Sumanta Pattanaik
Efficient Neural Representation of Volumetric Data using Coordinate-Based Networks
Jan 16, 2024Abstract:In this paper, we propose an efficient approach for the compression and representation of volumetric data utilizing coordinate-based networks and multi-resolution hash encoding. Efficient compression of volumetric data is crucial for various applications, such as medical imaging and scientific simulations. Our approach enables effective compression by learning a mapping between spatial coordinates and intensity values. We compare different encoding schemes and demonstrate the superiority of multi-resolution hash encoding in terms of compression quality and training efficiency. Furthermore, we leverage optimization-based meta-learning, specifically using the Reptile algorithm, to learn weight initialization for neural representations tailored to volumetric data, enabling faster convergence during optimization. Additionally, we compare our approach with state-of-the-art methods to showcase improved image quality and compression ratios. These findings highlight the potential of coordinate-based networks and multi-resolution hash encoding for an efficient and accurate representation of volumetric data, paving the way for advancements in large-scale data visualization and other applications.
Deep Learning based Super-Resolution for Medical Volume Visualization with Direct Volume Rendering
Oct 14, 2022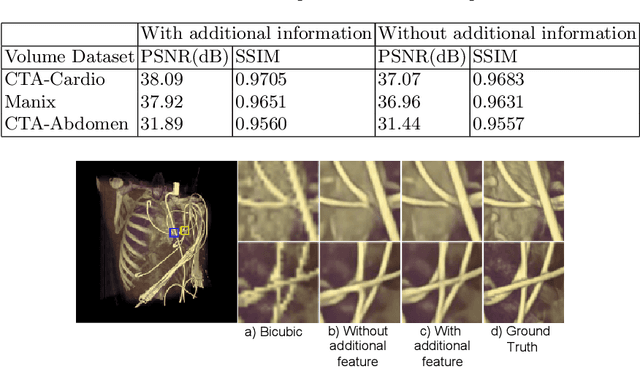
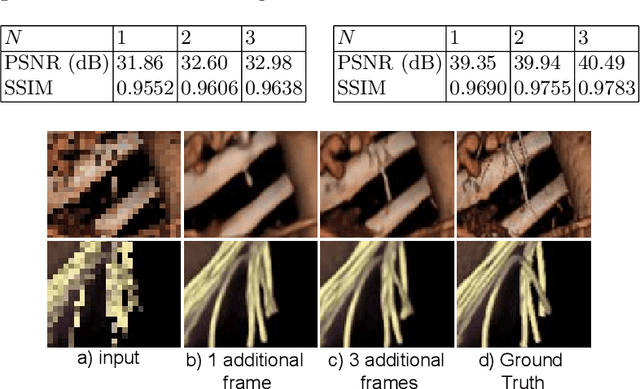
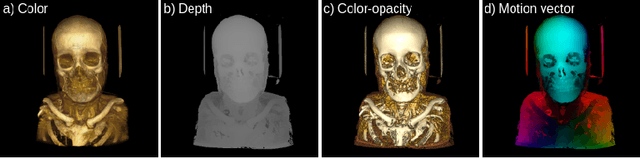
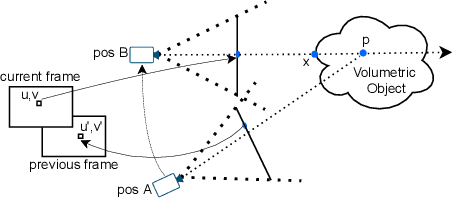


Abstract:Modern-day display systems demand high-quality rendering. However, rendering at higher resolution requires a large number of data samples and is computationally expensive. Recent advances in deep learning-based image and video super-resolution techniques motivate us to investigate such networks for high-fidelity upscaling of frames rendered at a lower resolution to a higher resolution. While our work focuses on super-resolution of medical volume visualization performed with direct volume rendering, it is also applicable for volume visualization with other rendering techniques. We propose a learning-based technique where our proposed system uses color information along with other supplementary features gathered from our volume renderer to learn efficient upscaling of a low-resolution rendering to a higher-resolution space. Furthermore, to improve temporal stability, we also implement the temporal reprojection technique for accumulating history samples in volumetric rendering.
Robust and fully automated segmentation of mandible from CT scans
Feb 23, 2017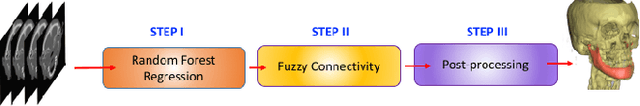
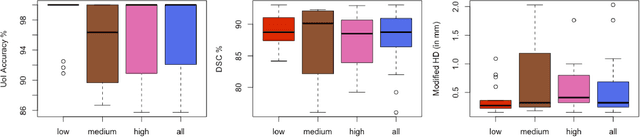
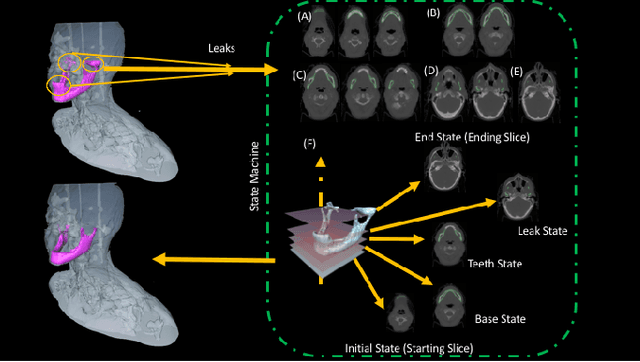
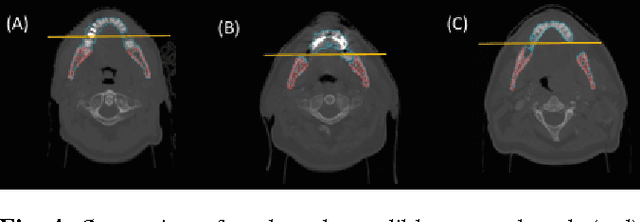
Abstract:Mandible bone segmentation from computed tomography (CT) scans is challenging due to mandible's structural irregularities, complex shape patterns, and lack of contrast in joints. Furthermore, connections of teeth to mandible and mandible to remaining parts of the skull make it extremely difficult to identify mandible boundary automatically. This study addresses these challenges by proposing a novel framework where we define the segmentation as two complementary tasks: recognition and delineation. For recognition, we use random forest regression to localize mandible in 3D. For delineation, we propose to use 3D gradient-based fuzzy connectedness (FC) image segmentation algorithm, operating on the recognized mandible sub-volume. Despite heavy CT artifacts and dental fillings, consisting half of the CT image data in our experiments, we have achieved highly accurate detection and delineation results. Specifically, detection accuracy more than 96% (measured by union of intersection (UoI)), the delineation accuracy of 91% (measured by dice similarity coefficient), and less than 1 mm in shape mismatch (Hausdorff Distance) were found.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge