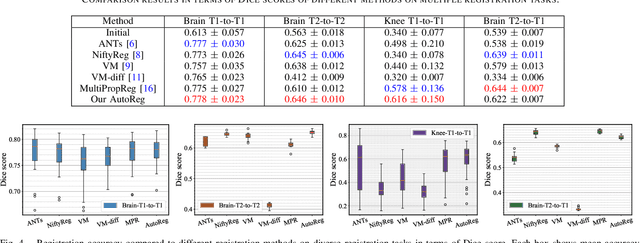
Deformable image registration plays a critical role in various tasks of medical image analysis. A successful registration algorithm, either derived from conventional energy optimization or deep networks requires tremendous efforts from computer experts to well design registration energy or to carefully tune network architectures for the specific type of medical data. To tackle the aforementioned problems, this paper proposes an automated learning registration algorithm (AutoReg) that cooperatively optimizes both architectures and their corresponding training objectives, enabling non-computer experts, e.g., medical/clinical users, to conveniently find off-the-shelf registration algorithms for diverse scenarios. Specifically, we establish a triple-level framework to deduce registration network architectures and objectives with an auto-searching mechanism and cooperating optimization. We conduct image registration experiments on multi-site volume datasets and various registration tasks. Extensive results demonstrate that our AutoReg may automatically learn an optimal deep registration network for given volumes and achieve state-of-the-art performance, also significantly improving computation efficiency than the mainstream UNet architectures (from 0.558 to 0.270 seconds for a 3D image pair on the same configuration).
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge